SCREENING OF LACTIC ACID BACTERIA FROM SELECTED SAUDI HONEY PRODUCED BY APIS MELLIFERA JEMENTICA
M. B. Hussain1⃰, Y. M. K. M. Aly1, A. A. M. Jiman-Fatani2, Zia Ullah1, I. A. Qureshi3, M. A Bakarman4, M. Yasir5, and Saleh M. Al-Maaqar6
1Department of Microbiology, Faculty of Medicine, Rabigh Branch, King Abdulaziz University (KAU), P.O. Box 80203, Jeddah 21589, Saudi Arabia (SA).
2Department of Medical Microbiology and Parasitology, Faculty of Medicine, King Abdulaziz University (KAU), P.O. Box 80203, Jeddah 21589, Saudi Arabia (SA).
3Department of Pathology, Faculty of Medicine, Rabigh Branch, King Abdulaziz University (KAU), P.O. Box 80203, Jeddah 21589, Saudi Arabia (SA).
4Department of Family & Community Medicine, Faculty of Medicine, Rabigh Branch, King Abdulaziz University (KAU), P.O. Box 80203, Jeddah 21589, Saudi Arabia (SA).
5Special Infectious Agents Unit, King Fahd Medical Research Center, King Abdulaziz University, Jeddah 21589, Saudi Arabia.
6Department of Biological Sciences, Faculty of Science, King Abdulaziz University, P.O. Box 80203, Jeddah, 21589, Saudi Arabia and Department of Biology, Faculty of Education, Al-Baydha University, Al-Baydha, Yemen.
*Corresponding Author’s email: mbhussain1@kau.edu.sa
ABSTRACT
Honey is receiving worldwide attention as an excellent source of probiotics because it contains lactic acid bacteria (LAB) derived from bees as well as multiple essential nutrients, including prebiotics, required for maintaining and enhancing the growth of probiotics. There is limited information on the presence of LAB in honey, especially in Saudi Arabia. Consequently, the purpose of the current study was to isolate and identify LAB from selected honey samples produced by Apis mellifera jementica obtained from various areas around Saudi Arabia using classical cultivation methods and 16S rDNA sequence analysis. The current study examined seventeen commercially sold honey samples, mainly from Saudi Arabia's Al-Baha, Aseer, and Jeddah (Hada Al-Sham) regions.LAB were isolated from honey samples and grown on De Man-Rogosa-Sharpe (MRS) agar. The phenotypic characterization of honey isolates was ascertained using Gram staining, microscopy, and the catalase test. Using 16S ribosomal RNA gene sequencing, isolates were genotypically described. Lactobacillusrhamnosus and Enterococcusfaecium were identified from Sidr, Talah, and Somra honey produced by Apis mellifera jementica. The P02, P14, and Con-6 strains were found to share > 99% of their DNA with Lactobacillus rhamnosus by 16S rRNA sequencing investigations, while the P01, P03, and P04 strains shared > 99% of their DNA with Enterococcus faecium. Additionally, the phylogenetic tree reveals that the honey strains isolated from Sidr (P02) and Talah (P14) were closely linked to Lactobacillus rhamnosus, whereas P01 (Sidr), P03 (Somra), and P04 (Somra) were most closely connected to Enterococcus faecium.According to the research, honey produced by Apis mellifera jementica is a beneficial source of probiotics and lactic acid bacteria.
Keywords: probiotics, lactic acid bacteria, honey, Sidr, Lactobacillus rhamnosus, prebiotics, Enterococcus faecium.
https://doi.org/10.36899/JAPS.2023.2.0616
Published first online December 18, 2022
INTRODUCTION
Food containing probiotics is in high demand around the world because of the continuous and vigorous generation of research evidence demonstrating their potential health advantages to its users. Probiotics are available in a variety of products, including dairy, functional foods, and food supplements with health claims (Peng et al., 2020). Foods as well as supplements appear to have been effective carriers for probiotics, promoting community health in general. However, probiotics obtained from supplements may be sensitive to bile salts and acids and cause antibiotic resistance (Wang et al., 2020). As a result, natural foods may be preferred over probiotic supplements due to buffering effects for probiotics while passing through the gastrointestinal tract, maintaining probiotic bacterial activity and efficacy through the availability of necessary nutrients, promoting probiotic growth through the synergistic effects of dietary components, and customer preferences for probiotic foods over tablets, capsules, and other medicinal forms(Homayoni Rad et al., 2016).
Probiotics are capable of surviving in simulated gastrointestinal settings, generate antibacterial activity against pathogenic microbes by producing a myriad of active antimicrobial compounds, and modulate gut microbiota (Stasiak-Różańska et al., 2021). We recently demonstrated that in a simulated gastric environment, pepsin and acidic pH had no negative effects on honey's antimicrobial activity against multidrug resistant (MDR) Shigella sonnei; rather, acidic pH increased honey's antibacterial capability (Al-Masaudi et al., 2020). Therefore, honey with probiotic bacteria may have far greater medicinal potential than honey or probiotic bacteria used alone, particularly against gastrointestinal tract and wound infections caused by MDR and pan-drug resistant (PDR) bacteria. Unfortunately, there is currently no established treatment regimen for patients with PDR bacterial infections (Karakonstantis et al., 2020). As a result, innovative therapeutic approaches for eradicating PDR bacteria are critical. Probiotics containing honey provide an effective, inexpensive, and safe solution in such difficult clinical settings because bacterial resistance to honey is extremely uncommon and has never been documented despite its widespread use as a topical therapy for recalcitrant wound infections, burn infections, diabetic foot wounds, and others (Bouzo et al., 2020).
Lactic acid bacteria (LAB) and Bifidobacterium, both from the Lactobacillus genus, are the most frequently utilized probiotic bacteria (Sharma et al., 2021). Other LAB that are classified as probiotics include Enterococcus, Lactococcus, and Streptococcus (Wang et al., 2021). Recent studies have shown a number of therapeutic impacts of probiotics on the human body on multiple levels (Sharifi-Rad et al., 2020). Some of these effects include prevention and treatment of gastrointestinal diseases, immune system activation, antibacterial activity, lowering serum cholesterol levels, prevention of cancer, antihypertensive effects, treatment of diarrhea caused by irritable bowel syndrome, and reduction of the clinical manifestations of lactose intolerance (Roobab et al., 2020). Recently, it has been revealed thatpostbiotics, a collection of compounds produced by probiotic microorganisms, have a protective role against diseases such as cardiovascular ailments, diabetes, and brain disorders (Diez-Gutiérrez et al., 2020).
