EFFICIENCY OF THYMOL AND CARVACROL IN ACTIVATION OF PATHOGENESIS RELATED GENES IN WHEAT AGAINST PUCCINIA STRIIFORMIS
E. I. El-Shafeey1, A. A. Aboulila2, E. H. Wheish1, M. A. Ashmawy3and M. M. Elsharkawy1*
1Department of Agricultural Botany, Faculty of Agriculture, Kafrelsheikh University, 33516 Kafr El-Sheikh, Egypt
2Department of Genetics, Faculty of Agriculture, Kafrelsheikh University, 33516 Kafr El-Sheikh, Egypt
3Wheat Diseases Research Department Plant Pathology Research Institute, Agricultural Research Center, Giza, Egypt
*Corresponding author Email: mohsen.abdelrahman@agr.kfs.edu.eg
ABSTRACT
Genetic resistance, which can be succeeded by combining several resistance genes, is the most helpful technique for controlling stripe rust disease (Puccinia striiformis). Nineteen wheat commercial cultivars were assessed for stripe rust resistance and the existence of four resistance genes were validated.Yr5 gene was found in all the studied genotypes in different sizes ranged from 467 bp to 581 bp, while Yr18 appeared only in four studied genotypes. The potential of thymol and carvacrol as bioactive compounds was investigated against stripe rust disease. Using real-time PCR, the regulation of defense genes expression was assessed. The findings of the present investigation indicated that both thymol and carvacrol had protective effects against stripe rust. Treatments with thymol and carvacrol boosted the incubation and latent periods of the disease. Additionally, in both treatments, the level of infection was lower than the control group. A significant up-regulation of the defense genes in wheat (PR5, PR6, PR8 and PR12) was recorded after treatment with thymol (100 mg/l) for 0, 1, 3, 4, 8 and 14 days after inoculation (DAI) and the fold change was close to the transcription of PR genes in the resistance monogenic line Yr15. After inoculation, carvacrol (100 mg/l) induced a significant up-regulation in the expression of PR genes at all days, except at 3 DAI (for PR6 and PR8) and 4 DAI (for PR6 and PR12). These results suggest that thymol and carvacrol may be utilized as elicitors of systemic resistance to stripe rust in wheat.
Keywords: Wheat; stripe rust; Yr Resistance genes; Thymol; Carvacrol; induced resistance; Pathogenesis related genes.
https://doi.org/10.36899/JAPS.2022.5.0543
Published first online April 26, 2022
INTRODUCTION
Wheat is a widely cultivated crop around the globe, with 218 million hectares harvested in 2017 (FAO, 2017). One of the most critical variables affecting wheat yield is the occurrence of plant diseases. Wheat rust diseases have long been considered one of the most severe biotic production impediments worldwide. Puccinia striiformis f. sp. tritici (causes stripe rust), P. graminis f. sp. tritici (causes stem rust), and P. triticina (causes stripe rust) are three pathogens of wheat rust diseases. Stripe rust is a common disease throughout the winter and early spring (Chen, 2005; Ali et al., 2017). It causes production losses ranging from 10% to 70% in most wheat-growing areas (Chen, 2020). Cultivating resistant cultivars, using the right fungicide, and following the right cultural practices may all help to manage the disease. For more effective, economical, and long-term management of stripe rust, more research is needed to establish additional cultivars with sustainable and high-level resistance to compete pathogen virulence.
The best approach for controlling stripe rust disease is the development of resistant cultivars, however, the traditional breeding technique is insufficient. A total of 43 resistance genes have been detected in wheat plants. There are a large number of genes that are race-specific, and wheat cultivars that contain some of these genes have considered an important tool in breeding programs (Cao et al., 2001). Since molecular markers for particular stripe rust genes have been developed, detection of these genes can now be done regardless of phenotype. Molecular breeding based on marker assisted selection (MAS) may substantially improve breeding efficiency (Yu et al., 2004). The availability of closed associated DNA markers is a requirement for using MAS in traditional breeding. This may increase the development of resistant genotypes and therefore, serves as an adequate method for plant breeding (Koebner et al., 2003). To improve parental selection for crossover and monitoring genes in segregating progeny, markers may be employed to better define parental material (William et al., 2007).
