Short Communication
CLONING OF BRAZZEIN GENE (SWEET PROTEIN) INTO PBI121 VECTOR THROUGH PROKARYOTIC EXPRESSION SYSTEM
B. Saleh, N. Huma, A. Azhar and S. Galani*.
Karachi Institute of Biotechnology and Genetic Engineering, University of Karachi, 75270, Pakistan.
Correspoding author’s email : Saddia.galani@kibge.edu.pk
ABSTRACT
Brazzein- a low-calorie natural sweetener is an effective substitute for deleterious effects of sucrose. Uptill now, limited production of recombinant brazzein protein using heterologous expression system along with solubility and low yield challenges are reported. Foreseeing the potential of brazzein gene in therapeutics, molecular strategies are required to optimize production of recombinant brazzein using different prokaryotic and eukaryotic expression systems. Therefore, in this study brazzein gene synthesized from Bio Basic Inc. (BBI) which was provided in pUC57 cloning vector. The gene was retrieved from cloning vector by performing PCR andcloned in plant expression vector pBI121 with CaMV35S promoter region. Heat shock or calcium chloride method was optimized for transformation into E. coli (DH5-alpha) with recombinant vector pBI121. Kanamycin resistence selection, colony PCR and transformation efficiency analysis were performed to analyze the successful cloning procedure. Transformed cultures having cloned brazzein gene in pBI121 vector with CaMV35S promoter could be used to transform different plants such as, sorghum, sugarcane and turnip for sweet taste enhancement in future and may provide plateform for commercial scale production of therapeutic proteins using crop plants as bioreactor.
Keywords: Brazzein, pBI121 vector; Recombinant technology; Transgenic plants.
https://doi.org/10.36899/JAPS.2023.3.0664
Published first online February 20, 2023
INTRODUCTION
Excessive sugar consumption derived from sucrose has adverse effects on human health leading to serious pathological issues (Aragno and Mastrocola, 2017). Therefore, an effective sucrose substitute in the form of artificial sweeteners is introduced to reduce the negative impacts of medical conditions related to obesity, heart diseases and irregular metabolism. Artificial sweeteners are FDA approved synthetic chemicals, having very low or zero calories with altered nutritive content (Spencer et al., 2016). However, their usage has unsatisfactory taste perception along with health issues such as diarrhoea, abnormal rates of metabolism, glucose intolerance and cancers in humans (NIH, 2016; US Food and Drug Administration, 2014).
Considerable efforts in the past decades led to the discovery of sweet proteins which have been reported to overcome the chemical ingestion of artificial sweeteners (Shil et al., 2020). Group of sweet proteins have been isolated from different plant sources which able to induce an intense sweetness including pentadin, monellin, brazzein, thaumatin, mabinlin and neoculin (Neiers et al., 2016). Among these sweet tasting proteins, Brazzein, a sweet protein isolated from the pulp during ripening stage of Pentadiplandra brazzeana plant has attracted much attention in 1994 (Ming and Hellekant, 1994). So far, two type of brazzein genes, major and minor forms have been found which differ one another at N-terminal amino acid residue. Minor brazzein type carries this unique ability of being the sweetest (500- 1200 times than sucrose) and smallest (Mol. Wt. 6473) sweet protein having only 54 amino acid residues (Choi et al., 2021). Extraction of genetic material of this protein from native plant to perform scientific experiments are difficult due to its geographical location. Geographical barrier, unavailability of plants (Pentadiplandra brazzeana) and heterologous expression system for commercial production are major hindrances towards the scientific investigations of brazzein.
Unfortunately, molecular studies aimed at quality, taste enhancement and molecular farming with recombinant brazzein expression are quite limited (Mérillon and Ramawat, 2018). Synthetic coding sequence responsible for coding brazzein protein have been used in this study to explore its characteristics.
Keeping in view these considerations, this research work focussed to optimize the ideal expression system pBI121-brz vector cloned with cDNA of brazzein gene in E. coli Fig.2. This transformed cassette could be used for the development of transgenic plants with improved quality and taste. It will also remove geographical barrier for the development and producing transgenic for this valuable protein on commercial scale.