Probiotics come in a variety of forms, each with their own set of health benefits. Comprehensive research on uncharted ecological niches, particularly indigenously fermented foods and others, could lead to the discovery of effective probiotics (Mishra et al., 2021). Dairy products have traditionally been employed as probiotic carriers, but these products are associated with some side effects, such as intolerance to lactose and allergy to milk proteins(Chaturvedi et al., 2021). As a result, non-dairy foods such as fruits, vegetables, cereals, and others are increasingly being investigated as probiotic carriers(Ansari and Pourjafar, 2021).
Honey is a popular food consumed all over the world and has significant nutritional and medicinal value (Hussain, 2018). Its matrix provides an ideal habitat for probiotic bacteria, as it contains prebiotics (oligosaccharides), organic acids, phenolic compounds, zinc and reducing sugars (Ebrahimi et al., 2021). These substances promote the growth of probiotics through oxidative stress protection, prebiotic activity, and a decrease in redox potential (Castro et al., 2021). In Saudi Arabia, honey is widely consumed as an alternative remedy for prophylaxis and treatment of several illnesses (Al-Ghamdi and Nuru, 2013). Apis mellifera jementica, a Saudi native, has been used in the majority of apiculture because it withstands local stressful conditions better than imported bees, especially in the summer when air temperatures soar (Alqarni et al., 2011a). Recently,Alatawy et al. (2020) identifiedLactobacillus and Bifidobacterium in the gut of Saudi Apis mellifera jementica (Alatawy et al., 2021). A lot of research carried out in different parts of the world also found LAB in bee stomachs (Olofsson et al., 2014; Aween et al., 2012). However, to our knowledge, there is limited information on the presence of LAB in honey, especially in Saudi Arabia. Consequently, the purpose of the current study was to isolate and identify LAB from selected honey samples produced by Apis mellifera jementica obtained from various areas around Saudi Arabia using classical cultivation methods and 16S rDNA sequence analysis.
MATERIALS AND METHODS
The current study examined seventeen commercially sold honey samples, mainly from Saudi Arabia's Al-Baha, Aseer, and Jeddah (Hada Al-Sham) regions (Table 1, Fig 1). The hilly regions of Al-Baha and Aseer in Saudi Arabia's southwest are the best places for beekeeping (Alqarni et al., 2011b). Hada Al-Sham is a small town in western Saudi Arabia that has one of the agricultural research stations of King Abdulaziz University and is situated sixty kilometers northeast of Jeddah. It has a large amount of underground fresh water as well as honey bee flora (Hakala et al., 2019). Acacia spp. and Ziziphus spina-christi are the two most common bee flora in Saudi Arabia, both of which may be found growing wild in most parts of the country (Al-Masaudi et al., 2020). According to the methods used in earlier studies, the botanical sources of honey samples were identified based on flowering plants, geographical location, season, aroma, and color of each honey (Davis, 2005). More definite identification of honeys by melissopalynology is expensive and is rarely done. Before analysis, the honey samples were kept at room temperature.
Inclusion criteria: honey produced by Apis mellifera jementica
Exclusion criteria: heated honey
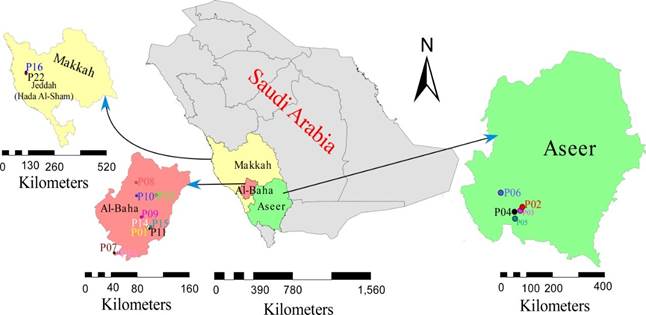
Fig. 1. Collection sites in Makkah, Al-Baha (left), and Aseer (right) regions of Saudi Arabia are shown on a map. Areas marked by circles are the sites for collection of honey.
Table 1. Profile of honey samples screened for lactic acid bacteria.
|
Code
|
Honey
|
Botanical name
|
Location
|
Season
|
|
P01
|
Sidr
|
Ziziphus spina-christi
|
Al-Baha, Baljurashi
|
October-2017
|
|
P02
|
Sidr
|
Ziziphus spina-christi
|
Aseer, Alsouda
|
October-2018
|
|
P03
|
Somra
|
Acacia tortilis
|
Aseer, Alsouda
|
April-2018
|
|
P04
|
Talah
|
Acacia spp.
|
Aseer, Rigal Almaa
|
April-2017
|
|
P05
|
Talah (comb)
|
Acacia spp.
|
Aseer, Rigal Almaa
|
April-2018
|
|
P06
|
Somra (comb)
|
Acacia tortilis
|
Aseer, Mahayil
|
April-2018
|
|
P07
|
Somra (comb)
|
Acacia tortilis
|
Al-Baha, Tahama
|
April-2018
|
|
P08
|
Sidr
|
Ziziphus spp.
|
Al-Baha, Bidah and Mahshuqa
|
October-2017
|
|
P09
|
Qatada
|
Acacia hamulosa
|
Al-Baha- Aqba
|
April-2018
|
|
P10
|
Dhuhyana
|
Acacia asak
|
Al-Baha, Bidah
|
April-2018
|
|
P11
|
Majra
|
Hypoestes forskaolii
|
Al-Baha, Baljurashi
|
April-2018
|
|
P12
|
Somra
|
Acacia tortilis
|
Al-Baha Al-Aqiq
|
April-2018
|
|
P13
|
Orange blossom
|
Citrus spp.
|
Al-Baha-Nawana
|
March-2018
|
|
P14
|
Talah
|
Acacia spp.
|
Al-Baha, Baljurashi
|
April-2018
|
|
P15
|
Sidr
|
Ziziphus spp.
|
Al-Baha, Baljurashi
|
October -2018
|
|
P16
|
Sidr
|
Ziziphus spp.
|
Jeddah, Hada Al-Sham
|
October-2017
|
|
P22
|
Sidr
|
Ziziphus spp.
|
Jeddah, Hada Al-Sham
|
October-2018
|
Isolation and identification of LAB: De Man-Rogosa-Sharpe (MRS) agar (Merck, Germany) was used to isolate LAB bacteria from honey samples following Aween et al. (2012). In a stomacher (Seaward model, Germany), 10 g of honey from each sample was poured into 90 ml of peptone water (0.1 percent w/v) and manually stirred. One milliliter of diluted honey was added to 10 ml ofMRS broth (Merck, Darmstadt, Germany) and incubated in an anaerobic jar at 30°C for 48 hrs. Then, 0.1 ml of incubated MRS broth was placed on MRS agar containing 0.9% CaCO3. In an anaerobic environment, the inoculated agar plates were incubated at 37°C for 48-72 hrs. The incubation period was prolonged in all cases until visible colonies formed.Protein Balance, a commercially available probiotic food supplement containingLactobacillus rhamnosus, was used as a control.