Thymol and carvacrol as natural plant volatile compounds showed high antifungal effects (Soković et al., 2009; Spadaro and Gullino 2014). These prospective fungicidal chemicals' mechanism of action has yet to be completely investigated. However, the phenolic structures of these compounds are very effective against pathogens. The performance of thymol and carvacrol is frequently due to the combination of various chemical compounds (Spadaro and Gullino 2014; Banani et al., 2018). However, nothing is known about PR gene induction's role of plant essential oils against plant pathogens.
Plants use a variety of immunological mechanisms, including pathogen detection, defense signal pathway activation, and the synthesis of antifungal substances such as PR proteins, all of which help to decrease pathogen invasion and reproduction (Sels et al., 2008). PR proteins, for example, are good targets for developing long-lasting and diverse resistance to fungal pathogens (Ali et al., 2017). Several transcriptional findings showed that PR genes are up-regulated in various crops following fungal infections, indicating that they play a role in disease resistance. PR gene expression is low under normal conditions, but it rises following fungal infection in both infected and non-infected parts of the plant, triggering resistance pathway (Návarová et al., 2012; Esmail et al., 2020; Salman et al., 2021). Furthermore, numerous in vivo experiments have shown that PR proteins target or hydrolyze fungal cell walls, causing cell death. PR5 and PR12 have long been thought to be the most effective antifungal proteins in plants. Furthermore, overexpression of PR genes, either alone or in combination, improves resistance against fungal pathogens in different crops (Dai et al., 2016). PR proteins are found across the plant kingdom and are made up of 17 different protein families. These proteins include thaumatin-like proteins (PR5), proteinase inhibitors (PR6), endochitinases (PR8) and defensins (PR12). Although these PR proteins are known to play key roles in plant resistance to different stresses, little is known about their involvement in wheat-P.striiformis interactions (Farrakh et al., 2018).
This study aimed to validate the roles of DNA markers (Yr5, Yr10, Yr15 and YR18) in stripe rust resistance in different wheat genotypes and to identify the effects of thymol and carvacrol as inducers of resistance against stripe rust through measuring the transcription levels of different PR genes.
MATERIALS AND METHODS
All the greenhouse experiments were performed in Agriculture Research Center, Giza, Egypt. Field experiments were conducted in two locations Sakha (6-27 ºC temperature and 36-55% relative humidity) and Gemmeiza (5-28 ºC temperature and 35-54% relative humidity).
Evaluation of the response of wheat varieties to stripe rust disease under field conditions: Wheat varieties containing 23 stripe rust resistance genes and another 19 varieties were tested for stripe rust resistance at adult plant stages. During the 2017/18 (8-27 ºC temperature and 36-54% relative humidity), 2018/19 (6-28 ºC temperature and 35-56% relative humidity), and 2019/2020 (5-26 ºC temperature and 36-55% relative humidity) growing seasons, adult plant resistance was assessed in the field on the same set of materials. During the season (at the booting stage), the required agricultural methods for inoculation were carried out according to Tervet and Cassell (1951). The modified Cobb's Scale was utilized to determine the severity of the disease (Peterson et al., 1948). According to Roelfs et al. (1992), infection responses were classified as susceptible (S), moderately susceptible (MS), moderately resistant (MR) and resistant (R).
Genomic DNA extraction: The genomic DNA of wheat plants was obtained from fresh leaves (100-150 mg) following the CTAB (Cetyltrimethylethyl Ammonium Bromide) method (Murray and Thompson, 1980). The A260/A280 ratio was verified for measuring DNA concentration. For PCR amplification, high quality DNA samples were used. The stocks were diluted to a final concentration of 40 ng/l of DNA before being utilized in further experiments.
DNA marker analysis: Four DNA primers related to Yr5, Yr10, Yr15 and Yr18 resistance genes as shown in Table (1) were screened. Results of these SSR markers were considered for evaluating molecular differences among 19 wheat genotypes with Yr5, Yr10, Yr15 and Yr18 monogenic lines as positive controls.