MATERIALS AND METHODS
This experiment was conducted at Plant Transformation Lab, Dr. A. Q. Khan Institute of Biotechnology and Genetic Engineering (KIBGE), University of Karachi, Pakistan.
Gene retrieval and isolation: Coding sequences (cDNA) of brazzein gene were identified from GenBank (GenBank: KF013250.1) and further modified for experiment. Cloning sites of BamHI, XbaI, SamI and SacI having 25ـ30 bp snucleotides for primer sequences (Designed by Oligo Calc software) were added on flanking region of identified brazzein gene sequence. cDNA brazzein gene (274 bp) Fig.1 wassynthesized by Bio Basic Inc. (BBI) and provided in pUC57 plasmid after cloning. Gene was recovered from pUC57 plasmid through specific primers (Table 1) which were added in flanking region by performing PCR with initial denaturation at 95 °C for 2 minutes followed by 30 cycles at 95 °C for 15 seconds, 55 °C for 30 seconds,72 °C for 18 seconds (1 min/kb) and final extension at 72 °C for 8 minutes. Amplified PCR product 273 bp was used in restriction digestion and ligation procedure.
Restriction digestion of PCR product: For restriction digestion, reaction mixture of 30 µl was prepared containing 12 µl Milli-Q water, 3 µl restriction enzyme buffer (10X tango), 8 µl PCR amplified product (850 ng/µl), 4 µl restriction enzyme SacI and 3 µl BamHI. Incubation in water bath was done for 3 hours at 37 °C. Results were observed at 1.4 % gel at 70 V for 30 minutes. to validate restriction digestion results of brazzein gene.
Gel extraction: Agarose gel purification was carried out for isolation of amplified gene. After electrophoresis, brazzien gene of 267 bp was cut out from gel. Separated gel was completely melted by keeping at 650C on water bath and mixed by vortex. Equal amount of phenol was added to make emulsion. Sample was centrifuged for 10 min. at 13000 rpm at room temperature. Top aqueous layer was kept separately. Equal volume of phenol-chloroform was added and centrifuged as previously mentioned. Again, aqueous layer was kept separate and 1/10 volume of 3 M sodium acetate with 2 volume ethanol (100%) was added. Tube was incubated for overnight. Next day, centrifugation was performed to pellet down DNA (Brazzien gene). Pellet was washed by 70% ethanol, incubated and allowed to air dry. Pellet was resuspended in TE buffer and stored at -200C for further use in process of ligation.
Restriction digestion of plasmid: E. coli DH5-α strain containing vector pBI121ـPro-Adh: TaPCS1:cmyc (Gong et al., 2003) had phytochelatin synthase gene (TaPCS1) in between SacI and BamHI restriction sites (purchased from Addgene (the non-profit plasmid repository organization). TaPCS1 gene was exchanged with digested complimentary DNA of brazzein gene which was derived from gel extraction after restriction digestion by using same restriction enzymes. Alkaline lysis was used for plasmid isolation with total volume of 30 µl containing 9 µl Miliq water, 3 µl 10X Tango buffer, plasmid sample (678 ηg/µl) 10 µl, restriction enzymes (3 µl BamHI and 4 µl SacI) (Ehrt and Schnappinger, 2003). Mixture was incubated at 37 °C in water bath for 3 hours. Results were confirmed through gel electrophoresis followed by purification as mentioned in previous section.
Ligation of digested pBI121 plasmid and brazzein gene: Digested pBI121ـPro-Adh: TaPCS1:cmyc plasmid and brazzein gene from BamHI and SacI were used for ligation. 10X DNA ligase buffer (2.5 µl), T4DNA ligase (2.5 µl), purified plasmid 1250 ηg/µl (10 µl) and DNA 215 ηg/µl (10 µl ) were used respectively. Mixture was incubated for overnight at 4 °C. Results were confirmed through gel electrophoresis followed by low melt specification agarose method. After ligation, ligated plasmid pBI121-brz was used for the transformation of DH5-alpha competent cells.