Phenotypic characterization:Gram staining (bioMérieux, Lyon, France), microscope examination, and the catalase test (bioMérieux, Lyon, France) were used to determine the phenotypic characterization of LAB isolates. Gram-positive and catalase-negative growth was preserved in MRS broth with 16% glycerol and stored at -20 ºC for further evaluation.
Genotypic Characterization
DNA extraction: Each overnight isolate culture (2 ml) was centrifuged at 14,000 g for 12 minutes at 4°C, and the resultant pellet was used to extract DNA using a QIAamp DNA Mini Kit (Qiagen, Germany). Spectrophotometer measurements at 260 and 280 nm (A260/A280) were used to evaluate the purity of DNA.
PCR reaction and program: Targeting the genus level of LAB, a partial 16S rDNA gene fragment was amplified with universal primers 27F (50-AGAGTTTGATCCTGGCTCAG-30) and 1492R (50-GGTTACCTTGTTACGACTT-30). These primers are designed against the conserved region and widely used universal primer for species-level identification (Lane, 1991). PCR was carried out in 20 ml reaction volumes, comprising 1.5 mM MgCl2, 0.25 mM forward primer, 1X Taq Master Mix, 0.25 mM reverse primer, and 0.4 ng of genomic DNA. The following were the temperature cycling conditions for PCR: 95 °C for 3 min, followed by 40 cycles of denaturation at 95 °C for 30 sec, annealing at 55 °C for 55 sec, extension at 72 °C for 1 min, and a 10-min final incubation at 72 °C. Eppendorf Master cycler gradient PCR (Eppendorf, Germany) was used to increase primer-annealing temperatures throughout the tests, resulting in a high degree of primer specificity. On an agarose 0.8% gel, the PCR-amplified fragments were resolved by electrophoresis at 75 V for 1 hr 30 min. After the gel was stained with ethidium bromide, the bands were visible under UV light at 254 nm. A PCR purification kit was used to purify the rest of the sample (Qiagen).
Sequencing and identification of isolate: An amount of 50 μL of each purified PCR product was sent to Macrogen in Seoul for nucleotide sequencing, where they were sequenced on an Automated Sequencer ABI Prism 3730XL (Korea). The isolates were assigned to likely genera or species using 785F and 907R primers as defined by the molecular operational taxonomic unit (MOTU). The partially sequenced 16S rDNA, which was obtained, was aligned and compared to homologous sequences in the GenBank database of the National Centre for Biotechnology Information (NCBI) for similarity using the Advanced Basic Local Alignment Search Tool (BLAST) search option on the NCBI homepage (http://www.ncbi.nlm.nih.gov/) (Altschul et al., 1997). Strains with an identification percentage of at least 97 percent compared to other strains in databases were assigned to the same species, as previously described (Hajigholizadeh et al., 2020).
Phylogenetic analysis:The Tamura-Nei model and the maximum likelihood (ML) approach were used to infer the evolutionary history (Tamura and Nei, 1993). Figures 2-6 depict the tree with the greatest log likelihood (-2784.77). The fraction of trees where the related taxa are joined together is depicted alongside the branches. The initial trees were built automatically, employing the BioNJ algorithms and neighbor-join on a matrix of pairwise distances calculated with the Tamura-Nei model. Next, the topology with the highest log probability value is selected. Per site, the number of substitutes is used to calculate branch lengths, and the tree is drawn to scale. A total of 20 nucleotide sequences were involved in this study. Using the complete deletion option, the gaps and missing data were removed from every spot.In the final dataset, there were a total of 928 sites. MEGA X was used to undertake phylogenetic analysis (Kumar et al., 2018).
Data analysis and species identification: The BLAST sequence tool was usedon the NCBI homepage (http://www.ncbi.nlm.nih.gov/) to analyze and determine the nucleotide sequences of the 16S rRNA gene in all of the isolates. Using MEGA X, a phylogenetic tree was created by processing thealignments and assessing sequence similarities using the neighbor-joining method.
RESULTS
Phenotypic characterization of LAB from honey: Five honey samples out of seventeen exhibited bacterial growth on MRS agar after 48-72 h of incubation in an anaerobic environment. The colonies of isolates were characterized as small, round, white, and opaque. Gram-positive bacilli were found in singles, pairs, and chains on microscopic examination. They were catalase-negative and facultative anaerobic (Table 2). These characteristics were similar to LAB's.
Table 2. Phenotypic profile of lactic acid bacteria isolated from honey samples.
|
Honey sample with code
|
Microbial count (CFU/g)
|
Characteristics
|
Tentative identification
|
Designated species as per16S rRNA
|
|
Sidr (P01)
|
1.2 × 102 ± 0.3
|
Gram +, rods in pairs and clusters, non-motile, non-sporulated, catalase negative
|
Lactobacillus spp.
|
Enterococcus faecium
|
|
Sidr (P02)
|
1.6 × 103 ± 0.2
|
Gram +, long rods, non-motile, non-sporulated, catalase negative
|
Lactobacillus spp.
|
Lactobacillus rhamnosus
|
|
Somra (P03)
|
1.2 × 104 ± 0.2
|
Gram +, long rods, non-motile, non-sporulated, catalase negative
|
Lactobacillus spp.
|
Enterococcus faecium
|
|
Somra (P06)
|
70 ± 13
|
Gram +, rods, non-motile, non-sporulated, catalase negative
|
Lactobacillus spp.
|
Enterococcus faecium
|
|
Talah (P14)
|
65 ± 15
|
Gram +, long rods, non-motile, non-sporulated, catalase negative
|
Lactobacillus spp.
|
Lactobacillus rhamnosus
|
⃰ LAB; lactic acid bacteria, CFU; colony forming units
Genotypic characterization: The partial 16S rDNA sequences were obtained by Automated Sequencer ABI Prism 3730XL sequencing at Macrogen, Seoul (Korea) and subjected to a BLASTN search to find a close match (data attached as supplementary files). As a result, numerous hits were found that shared significant similarities with distinct LAB species. Table 3 shows that there were three isolates belonging to Lactobacillus and three to Enterococcus spp. 16S rRNA sequencing studies found that the P02, P14 and Con-6L strains isolated from Sidr, Talah and Protexin Balance had > 99% identity with Lactobacillus rhamnosus, while P01 (Sidr), P03 (Somra) and P04 (Somra) showed >99% similarity with Enterococcus faecium. The 16S rRNA gene sequences of honey isolates were deposited in GenBank and assigned the following accession numbers: OL539410, OL539411, OL539412, OL539413, OL539414, and OL539415 (Table 3).
