Short Communication
POLYMORPHISMS IN BMPR1B AND INHβA GENES ARE ASSOCIATED WITH LITTER SIZE IN INDIGENOUS SHEEP OF BANGLADESH
R. Basrin1,2, R. Khanom1, M. M. Hossain1, M. S. Islam1,2, A. K. F. H. Bhuiyan1 and M. S. A. Bhuiyan1*
1Department of Animal Breeding and Genetics, Bangladesh Agricultural University, Mymensingh-2202, Bangladesh
2Department of Livestock Services, Farmgate, Dhaka-1215, Bangladesh
*Corresponding author’s Email: msabhuiyan@gmail.com
ABSTRACT
The purpose of the study was to detect the genetic polymorphisms in the fragments of four transforming growth factors β superfamily genes (BMPR1B, INHβA, BMP15 and GDF9) and to investigate their possible association with litter size trait in Bangladeshi indigenous sheep population. Previously reported six significantly associated single nucleotide polymorphisms (SNPs) of those aforementioned genes were included in this study. A panel of 12 samples were sequenced from each gene fragment for polymorphism detection. Multiple sequence alignment revealed polymorphic sites only for BMPR1B (c.746A>G) and INHβA (c.218A>G) gene fragments in indigenous sheep of Bangladesh. However, four other polymorphisms located in GDF9 [c.260G>A (FecG1) and c.1184C>T (FecG8)] and BMP15 [c.69C>T (FecX-H and c.718C>T (FecX-G)] gene fragments were absent in the studied populations. PCR-RFLP and resequencing were employed for genotyping of BMPR1B and INHβA polymorphisms, respectively. The mean litter size of ++, +B and BB genotypes of BMPR1B gene were 1.19±0.07, 1.44±0.09 and 2.13±0.09, respectively. Besides, the average litter size of AA, AG and GG genotypes of INHβA gene mutation was 2.11±0.39, 1.70±0.30 and 2.83±0.40, respectively. Litter size had significant association with BMPR1B (p<0.05) and INHβA (p<0.01) genotypes and regional population specific and therefore, could be used as molecular markers for improving prolificacy of Bangladeshi indigenous sheep.
Key words: Single nucleotide polymorphism, allele, genotype, sheep, prolificacy, Bangladesh
https://doi.org/10.36899/JAPS.2023.2.0632
Published first online December 18, 2022
INTRODUCTION
Prolificacy is an economically important reproductive trait that directly involved with production efficiency in sheep. Previous studies reported that prolificacy or litter size in sheep is regulated genetically either by a single gene with major effect or a set of genes each having small effect (Souza et al. 2001, Davis et al. 2002). Four members of transforming growth factors β (TGFβ) superfamily genes such as growth differentiation factor 9 (GDF9), bone morphogenetic protein receptor, type 1B (BMPR1B), bone morphogenetic protein 15 (BMP15) and Inhibin βA (INHβA) play pivotal roles in the control of folliculogenesis, ovulation rate and litter size in different sheep breeds (Hanrahan et al. 2004, Fabre et al. 2006). Hence, these genes are already proved as potential candidate genes for better prolificacy in sheep. BMPRIB or FecB or Booroola gene mutations was first reported in Merino sheep that influenced prolificacy traits (Davis et al. 1982). The A746G mutation in exon 6 of the BMPR1B gene resulted high prolificacy in Booroola Merino, Small Tailed Han and Hu, and Indian Garole sheep (Davis et al. 2002, Chu et al. 2007a and 2011). Nine mutations (G1 to G8, FecTT) so far identified in GDF9 or FecG gene where G1 and G8 had association with higher ovulation rate and litter size in different sheep breeds (Moradband et al. 2011, Zuo et al. 2013, Kolosov et al. 2015). In addition, polymorphisms in BMP15 and INHβA genes had significant effects on litter size in several sheep breeds (Hanrahan et al. 2004, Liu et al. 2006, Chu et al. 2007a and 2007b).