Bacterial transformation and transformation efficacy: Heat shock method was used to transform E. coli competent cells with 5µl of 20 ηg/µl ligated plasmid and incubated on ice for 30 minutes. After incubation, heat shock at 42°C for 1 min. was done followed by immediate transfer at ice for 2 min. For recovery of heat shocked cells, 1 ml LB medium was added and cells were incubated at 37 °C for 1 hour. Transformed cells (100µl) were spread onto kanamycin agar plate with control. Transformation efficiency was calculated by counting average no. of transformed colonies produced by spreading 100 µl heat shocked culture onto kanamycin containing plate.
Colony PCR analysis: Single transformed colony from each kanamycin containing plate was transferred to PCR tube. PCR was performed with same conditions as mentioned above except initial denaturation was done at 95 °C for 5 minutes. Result was confirmed through gel electrophoresis using 1.4 % agarose gel run at 70 V for 30 minutes.
RESULTS
Retrieval and restriction digestion of brazzein Gene: After PCR amplification, clear band of brazzein gene (273 bp) was observed in lane 2 compared with 100 bp DNA ladder in lane 1 Fig.3A. PCR amplicons of brazzein gene were than used in restriction digestion. BamHI and SacI restriction enzyme generated sticky ends for ligation experiment. Difference between digested and undigested PCR amplified product is shown in Fig.3B. 273 bp undigested and 267 bp digested gene product are present in lane 2 and 3 indicating size differences due to restriction digestion.
Restriction digestion of pBI121ـPro-Adh:TaPCS1:cmyc vector: Isolated vector was used in restriction digestion by using SacI and BamHI. Two bands of vector backbone ~2008 bp of TaPCS1 and ~13,000 bp were observed after restriction digestion in lane 2 and 3 indicated in Fig.3C. Presence of two bands after restriction digestion represent successful digestion of pBI121ـPro-Adh:TaPCS1:cmyc vector. This digested pBI121 vector and digested brazzein gene were used in ligation process.
Ligation of digested pBI121 plasmid and brazzein gene: Digested pBI121 vector and brazzein gene were used for ligation. Ligated plasmid and brazzein gene sample of ~13267 bp in lane 2 represents successful ligation. While digested plasmid (~13kb) and gene (267 bp) in lane 3 and 4 were loaded as control shown in Fig.3D. TaPCS1 gene which was initially inserted in pBI121ـPro-Adh:TaPCS1:cmyc vector is now being replaced by brazzein gene Fig.2. In the result of restriction ligation, new vector pBI121-brz was designed that could be used for transformation of bacteria.
Transformation efficiency of E. coli through heat shock method: Transformation was performed through pBI121-brz plasmid. Kanamycin containing plates were used to identify transformed culture with simple LB media plate as control. After overnight incubation, 181, 123, and 146 transformed colonies were observed on B, C and D kanamycin LB plate respectively mentioned in Fig.4. Transformation efficiency was calculated using kanamycin containing media plates having transformed colonies, that was 1.5x10 (Table 2).
Colony PCR analysis: Transformation was confirmed by antibiotic screening and colony PCR. 288 bp of colony PCR product was observed in lane 2 to 4 confirms the presence of brazzein gene Fig.3E. These bands suggested that picked colonies from supplemented plate were transformed from pBI121-brz vector.
Table 1. Details of primer sequances used in retriving brazzein gene and colony PCR.
|
Name
|
Sequence
|
Length
|
Tm°C
|
gc%
|
Product size
|
|
brz-A1 F
|
ACGGATCCTATAGCTAGACAC
|
21
|
60.6
|
47.6
|
273 bp
|
|
brz-A1 R
|
GAGAGCTCTACGTGTGTGTT
|
20
|
60.4
|
50
|
|
brz-1 F
|
CAAATTTGCTACCGGGTCAT
|
20
|
59.82
|
45
|
228 bp
|
|
brz-1 R
|
GCCTAGTTTAAAGATGCTTTGCTC
|
24
|
59.95
|
41.6
|
Table 2. Mathematical expression for calculation of transformation efficiency.
|
No. of TC identified on antibiotic supplemented plate
|
Average No. of transformed colonies
|
Transformation efficacy (TE)
|
|
Plate B
|
123
|
123+146+182 = 150
3
|
TE=TC/DS/DF
TE=150/1/0.1
=1.5x103
|
|
Plate C
|
146
|
|
Plate D
|
181
|
*TC= Transformed colonies
DS= DNA Sample
DF= Dilution factor

Fig. 1: Artificially synthesized 274 bp gene containing 165 bp coding sequence of brazzein, BamHI, Xba1, Sma1, Sac1 restriction sites and brz-A1 F, brz-A1 R, brz-1 F, brz-1 R primer sequences.