Table 1: List of four DNA marker sequences used in validation of resistance genes in wheat.
|
Resistance gene
|
Marker
|
Marker type
|
Primer Sequence (5ﹶ → 3ﹶ)
|
Expected product size (bp)
|
Reference
|
|
Yr5
|
STS-7
STS-8
|
STS
|
GTACAATTCACCTAGAGT
GCAAGTTTTCTCCCTATT
|
478
|
Chen et al. (2003)
|
|
Yr10
|
Xpsp3000
|
SSR
|
F: GCAGACCTGTGTCATTGGTC
R: GATATAGTGGCAGCAGGATACG
|
260
|
Wang et al. (2002)
|
|
Yr15
|
Xbarc8
|
SSR
|
F: GCGGGAATCATGCATAGGAAAACAGAA
R: GCGGGGGCGAAACATACACATAAAAACA
|
190
|
Murphy et al. (2009)
|
|
Yr18
|
Cssfr1
|
SSR
|
F: TTGATGAAACCAGTTTTTTTTCTA
R: GCCATTTAACATAATCATGATGGA
|
517
|
Lagudah et al. (2009)
|
All PCR products were run on 3 % agarose gel containing 5 µl of ethidium bromide (1 µg/ml). Along with the samples, a conventional DNA marker was run. The separated bands were photographed using a photo Doc-ItTM Imaging System and examined after being verified with a Benchtop UV-transilluminator.
Band scoring: Data for DNA markers were scored for computer analysis and the numbers and sizes of alleles produced by each primer were recorded and analyzed by GelAnalyzer3 program. DNA ladder marker 1 kb (+) (enzynomics) was used as the reference. Each polymorphic DNA fragment was treated as a separate locus since it corresponded to a distinct position in the genome.
Control of stripe rust disease by thymol and carvacrol under greenhouse conditions: Wheat (Triticum aestivum L.) seedlings of Egyptian genotype Gemmeiza-11 were put in a growth chamber at 20 ºC and subjected to a photoperiod of 12 h. Seedlings after seven days were transferred to a greenhouse, and infected with rust urediniospores. Depending on the infection types generated, seedling response was recorded as resistant (R) or susceptible (S) (Table 1). Seedlings were sprayed with 100 mg/l thymol (base chemicals India company) and carvacrol (Sigma-Aldrich Co.) at 24 h before and after inoculation. The period from inoculations to the onset of the first pustule is considered the incubation period (IP). The latent period (LP) was determined according to Parlevliet (1975) by adding up the number of pustules seen on marked leaves per day until no pustules appeared. In addition, the period from inoculation to the development of 50% of the pustules was reported. The size of 20 pustules were measured to compare pustule size on the first leaf. The number of pustules per unit leaf area (2.0 × 0.5 cm2) on the top surface of the leaves was calculated using the technique reported by Parlevliet and Kuiper (1977).
RNA isolation and RT-qPCR procedures: Leaves were harvested and washed to remove thymol and carvacrol excess. Leaves were harvested at the following time points; 0, 1, 3, 4, 8 and 14 days after inoculation (DAI). These time intervals were chosen based on the identified stages of the infection process: [1] interaction between P. striiformis and plants (0 DAI, just inoculation), [2] P. striiformis entered plant tissue (1 DAI), [3] haustoria formation and starting of the life cycle (3 and 4 DAI), [4] disease symptom appearance (8 DAI) and [5] sporulation (14 DAI) (Farrakh et al., 2018; Zhang et al., 2012). For Gemmieza-11 genotype, control plants (inoculated only with P. striiformis) were subjected to the same procedure, using distilled water. Inoculated plants without thymol and carvacrol treatments of Yr15 resistance monogenic line were used as a positive control and the non-inoculated plants of both genotypes (Gemmieza-11 and Yr15) at all different time points were used to normalize folding changes of each gene. The leaves were wrapped in labeled aluminum foil and cleaned with 95% ethanol before being frozen in liquid nitrogen and kept at –80°C until RNA isolation.