Indigenous sheep is an important small ruminant species after goat in Bangladesh which is being reared mainly for mutton production. They are foundthroughout the country but their concentration is relatively higher in coastal regions (COR), Sundarban delta regions (SDR), barind tracts (BAT) and Jamuna River basin (JRB) areas (Hasan and Talukder, 2011). Indigenous sheep of Bangladesh are capable of bi-annual lambing with multiple births where the average lambing interval was reported to be 179±1.06 days (Islam et al. 2018). They alsohave withstood ability to utilize low quality feed stuff and well adapted under tropical climatic conditions. Alongside, the breeding tract of highly prolific small sized Garole sheep is in and around the Sundarban delta regions of Bangladesh and India (Sharma et al. 1999) and is considered as the progenitor of high prolificacy Booroola (FecB) gene. It is notable to mention that prolificacy traits differed largely within and among the indigenous sheep populations of Bangladesh (Hassan and Talukder, 2011, Islam et al. 2018). Therefore, it would be worthwhile to unveil the genetic architecture of candidate genes for prolificacy traits, their polymorphisms and associations with litter size traits in Bangladeshi indigenous sheep. However, polymorphisms in TGFβ superfamily genes and their association with prolificacy traits remain unexplored until to date. Here, the objectives of this study were to screened polymorphisms in six selected gene fragments of four TGFβ superfamily genes and their possible associations with litter size trait in indigenous sheep of Bangladesh.
MATERIALS AND METHODS
Blood sampling and DNA extraction: This study was conducted as per guidelines of the university research system and approval was taken from the university ethical committee (No. 838/BAURES/2020 ESRC/AH/01). Blood samples were collected from indigenous sheep populated regions namely COR (n=57), SDR (n=36) and JRB (n=64) areas having both high and low fecundity ewes. From each flock, two or three blood samples were collected from unrelated mature individuals those having litter size records up to 3rd parities. A total of 157 blood samples (3-4 ml) were collected aseptically from jugular vein using vacutainer with EDTA as anticoagulant. DNA was extracted using Prime PrepTM genomic DNA isolation kit (GeNet Bio Co. Ltd., South Korea). The concentration and purity of extracted DNA samples were quantified by Nanodrop spectrophotometer (ND2000,Thermo Fisher Scientific, USA).
PCR amplification and sequencing: The selected polymorphisms were c.746A>G of BMPR1B, c. 218 A>G mutation of INHβA, c.69C>T and c.718C>T polymorphisms of BMP15 and c. 260G>A and c.1184C>T of GDF9. Accordingly, six primer pairs were selected based on previously reported sequence information by Hanrahan et al. (2004), Chu et al. (2007a and 2011) and Zuo et al. (2013).PCR amplification was carried out in 20 µl volume containing 10× buffer, 1.5 mM MgCl2, 10 mM dNTP mixtures, 10 µM of each primer (Bionics, South Korea), 1 U prime Taq DNA polymerase (GeNet Bio, South Korea) and ~50 ng of genomic DNA. The gene fragments were amplified using Biometra T-gradient thermocycler with the following cyclic conditions: initial denaturation at 94 °C for 10 min, followed by 35 to 37 cycles for denaturation, annealing and extension steps at 94 °C for 30 sec, at 59-62 °C for 30 sec and at 72 °C for 45 sec, respectively and a final extension at 72 °C for 10 min. Details on primer sequence information, amplicon size and optimized annealing temperature are presented in Supplementary Table S1.A set of purified PCR products (12 samples) from each gene fragment were then sequenced both in forward and reverse directions from a commercial sequencing service provider (Solgent Co. Ltd., South Korea). Sequencing was performed by using automated Genetic Analyzer 3130 ×l (Applied Biosystems, USA). The generated raw sequences were aligned and edited using bioinformatic tools BioEdit (Hall, 1999) and CLASTALW (https://www.genome.jp/tools-bin/clustalw) for mutation scoring in indigenous sheep of Bangladesh as well as to ascertain the sequence of respective gene.