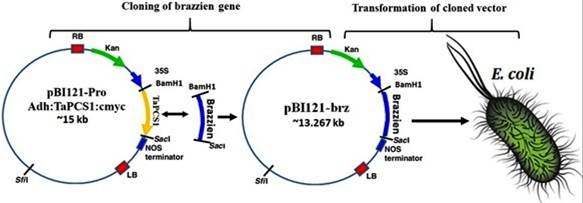
Fig. 2: During cloning of brazzein gene, digested brazzein gene from BamHI and SacI is inserted into digested pBI121ـPro-Adh:TaPCS1:cmyc plasmid with the same restriction endonuclease to make pBI121-brz plasmid. E. coli was transformed with ligated plasmid in transformation of cloned section which was confirmed though antibiotic selection and colony PCR analysis.

Fig. 3. Electrophoresis results. A) Brazzein gene amplification by PCR: lane 1 , 100 bp ladder, Lane 2, PCR Product B) Restriction digestion of brazzein gene: Lane 1, 100 bp ladder, Lane 2, brazzein gene, lane 3, digested brazzein gene. C) Restriction digestion of pBI121-Pro-Adh:TaPCS1:cmyc vector: Lane 1, 1kb DNA ladder, Lane 2, 3 and 4 contains digested pBI121 vector sample. D) Ligation of digested brazzein gene and pBI121 vector: Lane 1, 1kb ladder, lane 2, ligated Brazzein gene and vector sample, lane 3, digested vector (control), lane 4, digested brazzein gene. E) Colony PCR for the confirmation of transformation. Lane 1, 100 bp DNA ladder, lane 2 to 4, colony PCR product.
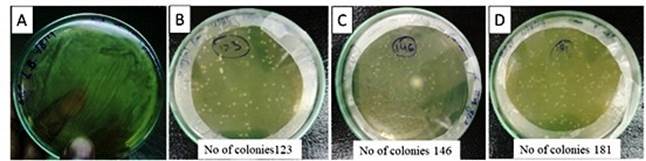
Fig. 4. E. coli DH5-α transformation by using pBI121-brz vector. A) LB plate (control): without kanamycin. Enormous bacterial growth represnet no damge or culture is living in respone of given heat shock. Antibiotic suplimented LB plate B), C) and D) for slection of trasformed becterial colonies. Pin pointed isolated colonies with distance represent the transformed culture having pBI121-brz vector.
DISCUSSION
Stability of cloned sequence is the foremost requirement for developing steady expression system to produce recombinant product. Moreover, appropriate selection and modification in microorganism, vector and expression system are also required. In this study, coding sequence of brazzein gene responsible for producing type 3 brazzein was cloned in binary expression vector pBI121. E. coli was selected for transformation due to short doubling time for efficient propagation of cloned vector (Watson et al., 2016; Gupta and Shukla, 2016). E. coli is also well studied having the ability to grow at broad ranges of temperature and pH making it convenient to handle at lab scale (Rosano and Ceccarelli, 2014). Binary expression vector pBI121 was selected as it has different elements to facilitate transformation of plant (Nakagawa et al., 2007). Previously, this vector is reported for soybean plant transformation with chitinase (chi) using cauliflower mosaic virus (CaMV) (Salehi et al., 2005). Similarly, diploid potato genotype transformation and transgenic cotyledon plants were successfully transformed by utilizing pBI121 vector (Sharma and Anjaiah, 2000).