RNA extraction and cDNA synthesis: Total RNA was purified from wheat samples using Trizol reagent (iNtRON Biotechnology, Inc.) as directed by the manufacturer. Electrophoresis on a 2% ethidium bromide stained agarose gel was used to confirm the integrity of the RNA. The TopscriptTM CDNA synthesis kit (enzynomics, Korea) was used to synthesize cDNA from a concentration of 2 µg RNA (Elsharkawy et al., 2012).
qRT-PCR assay: For qRT-PCR analysis, four specific primers were used to determine the expression of pathogenesis related genes PR5, PR6, PR8 and PR12, with the β-tubulin as a housekeeping gene. Primers were used from previous studies as presented in Table (2). Ten microliters (μL) of TOPreal™ qPCR 2X PreMIX (SYBR green with low ROX, enzynomics, Korea), 0.5 μM of each primer, and 2 μL of cDNA were used in the reaction. Detection of gene expression was done in triplets using the Thermocycler (Bio-Rad, C-1000) real-time PCR system. The program was set up in the first of an initial denaturation at 94 °C for 5 min followed by 40 cycles comprising of: 94 °C for 30 s, annealing at 55 °C for 30 s and 72 °C for 1 min. The relative expressions were calculated by the 2−ΔΔct method described by Livak and Schmittgen (2001) and values of each gene have normalized its folding changes against the housekeeping gene (β-tubulin) and the control values (non-inoculated plants) at each time point.
Statistical analysis: The experiments were conducted using a completely randomized approach and one-way ANOVA was used. A total of three repetitions were conducted of all experiments. Fisher's least significant difference test was used to statistically analyze the data. Analysis was done by XLSTAT Pro statistical analysis software (Addinsoft, New York, USA) at P ≤ 0.05.
Table 2: Accession numbers, function and annealing temperatures of PR genes used for qRT-PCR assay.
RESULTS AND DISCUSSION
Efficiency of Yr genes against stripe rust disease in adult plants: Under natural infection, the efficiency of resistance genes (Yr's) in the adult plant stage was evaluated. During three seasons 2017/18, 2018/19, and 2019/20, the genotypes were tested at two sites, Sakha and El-Gemmeiza. During the first growing season, the monogenic lines Yr1, Yr5, Yr10, Yr15, Yr17, Yr27, Yr CV and YrSP grown in Sakha and El- Gemmeiza were resistant to stripe rust. Meanwhile, Yr1, Yr5, Yr10, Yr15, Yr17, Yr27, Yr CV and YrSP were resistant to stripe rust in the second growing season. Lastly, the lines Yr1, Yr5, Yr10 and Yr15 were resistant to stripe rust in the third growing season (Table 3).
The evaluation of stripe rust resistance in the field indicated that Yr5, Yr10, Yr15 and Sakha 61 were resistant genotypes in the three studied seasons at different locations Sakha and Gemmeiza. However, this study indicates that the resistance genes Yr5, Yr10, Yr15 and Yr18 and other Yr genes could contribute to resistance for stripe rust pathogen. The obtained results are in accordance with those reported by Sadravi (2008); Pourkhorshid et al. (2014); Zhou et al. (2014). Since Yr5 is a race-specific resistance gene, it should be combined with other potent genes and/or race- non-specific adult-plant resistance genes. A combination like this may offer excellent resistance against stripe rust (Yan et al., 2003).
Field assessment of Egyptian wheat cultivars: The varieties Sakha 61, Sakha 94, Sakha 95, Giza 168, Giza 171, Gemmeiza 12, Sids 14 and Misr 3 were resistant to stripe rust disease in the first growing season (Table 4). Meanwhile, Sakha 61, Sakha 95, Giza 171, Sakha 94, Misr 3 and Gemmeiza 12 were resistant in the second growing season and Sakha 93, Sakha 8, Sakha 69, Sids 11, Sids 13, Sids 14, Gemmeiza 7, Gemmeiza 9, Misr 1 and Misr 2 were resistant in the third growing season (Table 4).