Genotyping: The PCR-RFLP method was employed for genotyping ofc.746A>G polymorphism of BMPR1B gene.The PCR products were digested with 6 U of Ava II restriction enzyme (New England Biolabs, MA, USA) at 37°C for 3 hours in a 20µl of reaction volume containing 10 µl of PCR product, 0.60 µl of AvaII enzyme (10 U/µl), 2.0 µl of buffer and 7.40 µl of deionized water. The resulting fragments were separated by 3% agarose gel. However, re-sequencing of PCR fragments was performed for genotyping of the animals for c.218A>G in INHβA gene polymorphism due to lack of restriction site.
Statistical analysis: Genotypic and allelic frequencies were estimated based on the resultant PCR-RFLP fragment pattern and sequence information according to Falconer and Mackay (1996). Single marker association analysis was performed to evaluate the relationships between genotypes and average litter size using general linear model (GLM) procedure of SAS for windows 9.1.3. Significant difference between means was performed using Duncan’s Multiple Range Test (DMRT). The following statistical model was employed for association analysis;
Yijk = µ + Li + Gj + Pk + eijk
Where Yijkis the phenotypic value of litter size; µ is the population mean, Li is the fixed effect of ith locations (i = 1, 2, 3), Gj is the fixed effect of jth genotypes (j = 1, 2, 3), Pk is the effect of kth parity (k = 1, 2, 3) and eijk is the random residual error of each observation.
RESULTS AND DISCUSSION
PCR amplification, polymorphism detection and genotyping: The generated DNA sequences were aligned with NCBI database using BLAST to ascertain the position of six gene fragments in the sheep genome. The multiple sequence alignment and chromatogram data revealed polymorphic sites only for BMPRIB (c.746A>G) and INHβA (c.218A>G) gene fragments in indigenous sheep of Bangladesh (Figure 1). However, sequence alignment showed four other selected mutations in GDF9 [c.260G>A (FecG1) and c.1184C>T (FecG8)] and BMP15 [c.69C>T (FecXG and c.718C>T (FecXHIL)] gene fragments were monomorphic in the studied populations that limits for further PCR-RFLP genotyping. Similar to the present findings, Hossain et al. (2020) reported the absence of polymorphisms in BMP15 and GDF9 genes of indigenous sheep of Bangladesh. This finding partially agrees with the results of Roy et al. (2011) who reported all considered loci of BMP15 and GDF9 genes were monomorphic except two (G1 and G4) of GDF9 were found to be polymorphic in Indian Bonpala sheep. However, several polymorphisms were reported in GDF9 and BMP15 genes among the sheep breeds of Cambridge, Belclare, Baluchi, Small Tailed Han and Bayanbulak (Hanrahan et al.2004, Chu et al. 2007a, Moradband et al. 2011 and Zuo et al. 2013) and is inconsistent to this study. Taken together, the reported polymorphisms in GDF9 and BMP15 genes were possibly sheep breed or population specific and were absent in the indigenous sheep of Bangladesh.According to Zhu et al. (2015), the population or breed specific SNP which is common in the individuals of one geographic region might be much rarer or absent in another. Therefore, further investigation was limited with only the polymorphisms inBMPRIB and INHβA gene fragments to detect possible association with prolificacy traits in indigenous sheep.