Table 3. 16S sequence divergence among study isolates and reference isolates.
|
Isolate no
|
Designated species
|
Reference type strain
|
Similarity %
|
Accession numbers
|
Isolated from
|
|
P01-1L
|
Enterococcus faecium
|
Enterococcus faecium LMG 11423
|
99.42
|
OL539410
|
Sidr
|
|
P0-2L
|
Lactobacillus rhamnosus
|
Lactobacillus rhamnosus JCM 1136
|
99.71
|
OL539411
|
Sidr
|
|
P0-3L
|
Enterococcus faecium
|
Enterococcus faecium LMG 11423
|
99.57
|
OL539412
|
Somra
|
|
P0-4L
|
Enterococcus faecium
|
Enterococcus faecium LMG 11423
|
99.64
|
OL539413
|
Somra
|
|
P14-5L
|
Lactobacillus rhamnosus
|
Lactobacillus rhamnosus JCM 1136
|
99.63
|
OL539414
|
Talah
|
|
Con-6L
|
Lactobacillus rhamnosus
|
Lactobacillus rhamnosus JCM 1136
|
99.86
|
OL539415
|
Protexin⃰
|
⃰A commercially available probiotic product
Phylogenetic analysis:The phylogenetic tree shows that the honey strains isolated from P14-5L (Talah), P0-2L (Sidr) and P06-6L (Protexin Balance; control) were closely related to Lactobacillus rhamnosus, whereas P0-IL (Sidr), P0-3L (Somra) and P0-4L (Somra) were closest to Enterococcus lactis(Fig. 2-6).
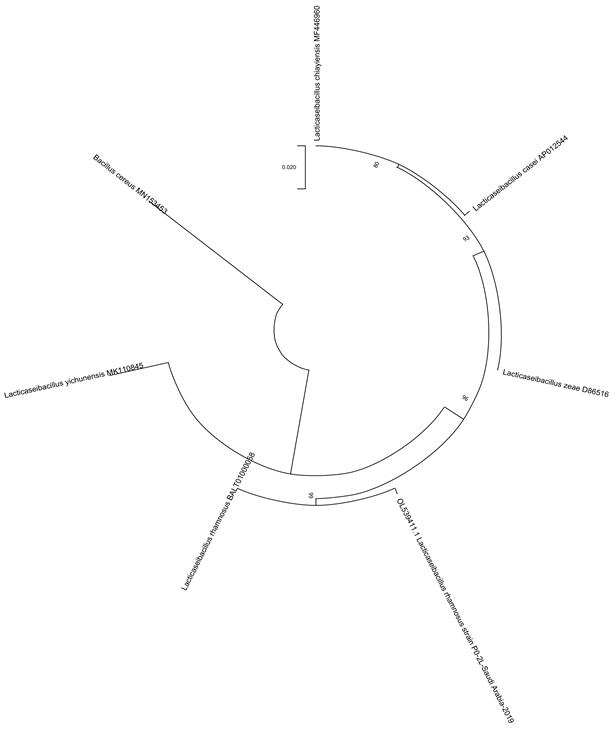
Fig 2. Phylogenetic tree of honey isolate Lactobacillus rhamnosus, isolated from Sidr honey (P0-2L).
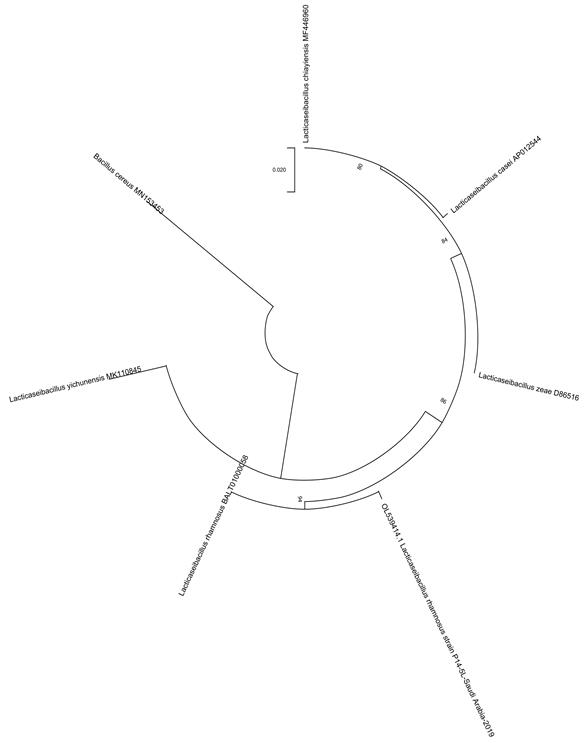
Fig 3. Phylogenetic tree of honey isolate Lactobacillus rhamnosus, isolated from Talah honey (P14-5L).
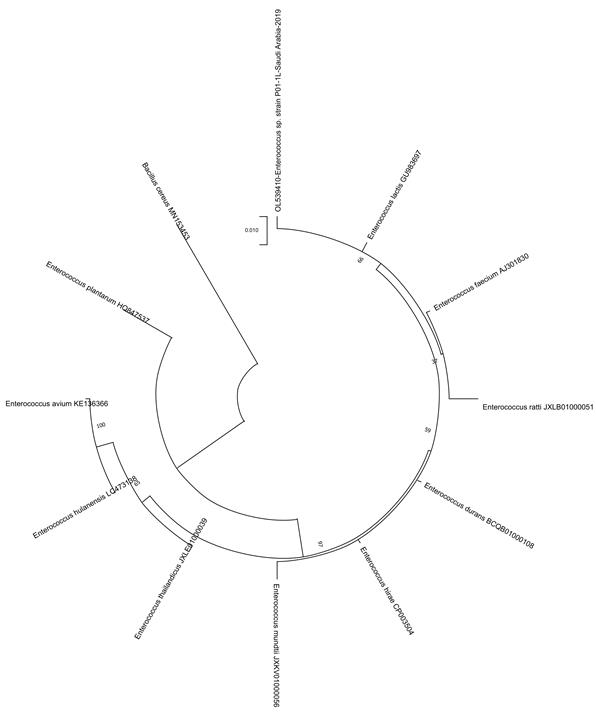
Fig 4. Phylogenetic tree of honey isolate Enterococcus species, isolated from Sidr honey (P01-1L).
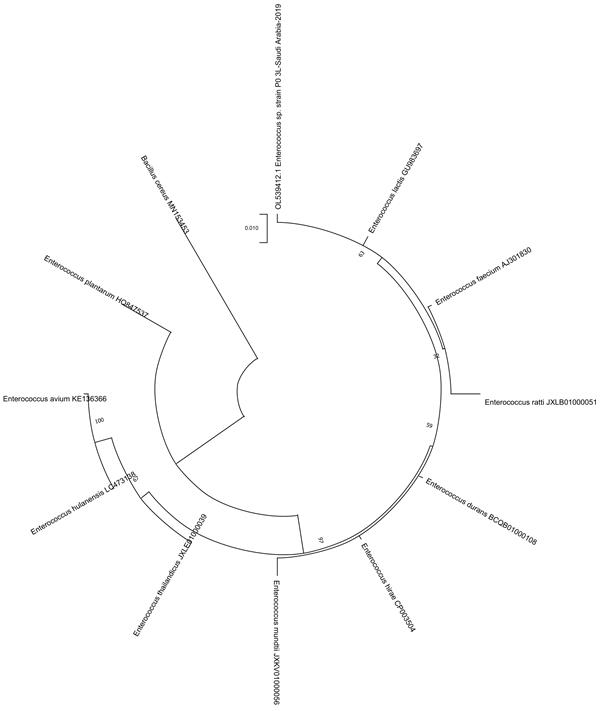
Fig 5. Phylogenetic tree of honey isolate Enterococcus species, isolated from Somra honey (P0-3L).