The collected data suggested that the severity of stripe rust was lower in the growing season 2017/18 than in 2018/19 and 2019/20 growing seasons. The rust data also revealed that the severity of disease at Sakha was significantly higher than at El- Gemmeiza. Major or minor genes, or both, control disease resistance; nevertheless, a complimentary impact between major genes may increase the variety's response and result in higher degrees of resistance. Variable variations have diverse reactions depending on the genetic composition of the major genes (Simons et al., 1978; Omara et al., 2021).
Validation of stripe rust resistance genes: The molecular analysis of 19 Egyptian wheat genotypes was performed using four stripe rust resistance genes selected based on their tight linkages at adult stage. For Yr5 resistance gene, the results indicated that monogenic line Yr5 separated a DNA fragment of 478 bp as a positive control (Figure 1), while it separated four alleles in the studied genotypes ranged from 467 to 581 bp with 114 bp size differences. Ten out of the 19 studied genotypes amplified a fragment of 478 bp as present in positive control indicating the presence of Yr5 in these genotypes as shown in Table 3. The evaluation of stripe rust resistance in the field indicated that Yr5, Yr10, Yr15 and Sakha 61 are resistant genotypes in the three studied seasons at different locations (Sakha and El-Gemmeiza).
Table 3: Response of 23 monogenic lines against stripe rust disease under field conditions at Sakha and El-Gemmeiza locations.
|
No.
|
Monogenic Lines
|
Rust reaction
|
|
Sakha
|
Gemmeiza
|
|
2017/18
|
2018/19
|
2019/20
|
2017/18
|
2018/19
|
2019/20
|
|
1
|
Yr 1
|
0
|
0
|
0
|
0
|
0
|
0
|
|
2
|
Yr2
|
20S
|
40S
|
30S
|
20S
|
50S
|
40S
|
|
3
|
Yr3
|
30S
|
50S
|
40S
|
30S
|
40S
|
50S
|
|
4
|
Yr4
|
40S
|
60S
|
50S
|
30S
|
50S
|
40S
|
|
5
|
Yr5
|
0
|
0
|
0
|
0
|
0
|
0
|
|
6
|
Yr6
|
80S
|
100S
|
90S
|
70S
|
100S
|
90S
|
|
7
|
Yr7
|
80S
|
100S
|
90S
|
90S
|
100S
|
80S
|
|
8
|
Yr8
|
70S
|
90S
|
80S
|
70S
|
80S
|
70S
|
|
9
|
Yr9
|
80S
|
100S
|
70S
|
70S
|
100S
|
80S
|
|
10
|
Yr10
|
0
|
0
|
0
|
0
|
0
|
0
|
|
11
|
Yr15
|
0
|
0
|
0
|
0
|
0
|
0
|
|
12
|
Yr17
|
0
|
40S
|
30S
|
10S
|
30S
|
20S
|
|
13
|
Yr18
|
70S
|
90S
|
100S
|
60S
|
80S
|
80S
|
|
14
|
Yr21
|
50S
|
60S
|
50S
|
40S
|
70S
|
60S
|
|
15
|
Yr25
|
40S
|
70S
|
60S
|
30S
|
70S
|
60S
|
|
16
|
Yr27
|
0
|
50S
|
30S
|
0
|
40S
|
30S
|
|
17
|
Yr28
|
20S
|
60S
|
50S
|
10S
|
50S
|
40S
|
|
18
|
Yr29
|
20S
|
60S
|
40S
|
30S
|
50S
|
50S
|
|
19
|
Yr CV
|
5R
|
20S
|
30S
|
5R
|
20S
|
10S
|
|
20
|
Yr SP
|
10R
|
20S
|
20S
|
5R
|
30S
|
10S
|
|
21
|
Yr SU
|
30S
|
50S
|
40S
|
20S
|
40S
|
20S
|
|
22
|
Yr SD
|
40S
|
60S
|
50S
|
30S
|
50S
|
40S
|
|
23
|
Yr A
|
80S
|
100S
|
90S
|
70S
|
100S
|
100S
|
|
L.S.D 5%
|
12.360
|
10.182
|
8.401
|
9.700
|
8.401
|
8.010
|
Table 4: Response of 19 wheat varieties for strip rust under field conditions in 2017/18, 2018/19 and 2019/2020 growing seasons.