PCR-RFLP genotyping of c.746A>G polymorphism detected three genotypes as wild type (190 bp), heterozygous (190, 160 and 30 bp) and homozygous mutants (160 and 30 bp) in indigenous sheep of Bangladesh for BMPR1Bgene fragment(Figure 1). Besides, resequencing revealed one A to G nucleotide silent mutation at base 218 of exon 2 of INHβA gene and thereby the resultant genotypes AA, AG and GG were detected in indigenous sheep of Bangladesh (Figure 1). The present findings are in agreement with the previous reports of Chu et al. (2007a) in Chinese Hu and Small Tailed Han sheep, Roy et al. (2011) in Indian Bonpala sheep, Sudhakar et al. (2013) in Indian Nilgiri sheep, Zuo et al. (2013) in Chinese Bayanbulak sheep and Moradband et al. (2011) in Iranian Baluchi sheep. However, previous studies reported the absence of FecB mutation in various sheep breeds such as Dorper, Texel, Dorset and South African Meat Murray sheep (Chu et al. 2009) and contradicts to this study. On the other hand, c218A>G polymorphism of INHβA gene was reported in Chinese Small Tailed Han and Hu sheep as well as in German Mutton Merino and Corriedale sheep (Chu et al. 2007b) and support the present findings. It is found that the above mentioned two polymorphisms in BMPR1B and INHβA genes were predominant in different Asian sheep breeds including Bangladeshi sheep and are considered as major genes for influencing the high lambing rate in sheep and could be used as molecular markers for improving prolificacy traits in indigenous sheep of Bangladesh.
Genotype and allele frequencies: Genotype and allele frequencies of the BMPR1B and INHβA gene mutations are presented in Table 1. For c.746A>G polymorphism of BMPR1B gene, the ++, +B and BB genotype frequencies were 0.31, 0.26, and 0.43, respectively. On the other hand, frequencies of genotypes AA, AG and GG for INHβA gene mutation were 0.36, 0.40, and 0.24, respectively in the studied populations. The genotype and allele frequencies observed in the present study differed slightly from those estimated in Iranian Kalehkoohi (Mahdavi et al. 2014), Indian Bonpala (Roy et al. 2011) and Chinese sheep breeds Hu and Small Tailed Han (Chu et al. 2007a) for c.746A>G polymorphism of BMPR1B gene. Two reasons such as the number of samples investigated and the distribution of polymorphisms in the respective breed or population could explain these differences among the studies. Moreover, the genotype and allele frequencies of INHβA gene for c.218A>G polymorphism in Kazakh sheep population were 0.36, 0.50 and 0.15, and 0.55 and 045, respectively (Zhao et al. 2019) and are very close to the present findings. However, compare to our finding, genotype and allele frequencies varied largely in Small Tailed Han, Hu, Texel, Dorset, German Mutton Merino and Corriedale sheep breeds for the aforesaid INHβA mutation (Chu et al. 2007b). Allele and genotype frequencies are breed or population specific and are expected to change across generations in a population with the influence of gene flow and mutation (Gillespie, 2004).
Effects of BMPR1Band INHβA gene mutation on litter size in indigenous sheep: Association analysis revealed that litter size was significantly influenced by BMPR1B (p<0.05) and INHβA (p<0.01) genotypes (Table 2). The mean litter size of ++, +B and BB genotypes of BMPR1B gene were 1.19±0.07, 1.44±0.09 and 2.13±0.09, respectively. Besides, the mean litter size of AA, AG and GG genotypes of INHβA gene were 2.11±0.39, 1.70±0.30 and 2.83±0.40, respectively (Table 2). Indigenous ewes with homozygous mutant genotypes (BB) had 0.94 more lambs per birth than those with wild genotype (++) in BMPR1B gene polymorphism. In INHβA gene, GG genotypes had 0.72 more lambs than the genotype AA. More particularly, the effect of the BMPR1B gene polymorphism was larger than that of the INHβA gene on the litter size trait in indigenous sheep of Bangladesh. Irrespective of the genotype of animal, the litter size performance of indigenous sheep of Bangladesh ranged between 1.60±0.20 and 1.90±0.44 (Hasan and Talukder, 2011). Islam et al. (2018) found relatively higher litter size as 1.99 ±0.02 in indigenous sheep populations of Bangladesh. The above stated results support the present findings. The present results are in accordance to the findings of Mishra et al. (2009), Chu et al. (2011), Maskur et al. (2016) who reported significant association (p<0.01) between c.746A>G polymorphisms of BMPR1B gene and litter size traits in Garole × Malpura crossbreds, Small Tailed Han and Indonesian fat-tailed sheep. Chu et al. (2011) reported that the ewes with genotypes BB and B+ had 1.51 (p<0.001) and 1.02 (p<0.001) lambs more compared to ++ genotype. Mishra et al. (2009) reported that 65.6% higher litter size recorded in ewes with BB genotype compared to non-carriers (++) in Garole × Malpura crossbred sheep. All of the above stated results are consistent with the present findings along with the effects of mutant genotypes in various degrees. On the other hand, Chu et al. (2007b) reported that the g.218A>G polymorphism of INHβA gene significantly associated with litter size in Small Tailed Han ewes where the genotype GG had 0.53 (p<0.05) or 0.63 (p<0.05) more lambs than those with genotype AG and AA, respectively and is consistent with the present findings.