So far, limited studies have reported production of recombinant brazzein protein using heterologous expression system along with solubility and low yield challenges. Previously, In transgenic maize high yield of recombinant brazzein was achieved but due to cross contamination low yield of purified product is reported (Neiers et al., 2016: Yun et al., 2016). Modifications with small Ubiquitin-like Modifier (SUMO) in vector still compromised production and purification of brazzein protein in E.coli. (Assadi Porter et al., 2008). Similarly, brazzein production using three Lactobacillus species and mice mammary gland was attempted yielding low gene expression with low sweet intensity (Yan et al., 2013). In K. lactis, unstable disulfide bond formation caused loss of activity and stability during protein production (Yun et al., 2016). However, development of transgenic pear producing sweet protein (thaumatin II) was also reported exhibiting alternative production strategies via transgenic vegetable with increased sweetness (Lebedev et al., 2000; Rommens et al., 2006).
It was evident from this study that 273 bp PCR product recovered from transformed culture was 100% identical with brazzein gene representing successful cloning and transformation of recombinant gene. Beside this, orientation of brz gene in PBI121 vector is correct despite multiple restriction digestions at the vector sites confirmed by DNA sequencing.
Conclusions: It is concluded that cloned brazzein gene in PbI121 vector (plant expression vector) may provide heterologous expression system as it may be used to produce for the expression of recombinant brazzien protein in various plant species irrespective of its native plant. The constructed vector may also serve for the source of this coding sequence of brazzein gene which can be retrieved and may used for other prokaryotic and eukaryotic expression systems. Currently there is need to utilize recombinant technologies to gaining maximum molecular insights towards brazzein as a sweet protein the less explored potential of brazzein gene. Utilizing brazzein as the product of interest, thus this study can potentially benefit the genetic engineering approaches required for the mass scale production of this important molecule. On the other hand, cloned brazzein gene in pBI121 vector with CaMV35S promoter, a plant expression vector Fig.2 could be used to transform different plants such as, sorghum, sugarcane and turnip for sweet taste enhancement in future and may serve as first step towards molecular farming for large scale production of recombinant protein, brazzein through plants.
REFERENCES
- Aragno, M., and Mastrocola, R. (2017). Dietary sugars and endogenous formation of advanced glycation endproducts: emerging mechanisms of disease. Nutr. 9(4), 385. https://doi.org/10.3390/nu9040385
- Assadi-Porter, F. M., Patry, S., and Markley, J. L. (2008). Efficient and rapid protein expression and purification of small high disulfide containing sweet protein brazzein in coli. Protein Expr. Purif. 58(2), 263-268.
- https://doi.org/10.1016/j.pep.2007.11.009
- Choi, H. E., Lee, J. I., Jo, S. Y., Chae, Y. C., Lee, J. H., Sun, H. J., ... and Kong, K. H. (2021). Functional expression of the sweet-tasting protein brazzein in transgenic tobacco. J. Food Sci. Technol., 42.
- https://doi.org/10.1590/fst.40521
- Ehrt, S., and Schnappinger, D. (2003). Isolation of plasmids from coli by alkaline lysis. In E. coli Plasmid Vectors, Humana Press, 75-78. https://doi.org/10.1385/1-59259-409-3:75
- Gong, J. M., Lee, D. A., and Schroeder, J. I. (2003). Long-distance root-to-shoot transport of phytochelatins and cadmium in arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 100(17), 10118-10123.
- https://doi.org/10.1073/pnas.1734072100
- Gupta, S. K., and Shukla, P. (2016). Advanced technologies for improved expression of recombinant proteins in bacteria: perspectives and applications. Crit. Rev. Biotechnol. 36(6), 1089-1098.
- https://doi.org/10.3109/07388551.2015.1084264
- Lebedev, V. G., Taran, S. A., Shmatchenko, V. V., and Dolgov, S. V. (2000). Pear transformation with the gene for supersweet protein thaumatin II. In VIII Inter. Symp. on Pear. 596, 199-202.