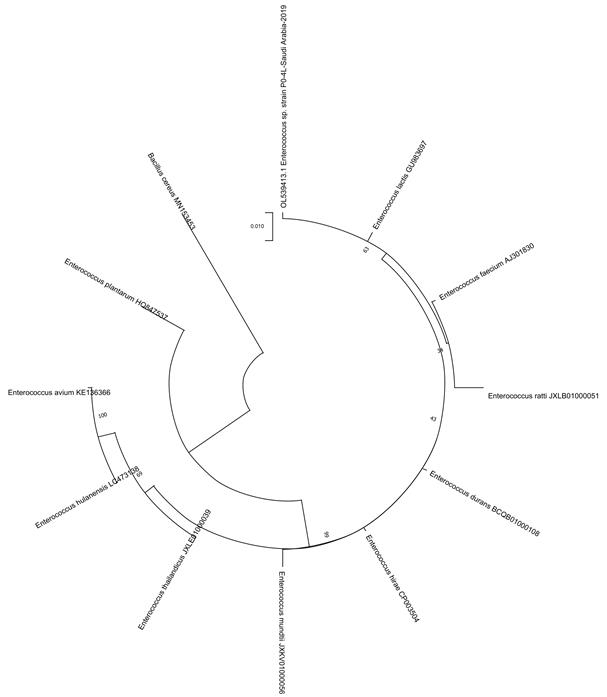
Fig 6. Phylogenetic tree of honey isolate Enterococcus species, isolated from Somra honey (P0-4L).
DISCUSSION
Generally, honey inhibits the growth of pathogenic bacteria due to its high osmolarity, reduced water activity, natural acidity, presence of H2O2 and phytochemical substances such as phenolic acids and flavonoids derived from honey bee flora (Hussain, 2018). In addition, honey also contains enzymes and antimicrobial peptides such as bee defensin-1 that originate from the bee gut (Kafantaris et al., 2021). These antibacterial factors are increasingly recognized as functioning in a synergistic fashion and targeting pathogenic bacteria on multiple sites (Brudzynski and Sjaarda, 2021). In addition, they have the capability to interfere with quorum sensing and disrupt bacterial biofilms, thus not letting bacteria generate resistance (Anju et al., 2021). Moreover, LAB present in some honey types also produce antibacterial compounds such as bacteriocins, benzoates, organic acids, diacetyl and H2O2, which can further complement the antibacterial potential of honey (Hernández-González et al., 2021). Therefore, honey that possesses LAB could serve as a valuable source of natural, broad-spectrum, and potent antibiotics, in addition to other clinical applications.
Several studies have examined the diverse microbiota of honey, revealing the presence of beneficial bacteria such as lactic acid bacteria (Lashani et al., 2020). In the current study, we described the bacterial diversity of LAB found in honey using traditional culture methods along with 16S rRNA sequencing and phylogenetic analysis. Lactobacillusrhamnosus and Enterococcus faecium were identified by 16S rRNA sequencing analysis of locally produced Sidr (02 samples), Somra (02 samples), and Talah (01 sample) honey (Table 3). The phylogenetic analysis also exhibited the closest resemblance of Lactobacillus rhamnosus and Enterococcus spp. with honey isolates (Fig 2-6). Interestingly, there was complete homology between the findings of the 16S rDNA similarity analysis and the phylogenetic analysis of honey isolates. Both identification analyses demonstrated that L. rhamnosus and Enterococcus spp. had the closest similarity to honey isolates (Table 3, Fig 2 to 6).
rhamnosus is one of the most widely studied probiotic strains and is available as a commercially marketed probiotic product (Stivala et al., 2021). The reason for its popularity as a probiotic strain, stems from its acid and bile resistance, excellent growth in the digestive system, modulation of the immune system and ability to attach to the intestinal epithelial layer (Tkhruni et al., 2020). Toll-like receptors are involved in the interactions between host immune cells and probiotic bacteria (Petrova et al., 2021). The positive health benefits of L. rhamnosus have been extensively studied in clinical trials and in vitro experiments (Petrova et al., 2021). Several studies have shown that L. rhamnosus has beneficial effects on diarrhea, infections, immune disorders, diabetes, systemic inflammation, obesity, cancer, irritable bowel disease, mental disorders, and other conditions (Manasian et al., 2020). Olofsson et al. (2008) identified bacteria from fresh honey samples and the stomachs of honeybees. They discovered a unique LAB flora that included six Lactobacillus and four Bifidobacterium phylotypes (Olofsson et al., 2008).
Recently, Lashani et al. (2020) isolated L. rhamnosus in one of the honey samples and showed that L. rhamnosus isolates inhibited foodborne pathogenic bacteria, including Salmonella enteritidis, Shiglla flexneri, Staphylococcus aureus, enteropathogenic Escherichia coli and Bacillus cereus (Lashani et al., 2020). L. rhamnosus was recently found to diminish conglycinin-allergy-induced apoptotic cells in BALB/c mouse intestinal contents by modulating the Bacteroides and bile secretion pathways. The outcomes of this study offer a foundation for the application of dietary intervention in the treatment of allergy illnesses as a novel approach for lowering cell apoptosis (Chen et al., 2021). Previous research has shown that the L. rhamnosus GG strain has antiproliferative properties against gastrointestinal cancer (Śliżewska et al., 2021).
Endo and Salminen (2013) recovered 66 isolates of Fructobacillus fructosus and L. kunkeei from honey, bees, flowers, and larvae. A study analyzed nine Lactobacillus strains isolated from honey in various locations and found them to have probiotic characteristics (Endo et al., 2012). In Japan in 2015, L. kunkeei have been recovered from the gastrointestinal tract of bees (Asama et al., 2015). In 2012, six L. acidophilus strains were identified from commercially available honey in Malaysia (Aween et al., 2012).
faecium was identified from Sidr (P01) and Somra (P03 & P04) honey (Table 3). Enterococci are used in animal feed as probiotics to promote growth, prevent diarrhea, and stimulate the innate as well as the adaptive immune system (Peng et al., 2019). Furthermore, enterococci have the capability to produce a number of antibacterial substances, such as bacteriocins, lactic acid and H2O2, which could explain their ability to fight pathogenic bacteria such as Listeria monocytogenes and Staphylococcus aureus (Dinçer and Kıvanç, 2021). Many studies have shown that enterococcal strains meet probiotic criteria for acid and bile tolerance, antibacterial substance production, and adhesion ability on the host gastrointestinal mucosal surface (Anania et al., 2021). Moreover, numerous Enterococcus strains have been commercially employed in probiotic formulations, such as E. faecium SF-68, E. faecalis Symbioflor 1, and E. faecium M74 (Dioso et al., 2020).