Based on geographic categorization, the mean litter size of JRB sheep population (2.83±0.40) was significantly higher (p<0.01) than those of SDR (1.71±0.09) and COR (1.09±0.04) for BMPR1B mutation (Table 3). The prolificacy attributed BB genotype frequency was the highest (80.95 %) in JRB population while it was absent in the COR population. The mean litter sizes were found to be 3.09±0.11, 2.67±0.33 and 1.00±0.00, respectively in the flocks of JRB, SDR and COR for INHβA genotypes (Table 3) and was differed significantly among those three sheep populations (p<0.01). The highest GG genotype frequency (45.46%) of INHβA mutation was also observed in JRB population. This result depicted that both polymorphisms had uneven distribution in indigenous sheep population and were predominantly available in the JRB flocks. The JRB flock had better reproductive attributes such as early sexual maturity and high prolificacy compared to other indigenous sheep population of Bangladesh (Hasan and Talukder, 2011; Islam et al., 2018).
Altogether, two polymorphisms were detected in BMPR1B (c.746A>G) and INHβA (c218A>G) genes in indigenous sheep of Bangladesh where both of them had significant association with litter size trait. Moreover, these mutations were population specific and were abundant in JRB population. However, no polymorphism was detected in GDF9 and BMP15 genes’ fragments. To best of our knowledge, this is the first report on polymorphisms detection and association analysis in Bangladeshi indigenous sheep. The discovery of mutations will allow to adopt breeding strategies for improving further the prolificacy of Indigenous sheep and this could also be used for marker-assisted selection programs for the genetic improvement of reproductive performances in the said population.
Table 1. Allelic and genotypic frequencies of BMPR1B and INHβA gene loci in indigenous sheep of Bangladesh.
|
Gene
|
No. of ewes
|
Allelic frequencya
|
Genotypic frequencyb, c
|
|
BMPR1B
|
121
|
+
|
B
|
++
|
+B
|
BB
|
|
0.44
|
0.56
|
0.31 (37)
|
0.26 (32)
|
0.43 (52)
|
|
INHβA
|
25
|
A
|
G
|
AA
|
AG
|
GG
|
|
0.56
|
0.44
|
0.36 (09)
|
0.40 (10)
|
0.24 (6)
|
aWild type allele (+) and mutant allele (B) for BMPR1B gene polymorphism; A and G denote wild and mutant allele, respectively for INHβA gene.
bHomozygous, heterozygous and non-carrier genotypes are BB, +B and ++, and GG, GA and AA, for BMPR1B and INHβA polymorphisms, respectively.
cNumbers in the parentheses indicate the individuals belong to the respective genotype.
Table 2. Mean±SE for litter size of different BMPR1B and INHβA genotypes in indigenous sheep of Bangladesh.
|
Gene
|
SNP
|
Genotype
|
No. of ewes
|
Litter size
|
|
BMPR1B
|
c.746A>G
(Q249R)
|
FecB++
|
37
|
1.19c±0.07
|
|
FecBB+
|
32
|
1.44b±0.09
|
|
FecBBB
|
52
|
2.13a±0.09
|
|
INHβA
|
c.218A>G
|
AA
|
9
|
2.11b±0.39
|
|
AG
|
10
|
1.70b±0.30
|
|
GG
|
6
|
2.83a±0.40
|
Means with different superscripts within a column differ significantly for BMPR1B (P<0.05) and INHβA (P<0.01) genotypes. For BMPR1B polymorphism; + and B represent wild and mutant alleles while A and G denote wild and mutant alleles for INHBA gene mutation.