|
No
|
Wheat variety
|
Location /Rust reaction
|
|
2017/18
|
2018/19
|
2019/20
|
|
Sakha
|
Gemmeiza
|
Sakha
|
Gemmeiza
|
Sakha
|
Gemmeiza
|
|
1
|
Sakha 8
|
90S
|
80S
|
90S
|
90S
|
80S
|
80S
|
|
2
|
Sakha 61
|
0
|
0
|
0
|
0
|
0
|
0
|
|
3
|
Sakha 69
|
80 S
|
70 S
|
90 S
|
90 S
|
80 S
|
70 S
|
|
4
|
Sakha 93
|
40S
|
30S
|
70S
|
80S
|
60S
|
50S
|
|
5
|
Sakha 94
|
5S
|
5S
|
10S
|
20S
|
5S
|
5S
|
|
6
|
Sakha 95
|
0
|
0
|
TrMS
|
TrMS
|
TrMS
|
TrMS
|
|
7
|
Gemmeiza 7
|
40S
|
50S
|
80S
|
90S
|
70S
|
70S
|
|
8
|
Gemmeiza 9
|
40S
|
50S
|
70S
|
80S
|
60S
|
50S
|
|
9
|
Gemmeiza 11
|
80S
|
80S
|
100S
|
100S
|
90S
|
80S
|
|
10
|
Gemmeiza 12
|
0
|
0
|
TrMS
|
TrMS
|
TrMS
|
TrMS
|
|
11
|
Giza 168
|
10S
|
10S
|
40S
|
50S
|
30S
|
40S
|
|
12
|
Giza 171
|
0
|
0
|
10S
|
20S
|
5S
|
5S
|
|
13
|
Sids 12
|
80S
|
90S
|
100S
|
100S
|
90S
|
100S
|
|
14
|
Sids 13
|
80S
|
70S
|
80S
|
90S
|
70S
|
50S
|
|
15
|
Sids 14
|
0
|
0
|
40S
|
50S
|
30S
|
20S
|
|
16
|
Shandaweil 1
|
0
|
0
|
50S
|
40S
|
40S
|
50S
|
|
17
|
Misr 1
|
10S
|
20S
|
80S
|
70S
|
80S
|
90S
|
|
18
|
Misr 2
|
10S
|
20S
|
90S
|
90S
|
90S
|
80S
|
|
19
|
Misr 3
|
0
|
0
|
TrMS
|
TrMS
|
TrMS
|
TrMS
|
|
L.S.D 5%
|
12.120
|
8.582
|
11.819
|
9.645
|
11.328
|
10.819
|

Figure 1. Band patterns of stripe rust resistance genes in 19 tested genotypes in addition to monogenic lines. M; 1 kb (+) DNA ladder.
As shown in Figure (1), stripe rust resistance dominant gene Yr10 was validated in 19 wheat genotypes, the monogenic line Yr10 (a band with size of 219 bp in the positive control) and two commercial varieties (Sakha 8 and Gemmeiza 7). Four alleles ranged in their molecular size from 190 to 300 bp were found in the rest genotypes.
For Yr15 resistance gene, the corresponding allele found in the positive control Yr15 monogenic line was 208 bp in size, while the 19 tested genotypes were divided into three groups. The first group containing the two genotypes, Sakha 8 and Sakha 61, presented an allele with molecular size of 229 bp, the second group (presented a 255 bp allele) appeared in 14 genotypes, while the rest three genotypes did not record any alleles for this gene.
In validation of Yr18 stripe rust resistance gene, only four genotypes (Sakha-8, Sakha-94, Sakh-95 and Sids-13) out of the 19 studied were found to carry this gene and presented the same allele appeared in monogenic line Yr18. Therefore, it is likely that if the genotype does not carry the corresponding alleles presented in the positive control, the genotypes' resistance could be due to the presence of another effective gene. On the other hand, susceptibility of the genotype that has the corresponding alleles of resistance gene indicates that this gene cannot be expressed in this genotype.