- https://doi.org/10.17660/ActaHortic.2002.596.27
- Ming, D., and Hellekant, G. (1994). Brazzein, a new high-potency thermostable sweet protein from pentadiplandra brazzeana. FEBS letters. 355(1), 106-108. https://doi.org/10.1016/0014-5793(94)01184-2
- Mérillon, J. M., and Ramawat, K. G. (2018). Brazzein: a natural sweetener. sweeteners-pharmacology, biotechnology and applications, Springer. 17-33. https://hal.science/hal-03248270
- National Institute of Health (2016). Artificial sweeteners and cancer. https://www.cancer.gov/about-cancer/causes-prevention/risk/diet/artificial-sweeteners-fact-sheet. Accessed August 10,2016
- Nakagawa, T., Kurose, T., Hino, T., Tanaka, K., Kawamukai, M., Niwa, Y., ... and Kimura, T. (2007). Development of series of gateway binary vectors, pGWBs, for realizing efficient construction of fusion genes for plant transformation. J. Biosci. Bioeng. 104(1), 34-41. https://doi.org/10.1263/jbb.104.34
- Neiers, F., Naumer, C., Krohn, M., and Briand, L. (2016). The recent development of a sweet-tasting brazzein and its potential industrial applications. Sweeteners; Springer: Berlin/Heidelberg, Germany . https:// DOI 10.1007/978-3-319-26478-3_2-1
- Rommens, C. M., Ye, J., Richael, C., and Swords, K. (2006). Improving potato storage and processing characteristics through all-native DNA transformation. J. Agric. Food Chem. 54(26), 9882-9887.
- https://doi.org/10.1021/jf062477l
- Rosano, G. L., and Ceccarelli, E. A. (2014). Recombinant protein expression in Escherichia coli: advances and challenges. Front. Microbiol. 5, 172. https://doi.org/10.3389/fmicb.2014.00172
- Salehi, A., Mohammadi, M., Okhovvat, S. M., and Omidi, M. (2005). Chitinase gene transformation through Agrobacterium and its explanation in soybean in order to induce resistance to root rot caused by Rhizoctonia solani. Commun. Agric. Appl. Biol. 70(3), 399-406. PMID: 16637205
- Shil, A., Olusanya, O., Ghufoor, Z., Forson, B., Marks, J., and Chichger, H. (2020). Artificial sweeteners disrupt tight junctions and barrier function in the intestinal epithelium through activation of the sweet taste receptor, T1R3. Nutr. 12(6), 1862. https://doi.org/10.3390/nu12061862
- Sharma, K. K., and Anjaiah, V. (2000). An efficient method for the production of transgenic plants of peanut (Arachis hypogaea) through Agrobacterium tumefaciens-mediated genetic transformation. Plant Sci. 159(1), 7-19. https://doi.org/10.1016/S0168-9452(00)00294-6
- Spencer, M., Gupta, A., Van Dam, L., Shannon, C., Menees, S., and Chey, W. D. (2016). Artificial sweeteners: a systematic review and primer for gastroenterologists. J. Neurogastroenterol Motil. 22(2), 168.
- https://doi: 10.5056/jnm15206
- US Food and Drug Administration (2014). High-Intensity Sweeteners. https://www.fda.gov/food/ food- additives-petitions/high-intensity-sweeteners. Accessed May 19,2014.
- Watson, M. R., Lin, Y. F., Hollwey, E., Dodds, R. E., Meyer, P., and McDowall, K. J. (2016). An improved binary vector and Escherichia coli strain for Agrobacterium tumefaciens-mediated plant transformation. G3: Genes, Genomes, Genetics. 6(7), 2195-2201. https://doi.org/10.1534/g3.116.029405
- Yan, S., Song, H., Pang, D., Zou, Q., Li, L., Yan, Q., . and Lai, L. (2013). Expression of plant sweet protein brazzein in the milk of transgenic mice. PloS One, 8(10), e76769. https://doi.org/10.1371/journal.pone.0076769
- Yun, C. R., Kong, J. N., Chung, J. H., Kim, M. C., and Kong, K. H. (2016). Improved secretory production of the sweet-tasting protein, brazzein, in Kluyveromyces lactis. J. Agric. Food Chem. 64(32), 6312-6316.
- https://doi.org/10.1021/acs.jafc.6b02446
|