Out of seventeen honey samples, twelve did not harbour LAB. This could be due to the low water content of the honey samples tested, as LAB cannot thrive at low water concentrations (Kňazovická et al., 2021). Honey is traditionally collected when the hives comprise a combination of almost ripe and mature honey with a water concentration of 22 to 30% (Olofsson et al., 2016b). In many regions of the world, this technique of gathering honey is still in use, and such honey contains a higher amount of viable LAB (108 cfu per gram of honey). According to a recent study, Apis laboriosa honey from Nepal and Apis mellifera honey from Africa had a high concentration of LAB (Vásquez et al., 2012b). Both honeys had more than 20% water content. LAB cannot survive in honey with a water content of less than 20%, so the honey samples with no viable LAB detected in this study could have less than 20% water content (Balzan et al., 2020). The water percentage of honey in Europe and many other countries is not permitted to exceed 20% (Thrasyvoulou et al., 2018). As a result, LAB are not viable in honey with a water content less than 20%. Therefore, to obtain the full benefit of honey along with its resident LAB symbionts, it is important for the health community to revise the criteria regarding the water content of honey, setting it an optimal level where LAB symbionts remain viable for an extended period of time.
The second possibility for negative LAB in tested honey could be heating the honey at a high temperature, which can also kill the LAB along with heat-sensitive antimicrobial components. To prevent honey crystallization, beekeepers in some parts of the world heat honey, which adversely affects the viability of LAB and the antimicrobial properties of honey (Scripcă and Amariei, 2021). Honey is currently being sterilized by gamma irradiation before its use as a topical therapeutic agent. This practice kills pathogenic microorganisms as well as LAB symbionts present in honey (Hermanns et al., 2020). Therefore, it is important to develop methods of selective sterilization of honey that target only pathogenic bacteria, sparing LAB symbionts. Heather honey, when compared to other honey types, was found to be more effective against equine wounds infected with drug-resistant pathogens. This could be because heather honey contains somewhat more water than other varieties of honey, allowing the LAB to be more active and viable (Olofsson et al., 2016a). To treat resistant wound infections, a well-controlled clinical trial using fresh honey with a determined amount of viable LAB is required. A recent study has shown that chestnut honey maintains the viability of a high concentration of probiotic bacteria for a prolonged period of time. Additionally, chestnut honey harboring probiotic bacteria exhibited a greater cytotoxic effect against malignant cells than honey or probiotics alone (Celebioglu, 2020b). The current study, as well as a number of previous studies, has shown that certain varieties of honey are an important source of probiotic bacteria and can be utilized as such (in raw form) for the treatment and prevention of human illnesses (Ebrahimi et al., 2021). An important point regarding using honey as a probiotic product is that honey should not be processed and heated because LAB can be killed at high temperatures. Similarly, bee-derived antibacterial compounds can also be destroyed by heating (Hussain et al., 2015). Therefore, honey intended to be used as a probiotic as well as an antibiotic should not be heated at all (Celebioglu, 2020b).
Conclusions: Since honey is produced by a variety of different types of honey bees from diverse bee flora and LAB originate from both insect and plant kingdoms, there is a possibility of a diverse range of LAB found in honey collected from different geographical areas. The results of the present study support this statement. The present study isolated and identified Lactobacillus rhamnosus, which has been used as a probiotic strain in multiple commercially available products.
Availability of data and materials: The data on the 16S rRNA gene sequence generated in this study can be found in the GenBank repository under accession numbers OL539410 to OL539415 (https://www.ncbi.nlm.nih.gov/ nuccore/OL539410).
Authors' contributions: The study was designed by Muhammad Barkaat Hussain and Yasser Mahmoud Kamel Mohamed Aly. Yasser Mahmoud Kamel, Imtiaz Ahmed Qureshi, Muhammad Yasir and Zia Ullah performed the experiments. Muhammad Barkaat Hussain, Asif Ahmad M. Jiman-Fatani, Marwan A Bakarman and Saleh M. Al-Maaqar drafted the manuscript. All of the authors approved the final manuscript.
Acknowledgements: This project was funded by the Deanship of Scientific Research (DSR) at King Abdulaziz University, Jeddah, under grant number G-167-828-1438. The authors, therefore, acknowledge and thank DSR for technical and financial support.
REFERENCES
- Al-Ghamdi, A., and A. Nuru (2013). Beekeeping in the Kingdom of Saudi Arabia past and present practices. Bee World. 90(2): 26-29.
- Al-Masaudi, S. B., Hussain, M. B., Al-Maaqar, S. M., Al Jaouni, S., & Harakeh, S. (2020). In vitro antibacterial activity of honey against multidrug-resistant Shigella sonnei. Complement Ther Clin Pract. 4: 101257.
- Alatawy, M., Al-Attas, S. G., Assagaf, A. I., Al-Hindi, R. R., Alghamdi, K. M., Mahyoub, J. A., Edris, S. (2021). Gut Microbiome of Two Different Honeybee Workers Subspecies in Saudi Arabia. Biosciences Biotechnology Research Asia. 17(4): 659.
- Alqarni, A., Hannan, M., Owayss, A., & Engel, M. (2011). The indigenous honey bees of Saudi Arabia (Hymenoptera, Apidae, Apis mellifera jemenitica Ruttner): Their natural history and role in beekeeping. ZooKeys. 134: 83-98.
- Altschul, S. F., Madden, T. L., Schäffer, A. A., Zhang, J., Zhang, Z., Miller, W., & Lipman, D. J. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25(17): 3389-3402.
- Anania, C., Di Marino, V. P., Olivero, F., De Canditiis, D., Brindisi, G., Iannilli, F., Duse, M. (2021). Treatment with a Probiotic Mixture Containing Bifidobacterium animalis Subsp. Lactis BB12 and Enterococcus faecium L3 for the Prevention of Allergic Rhinitis Symptoms in Children: A Randomized Controlled Trial. Nutrients. 13(4): 1315.
- Anju, V., Dyavaiah, M., and Siddhardha, B. (2021). Quorum Sensing as a Therapeutic Target in the Treatment of Chronic Wound Infections. In Wound Healing Research (pp. 643-659): Springer.