New progress in the molecular analysis of resistance genes have allowed the development of direct markers that integrate significant race-specific resistance with adult plant resistance genes (Sharma, 2012). Bariana et al. (2002) reported that the varieties with the Yr10 gene amplified a 258-260 bp fragment, but those without this gene only produce a 240 bp band fragment. This gene also expresses at the end of plant development, making it unsuitable for early selection of resistance to stripe rust. Additionally, this gene was present in the majority of cultivars and showed its effectiveness against the predominant and virulent pathogen 78S84 (Lagudah et al., 2009; Mukhtar et al., 2015). When combined with other resistance genes, this gene may improve protection against stripe rust (German and Kolmer, 1992). Based on previous studies, Yr5 and Yr10 genes were effective with high levels of resistance against all virulent races in North America (Bux et al., 2012), Iran (Afshari, 2008) and China (Chen, 2005). This fact indicated that Yr5 was a useful candidate for breeding programs.
Table 5: Absence/presence of corresponding alleles for four resistance genes in addition to other alleles in 19 Egyptian commercial wheat cultivars
|
Number
|
Genotype
|
Yr resistance genes
|
|
Yr5 (478 bp)
|
Yr10 (219 bp)
|
Yr15 (208 bp)
|
Yr18 (532 bp)
|
|
1
|
Sakha-8
|
+
|
+
|
229
|
+
|
|
2
|
Sakha-61
|
+
|
190
|
229
|
-
|
|
3
|
Sakha-69
|
467
|
190
|
255
|
-
|
|
4
|
Sakha-93
|
467
|
247
|
255
|
-
|
|
5
|
Sakha-94
|
+
|
247
|
-
|
+
|
|
6
|
Sakha-95
|
+
|
247
|
255
|
+
|
|
7
|
Gemmeiza-7
|
+
|
+
|
255
|
-
|
|
8
|
Gemmeiza-9
|
+
|
273
|
255
|
-
|
|
9
|
Gemmeiza -11
|
+
|
247
|
255
|
-
|
|
10
|
Gemmeiza-12
|
+
|
-
|
-
|
-
|
|
11
|
Giza-168
|
+
|
300
|
255
|
-
|
|
12
|
Giza-171
|
+
|
300
|
255
|
-
|
|
13
|
Sids-12
|
542
|
300
|
255
|
-
|
|
14
|
Sids-13
|
542
|
300
|
-
|
+
|
|
15
|
Sids-14
|
581
|
300
|
255
|
-
|
|
16
|
Shandaweil-1
|
581
|
300
|
255
|
-
|
|
17
|
Misr-1
|
581
|
273
|
255
|
-
|
|
18
|
Misr-2
|
581
|
300
|
255
|
-
|
|
19
|
Misr-3
|
581
|
300
|
255
|
-
|
|
Total No. of allels
|
4
|
5
|
2
|
1
|
|
Polymorphism %
|
100
|
100
|
100
|
100
|
|
Size range (bp)
|
114
|
110
|
26
|
0
|
+ indicates the presence of corresponding genes, - indicates the absence of genes
Effect of thymol and carvacrol on incubation, latent periods, and infection type: Figures 2 and 3 demonstrate the impacts of the thymol and carvacrol on stripe rust development, including incubation and latent periods and infection type. All parameters showed significant variations when different treatments were sprayed at 24 h before and after inoculation. Thymol and carvacrol treatments enhanced incubation and latent periods. Furthermore, as compared to the control, both treatments had the lowest infection type. However, the incubation and latent durations, as well as the infection type, grew more slowly than in the control treatment (Figs. 2 and 3).

Figure 2. Effect of thymol and carvacrol on stripe rust development (incubation, latent periods, and infection type) at seedling stage.
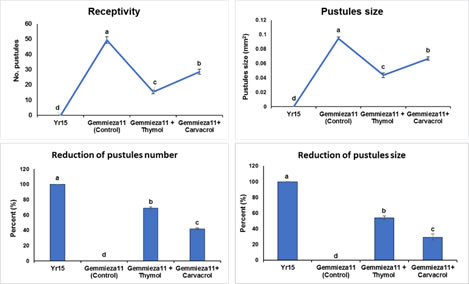
Figure 3. Effect of treatments on pustules size, number and receptivity of stripe rust urediniospores at seedling stage.