Table 3. Location wise average letter size and genotype frequency for BMPR1B and INHβA gene polymorphisms in indigenous sheep of Bangladesh.
|
Gene and SNP
|
Location
|
Litter size
|
BMPR1B genotype frequency (%)
|
|
FecB++
|
FecBB+
|
FecBBB
|
|
BMPR1B (c.746A>G)
|
COR
|
1.09c±0.04 (45)
|
71.11 (32)
|
28.89 (13)
|
0.00
|
|
JRB
|
2.24a±0.10 (42)
|
4.76 (02)
|
14.29 (06)
|
80.95 (34)
|
|
SDR
|
1.71b±0.09 (34)
|
8.82 (03)
|
38.24 (13)
|
52.94 (18)
|
|
INHβA
(c.218A>G)
|
COR
|
1.00b±0.00 (11)
|
36.36 (04)
|
54.55 (06)
|
9.09 (01)
|
|
JRB
|
3.09a±0.11 (11)
|
36.36 (04)
|
18.18 (02)
|
45.46 (05)
|
|
SDR
|
2.67a±0.33 (03)
|
33.33 (01)
|
66.67 (02)
|
0.00 (0)
|
Means with different superscripts within a column differ significantly for BMPR1B (P<0.05) and for INHβA (P<0.01) genotypes. COR = Coastal regions, SDR =Sundarban delta regions and JRB = Jamuna River basin areas.
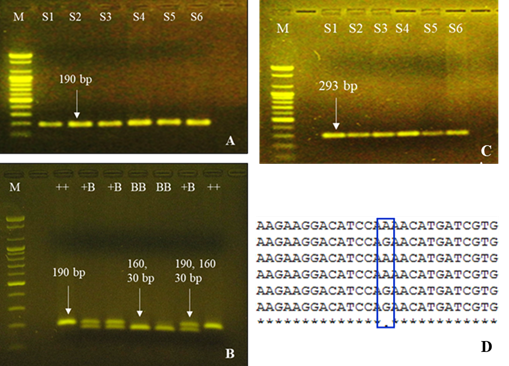
Figure 1. PCR amplification (A, C) and PCR-RFLP genotyping (B) of BMPR1B and multiple sequence alignment (D) of INHβA gene fragment in indigenous sheep of Bangladesh. M = 100 bp size marker, S1-S6 denote the samples’ ID. The ++, +B and BB represent wild, heterozygous and mutant individuals. Blue box in sequence alignment highlighted c.218A>G polymorphism.
Supplementary Materials
Table S1. Gene and primer sequence information for the selected fragments of this study.
|
Gene
|
Primer name
|
Sequence information (bp)
|
PCR fragment size
|
Annealing temp.(°C)
|
Amplicons
|
|
BMPR1B
|
FecB-F1
|
5'-CCAGAGGACAATAGCAAAGCAAA-3'
|
190 bp
|
59°C
|
Exon 6
|
|
FecB-R1
|
5'-CAAGATGTTTTCATGCCTCATCAACAGGTC-3'
|
|
INHβA
|
INHβA-F
|
5'-GCTACCACGCCAACTACTGT-3'
|
293 bp
|
60°C
|
Exon 2
|
|
INHβA-R
|
5'-TCTCTGGACCATCTCGCTC-3'
|
|
BMP15
|
FecXG -F
|
5'-CACTGTCTTCTTGTTACTGTATTTCAATGAGAC-3'
|
141 bp
|
62°C
|
Exon 1
|
|
FecXG -R
|
5'-GATGCAATACTGCCTGCTTG-3'
|
|
FecXHIL -F
|
5'-GGCAGTATTGCATCGGAAGT-3'
|
216 bp
|
54°C
|
Exon 2
|
|
FecXHIL -R
|
5'-GATGGCATGATTGGGAGAAT-3'
|
|
GDF9
|
FecG1-F
|
5'-GAAGACTGGTATGGGGAAATG-3
|
462 bp
|
62°C
|
Exon 1
|
|
FecG1-R
|
5'-CCAATCTGCTCCTACACACCT-3'
|
|
FecG8-F
|
5'-CCATGACTTTAGACTTAGC-3'
|
324 bp
|
55°C
|
Exon 2
|
|
FecG8-R
|
5'-TGGTTTTACTTGACAGGAG-3'
|
Acknowledgements: The study was supported by Bangladesh Academy of Sciences, Bangladesh (Project No.: BAS-USDA PALS LS-19). Special thanks to sheep farmers for providing valuable data on their flock performances and blood samples.