- Ansari, F., & Pourjafar, H. (2021). Nondairy Foods as Potential Carriers of Probiotic Bacteria and Postbiotics. Probiotic Bacteria and Postbiotic Metabolites: Role in Animal and Human Health. 351-373.
- Asama, T., Arima, T. H., Gomi, T., Keishi, T., Tani, H., Kimura, Y., Hashimoto, K. (2015). Lactobacillus kunkeei YB 38 from honeybee products enhances IgA production in healthy adults. J Appl Microbiol. 119(3): 818-826.
- Aween, M. M., Hassan, Z., Muhialdin, B. J., Eljamel, Y. A., Al‐Mabrok, A. S. W., & Lani, M. N. (2012). Antibacterial Activity of Lactobacillus acidophilus Strains Isolated from Honey Marketed in Malaysia against Selected Multiple Antibiotic Resistant (MAR) Gram‐Positive Bacteria. J Food Sci. 77(7): M364-M371.
- Balzan, S., Carraro, L., Merlanti, R., Lucatello, L., Capolongo, F., Fontana, F., Cardazzo, B. (2020). Microbial metabarcoding highlights different bacterial and fungal populations in honey samples from local beekeepers and market in north-eastern Italy. Int J Food Microbiol. 334: 108806.
- Bouzo, D., Cokcetin, N. N., Li, L., Ballerin, G., Bottomley, A. L., Lazenby, J., Harry, E. J. (2020). Characterizing the mechanism of action of an ancient antimicrobial, manuka honey, against Pseudomonas aeruginosa using modern transcriptomics. MSystems. 5(3): e00106-00120.
- Brudzynski, K., & Sjaarda, C. P. (2021). Colloidal structure of honey and its influence on antibacterial activity. Compr Rev Food Sci Food Saf. 20(2): 2063-2080.
- Castro, V. M. R., da Mota Silva, M., de Souza, E. R. P., Guerra, A. F., Riger, C. J., Laureano-Melo, R., & Luchese, R. H. (2021). Role of milk and honey in the tolerance of lactobacilli to oxidative stress. Braz J Microbiol. 52(2): 883-893.
- Celebioglu, H. U., (2020b). Probiotic bacteria grown with chestnut honey enhance in vitro cytotoxicity on breast and colon cancer cells. Arch Biol Sci. 72(3), 329-338.
- Chaturvedi, S., Chakraborty, S., & Technology. (2021). Review on potential non‐dairy synbiotic beverages: a preliminary approach using legumes. Int J Food Sci. 56(5): 2068-2077.
- Chen, X., Wu, Y., Hu, Y., Zhang, Y., & Wang, S. (2021). Lactobacillus rhamnosus GG Reduces β-conglycinin-Allergy-Induced Apoptotic Cells by Regulating Bacteroides and Bile Secretion Pathway in Intestinal Contents of BALB/c Mice. Nutrients. 13(1): 55.
- Davis, C. (2005). The use of Australian honey in moist wound management. A report for the Rural Industries Research and Development Corporation. Kingston, Australia.
- Diez-Gutiérrez, L., San Vicente, L., Barron, L. J. R., del Carmen Villaran, M., & Chávarri, M. (2020). Gamma-aminobutyric acid and probiotics: Multiple health benefits and their future in the global functional food and nutraceuticals market. J Funct Foods. 64: 103669.
- Dinçer, E., & Kıvanç, M. (2021). In vitro evaluation of probiotic potential of Enterococcus faecium strains isolated from Turkish pastırma. Arch Microbiol. 1-11.
- Dioso, C. M., Vital, P., Arellano, K., Park, H., Todorov, S. D., Ji, Y., & Holzapfel, W. (2020). Do Your Kids Get What You Paid for? Evaluation of Commercially Available Probiotic Products Intended for Children in the Republic of the Philippines and the Republic of Korea. Foods. 9(9): 1229.
- Ebrahimi, M., Sadeghi, A., Rahimi, D., Purabdolah, H., & Shahryari, S. (2021). Postbiotic and Anti-aflatoxigenic Capabilities of Lactobacillus kunkeei as the Potential Probiotic LAB Isolated from the Natural Honey. Probiotics Antimicrob Proteins. 13(2): 343-355.
- Endo, A., Irisawa, T., Futagawa-Endo, Y., Takano, K., du Toit, M., Okada, S., & Dicks, L. M. (2012). Characterization and emended description of Lactobacillus kunkeei as a fructophilic lactic acid bacterium. Int J Syst Evol Microbiol. 62(3): 500-504.
- Endo, A., & Salminen, S. (2013). Honeybees and beehives are rich sources for fructophilic lactic acid bacteria. Syst Appl Microbiol. 36(6): 444-448.
- Hajigholizadeh, M., Mardani, K., Moradi, M., & Jamshidi, A. (2020). Molecular detection, phylogenetic analysis, and antibacterial performance of lactic acid bacteria isolated from traditional cheeses, North‐West Iran. Food Sci Nutr. 8(11): 6007-6013.
- Hakala, S., Alghamdi, M. A., Paasonen, P., Vakkari, V., Khoder, M. I., Neitola, K., Shabbaj, I. I. (2019). New particle formation, growth and apparent shrinkage at a rural background site in western Saudi Arabia. Atmos Chem Phys Discuss. 19(16): 10537-10555.
- Hermanns, R., Mateescu, C., Thrasyvoulou, A., Tananaki, C., Wagener, F. A., & Cremers, N. (2020). Defining the standards for medical grade honey. J Apic Res. 59(2): 125-135.
- Hernández-González, J. C., Martínez-Tapia, A., Lazcano-Hernández, G., García-Pérez, B. E., & Castrejón-Jiménez, N. S. (2021). Bacteriocins from Lactic Acid Bacteria. A Powerful Alternative as Antimicrobials, Probiotics, and Immunomodulators in Veterinary Medicine. Animals. 11(4): 979.
- Homayoni Rad, A., Vaghef Mehrabany, E., Alipoor, B., & Vaghef Mehrabany, L. (2016). The comparison of food and supplement as probiotic delivery vehicles. Crit Rev Food Sci Nutr. 56(6): 896-909.
- Hussain, M. B. (2018). Role of honey in topical and systemic bacterial infections. J Altern Complement Med. 24(1): 15-24.
- Hussain, M. B., Hannan, A., Akhtar, N., Fayyaz, G. Q., Imran, M., Saleem, S., & Qureshi, I. A. (2015). Evaluation of the antibacterial activity of selected Pakistani honeys against multi-drug resistant Salmonella typhi. BMC Complement Altern Med. 15(1): 1-9.
- Kafantaris, I., Amoutzias, G. D., & Mossialos, D. (2021). Foodomics in bee product research: a systematic literature review. Eur Food Res Technol. 247(2): 309-331.