Conflict of interest: The authors disclose no conflicts of interest
- Chu, M. X., Z. H. Liu, C. L. Jiao, Y. Q. He, L. Fang, S. C. Ye, G. H. Chen and J. Y. Wang (2007a). Mutations in BMPRIB and BMP15 genes are associated with litter size in Small Tailed Han sheep (Ovis aries). J. Anim. Sci.85(3): 598-603.
- Chu, M. X., C. T. Xiao, Y. Fu, L. Fang and S. C. Ye (2007b). PCR-SSCP Polymorphism of Inhibin: A Gene in Some Sheep Breeds. Asian-Australas. J. Anim. Sci.20(7): 1023-1029.
- Chu, M. X., X. H. Zhao, Y. J. Zhang, M. Jin, J. Y. Wang, R. Di, G. L. Cao, T. Feng, L. Fang, Y. H. Ma and K. Li (2011). Polymorphisms of BMPR-IB gene and their relationship with litter size in goats. Mol. Biol. Rep. 37: 4033-4039.
- Chu, M.X., R. Di, S.C. Ye et al. (2009). Establishment of molecular detection methods for high prolificacy major gene FecB in sheep and its application. J. Agric. Biotechnol. 17:52-58.
- Davis, G.H., G.W. Montgomery, A. J. Allison, R.W. Kelly and A.R. Bray (1982). Segregation of a major gene influencing fecundity in progeny of Booroola sheep. New Zealand J. Agric. Res. 25:525-529.
- Davis, G. H., S. M. Galloway, I. K. Ross, S. M. Gregan, J. Ward, B. V. Nimbkar, P. M. Ghalsasi, C. Nimbkar, G. D. Gray, I. Inounu and B. Tiesnamurti (2002). DNA tests in prolific sheep from eight countries provide new evidence on origin of the Booroola (FecB) mutation. Biol. Reprod. 66(6): 1869-1874.
- Fabre, S., A. Pierre, P. Mulsant, L. Bodin, E. Di Pasquale, L. Persani and D. Monniaux (2006). Regulation of ovulation rate in mammals: Contribution of sheep genetic models. Reprod. Biol. Endocrin. 4: 20.
- Falconer, D. S. and T. F. C. Mackay (1996). Introduction to Quantitative Genetics (4th edn). Prentice Hall, London, UK. pp.1-21.
- Gillespie, J. H. (2004). Population genetics: a concise guide (2nd edn). The Johns Hopkins University Press. Baltimore and London. pp.10-18.
- Hall, T.A. (1999). BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41: 95-98.
- Hanrahan, J. P., S. M. Gregan, P. Mulsant, M. Mullen, G. H. Davis, R. Powell and S. M. Galloway (2004). Mutations in the genes for oocyte-derived growth factors GDF9 and BMP15 are associated with both increased ovulation rate and sterility in Cambridge and Belclare sheep (Ovis aries). Biol. Reprod. 70: 900-909.
- Hassan, M. R. and M. A. I. Talukder (2011). Comparative performance of different regional native sheep in Bangladesh. The Bangladesh Veteri. 28(2): 85-94.