- Karakonstantis, S., Gikas, A., Astrinaki, E., & Kritsotakis, E. I. (2020). Excess mortality due to pandrug-resistant Acinetobacter baumannii infections in hospitalized patients. J Hosp Infect. 106(3): 447-453.
- Kňazovická, V., Gábor, M., Miluchová, M., Bobko, M., & Medo, J. (2021). Diversity of bacteria in Slovak and foreign honey, with assessment of its physico-chemical quality and counts of cultivable microorganisms. J Microbiol, Biotechnol and Food Sciences. 2021: 414-421.
- Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 35(6): 1547-1549.
- Lane, D. (1991). 16S/23S rRNA sequencing. Nucleic acid techniques in bacterial systematics. 115-175.
- Lashani, E., Davoodabadi, A., & Dallal, M. M. S. (2020). Some probiotic properties of Lactobacillus species isolated from honey and their antimicrobial activity against foodborne pathogens. Paper presented at the Veterinary Research Forum.
- Manasian, P., Bustos, A.-S., Pålsson, B., Håkansson, A., Peñarrieta, J. M., Nilsson, L., & Linares-Pastén, J. A. (2020). First Evidence of Acyl-Hydrolase/Lipase Activity From Human Probiotic Bacteria: Lactobacillus rhamnosus GG and Bifidobacterium longum NCC 2705. Front Microbiol. 11: 1534.
- Mishra, S. S., Das, U., Biswal, R., & Behera, S. S. (2021). Probiotic beverages in India: History and current developments. In Probiotic Beverages (pp. 9-33): Elsevier.
- Olofsson, T. C., Butler, E., Lindholm, C., Nilson, B., Michanek, P., & Vásquez, A. (2016). Fighting off wound pathogens in horses with honeybee lactic acid bacteria. Curr Microbiol. 73(4): 463-473.
- Olofsson, T. C., Butler, È., Markowicz, P., Lindholm, C., & Larsson, L. (2016). Lactic acid bacterial symbionts in honeybees–an unknown key to honey's antimicrobial and therapeutic activities. Int Wound J. 13(5): 668-679.
- Olofsson, T. C., Butler, È., Markowicz, P., Lindholm, C., Larsson, L., & Vásquez, A. (2014). Lactic acid bacterial symbionts in honeybees–an unknown key to honey's antimicrobial and therapeutic activities. Int Wound J. 13(5): 668-679.
- Olofsson, T.C., and Vásquez, A., (2008). Detection and identification of a novel lactic acid bacterial flora within the honey stomach of the honeybee Apis mellifera. Curr Microbiol, 57(4): 356-363.
- Oruc, O., Ceti̇n, O., Darilmaz, D.O. and Yüsekdag, Z.N. (2021). Determination of the biosafety of potential probiotic Enterococcus faecalis and Enterococcus faecium strains isolated from traditional white cheeses. LWT. 148:111741.
- Peng, M., Tabashsum, Z., Anderson, M., Truong, A., Houser, A. K., Padilla, J., Safety, F. (2020). Effectiveness of probiotics, prebiotics, and prebiotic‐like components in common functional foods. Compr Rev Food Sci Food Saf. 19(4): 1908-1933.
- Peng, X., Wang, R., Hu, L., Zhou, Q., Liu, Y., Yang, M., Feng, B. (2019). Enterococcus faecium NCIMB 10415 administration improves the intestinal health and immunity in neonatal piglets infected by enterotoxigenic Escherichia coli K88. J Anim Sci Biotechnol. 10(1): 1-15.
- Petrova, M. I., Reid, G., & Ter Haar, J. A. (2021). Lacticaseibacillus rhamnosus GR-1, aka Lactobacillus rhamnosus GR-1: Past and Future Perspectives. Trends Microbiol.
- Roobab, U., Batool, Z., Manzoor, M. F., Shabbir, M. A., Khan, M. R., & Aadil, R. M. (2020). Sources, formulations, advanced delivery and health benefits of probiotics. J Current Opinion in Food Science. 32: 17-28.
- Scripcă, L. A., & Amariei, S. (2021). The Use of Ultrasound for Preventing Honey Crystallization. J Foods. 10(4): 773.
- Sharifi-Rad, J., Rodrigues, C. F., Stojanović-Radić, Z., Dimitrijević, M., Aleksić, A., Neffe-Skocińska, K.,Milton Prabu, S. (2020). Probiotics: Versatile bioactive components in promoting human health. J Medicina. 56(9): 433.
- Sharma, M., Wasan, A., & Sharma, R. K. (2021). Recent developments in probiotics: An emphasis on Bifidobacterium. J Food Bioscience. 100993.
- Śliżewska, K., Markowiak-Kopeć, P., & Śliżewska, W. (2021). The Role of Probiotics in Cancer Prevention. Cancers. 13(1): 20.
- Stasiak-Różańska, L., Berthold-Pluta, A., Pluta, A. S., Dasiewicz, K., Garbowska, M., & Health, P. (2021). Effect of Simulated Gastrointestinal Tract Conditions on Survivability of Probiotic Bacteria Present in Commercial Preparations. Int J Environ Res Public Health. 18(3): 1108.
- Stivala, A., Carota, G., Fuochi, V., & Furneri, P. M. (2021). Lactobacillus rhamnosus AD3 as a Promising Alternative for Probiotic Products. Biomolecules. 11(1): 94.
- Tamura, K., & Nei, M. (1993). Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol. 10(3): 512-526.
- Thrasyvoulou, A., Tananaki, C., Goras, G., Karazafiris, E., Dimou, M., Liolios, V., Gounari, S. (2018). Legislation of honey criteria and standards. J Apic Res. 57(1): 88-96.
- Tkhruni, F. N., Aghajanyan, A. E., Balabekyan, T. R., Khachatryan, T. V., & Karapetyan, K. J. (2020). Characteristic of bacteriocins of Lactobacillus rhamnosus BTK 20-12 potential probiotic strain. Probiotics Antimicrob Proteins. 12(2): 716-724.
- Vásquez, A., Forsgren, E., Fries, I., Paxton, R.J., Flaberg, E., Szekely, L. and Olofsson, T.C., (2012). Symbionts as major modulators of insect health: lactic acid bacteria and honeybees. PloS one, 7(3): p.e33188.
- Wang, Y., Jiang, Y., Deng, Y., Yi, C., Wang, Y., Ding, M.,Wong, A. (2020). Probiotic Supplements: Hope or Hype? Front Microbiol. 11, 160. doi:10.3389/fmicb.2020.00160.
- Wang, Y., Wu, J., Lv, M., Shao, Z., Hungwe, M., Wang, J., Geng, W. (2021). Metabolism Characteristics of Lactic Acid Bacteria and the Expanding Applications in Food Industry. Front Bioeng Biotechnol. 9: 1-19.
|