- Hossain, M. I., M. K. I. Khan, M. M. Momin and A. Das (2020). Effects of protein supplements on fertility and assessment of the fertility genes (GDF9 and BMP15) in indigenous sheep of Bangladesh. J. Appl. Anim. Res. 48(1): 484-491.
- Islam, M. S., A. K. F. H. Bhuiyan, M. Ersaduzzaman, M. M. Hossain, S. H. Lee and M. S. A. Bhuiyan (2018). Morphometric features, production and reproduction potentials of indigenous sheep genetic resources of Bangladesh. J. Anim. Breed. Genom.2(2): 1-8.
- Kolosov, Y. A., L. V. Genmantseva, N. V. Shirockova and A. Klimenko (2015). Polymorphism of the GDF9 Gene in Russian Sheep Breeds. J. Cytol. Histol.6(4): 01-04.
- Liu, S. F., H. B. Li, X. M. Song, A. H. Wang, C. H. Wei and L. X. Du (2006). Candidate genes with ovulation by differential display PCR in Small Tail Han sheep. Asian-Australas. J. Anim. Sci.19(9): 1229-1233.
- Mahdavi, M., S. Nanekarani and S. D. Hosseini (2014). Mutation in BMPRIB gene is associated with litter size in Iranian Kalehkoohi sheep. Anim. Reprod. Sci.147(3-4): 93-98.
- Maskur, M., R. Tapaul and L. Kasip (2016). Genetic polymorphism of bone morphogenetic protein receptor 1B (BMPR1B) gene and its association with litter size in Indonesian fat-tailed sheep. African J. Biotechnol.15(25): 1315-1319.
- Mishra, A. K., A. L. Arora, S. Kumar and L. L. L. Prince (2009). Studies on effect of Booroola (FecB) genotype on lifetime ewes’ productivity efficiency, litter size and number of weaned lambs in Garole×Malpura sheep. Anim. Reprod. Sci.113(1-4): 293-298.
- Moradband, F., G. Rahimi and M. Gholizadeh (2011). Association of polymorphisms in fecundity genes of GDF9, BMP15 and BMPR1B with litter size in Iranian Baluchi sheep. Asian-Australas. J. Anim. Sci.24(9): 1179-1183.
- Roy, J., S. Polley, S. De, A. Mukherjee, S. Batabyal, S. Pan, B. Brahma, T. K. Datta and S. L. Goswami (2011). Polymorphism of fecundity genes (FecB, FecX, and FecG) in the Indian Bonpala sheep. Anim. Biotechnol. 22(3): 151-162.
- Sharma, R. C., A. L. Arora, H. K. Narula and R. N. Singh (1999). Characteristics of Garole sheep in India. Anim. Genet. Resour. Infor. 26: 57-64.
- Souza, C. J. H., C. MacDougall, B. K. Campbell, A. S. McNeilly and D. T. Baird (2001). The Booroola (FecB) phenotype is associated with a mutation in the bone morphogenetic receptor type 1B (BMPR1B) gene. J. Endocrinol. 169(2): R1-R6.
- Sudhakar, A., R. Rajendran and P. S. Rahumathulla (2013). Detection of Booroola (FecB) mutation in Indian sheep-Nilagiri. Small Rumin. Res. 113: 55-57.
- Zhao, Z., M. Zhu, S. Cao, M. Zhai, H. Yang and Y. Nan (2019). Effect of Inhibin-βA subunit gene on reproductive performance of Kazakh Sheep in non-breeding season. Kafkas Universitesi Veterinary Fakultesi Dergisi. 25: 611-618.
- Zhu Z., D. Yuan, D. Luo, X. Lu and S. Huang (2015). Enrichment of Minor Alleles of Common SNPs and Improved Risk Prediction for Parkinson's Disease. PLoS One 10(7): e0133421.
- Zuo B., Qian H., Wang Z., Wang X., Nisa N., Bayier A., Ying S., Hu X., Gong C., Guo Z. and Wang F. (2013). A study on BMPRIB genes of Bayanbulak sheep. Asian-Australas. J. Anim. Sci. 26(1): 36-42.
|