Plant extracts substantially extended the incubation and latent periods of wheat leaf rust (Omara et al., 2015 and 2019). Since thymol and carvacrol are currently extensively used as feed additives, these compounds may be useful candidates in the control of P. striiformis in wheat. Several researchers examined the function of thymol and carvacrol against bacterial infection through down-regulating the virulence factor production (Inamuco et al., 2011; Burt et al., 2016). Such molecules may have direct antifungal activity that can prevent P. striiformis growth via inactivation of virulence genes but this needs further studies.
Effect of thymol and carvacrol on PR protein genes expression: In order to analyze the up-regulation of PR genes involved in thymol and carvacrol- stimulated resistance in wheat, the expression of PR5, PR6, PR8 and PR12 was analyzed in treated wheat plants at 0, 1, 3, 4, 8 and 14 days after inoculation (Tables 6, 7, 8 and 9).
All of the pathogenesis related genes that were examined showed up-regulation by thymol and carvacrol at different time points compared with non-treated plants (Figure 4). Treatment with 100 mg/L thymol in susceptible Gemmiza 11 genotype induced up-regulation of the expression of all tested PR genes at 0, 1, 3, 4, 8 and 14 days after pathogen inoculation (DAI) compared to inoculated control plants. The fold change was very close to the expression of PR genes in the resistance monogenic line Yr15. While, carvacrol treatment (100 mg/L) showed significant differences of relative expression of PR genes at all days of inoculation compared to control, except after 3 DAI in PR6 and PR8 and at 4 DAI in PR6 and PR12. Also, treatment with carvacrol recorded down-regulation of PR8 at 14 DAI and PR12 at 1 and 3 DAI. These results suggest that thymol treatment was better than carvacrol in inducing resistance against P. striiformis in wheat.

Figure 4. Expression of pathogenesis-related protein genes PR5, PR6, PR8 and PR12 in wheat treated with thymol and carvacrol and inoculated plants at different days of inoculation.
Data from this study supported the effect of thymol and carvacrol in regulation of gene expression. In wheat, thymol and carvacrol have properties of inducers that can activate the resistance genes. Thyme oil enhanced apple resistance to B. cinerea because it encouraged the expression of PR8. Although this signal stimulated the plant, thyme produced a modest induction of resistance (PR8 expression level was 2.5-fold greater than the expression in water-treated fruits) after 6 h of pathogen inoculation (Banani et al., 2018). Also, Giovagnoni et al. (2020) confirmed that thymol and carvacrol concentrations of 0.47 mM inhibited cell invasion by lowering the expression of virulence genes such as SipA and they indicated that treatment with carvacrol showed expression significantly lower than thymol in all studied genes, except SipC.
Table 6: Relative expression of pathogen-related protein gene PR5 in Gemmieza-11 wheat genotype at different days after inoculation (DAI) with P. striiformis at seedling stage in comparison with Yr15 resistance monogenic line.
|
Days after inoculation
|
Yr15 gene
|
Gemmieza-11
|
|
Control
|
Thymol
|
Carvacrol
|
|
0
|
3.34
|
1.07c
|
2.96a
|
1.73b
|
|
1
|
7.79
|
0.42c
|
3.14a
|
2.12b
|
|
3
|
13.25
|
0.84c
|
5.48a
|
1.85b
|
|
4
|
9.18
|
0.21c
|
8.57a
|
6.89b
|
|
8
|
21.38
|
0.16b
|
4.81a
|
4.72a
|
|
14
|
8.65
|
0.08c
|
5.22a
|
3.99b
|
Table 7: Relative expression of pathogen-related protein gene PR6 in Gemmieza-11 wheat genotype at different days after inoculation (DAI) with P. striiformis at seedling stage in comparison with Yr15 resistance monogenic line.
|
Days after inoculation
|
Yr15 gene
|
Gemmieza-11
|
|
Control
|
Thymol
|
Carvacrol
|
|
0
|
1.26
|
0.06c
|
1.76a
|
0.44b
|
|
1
|
6.39
|
0.12c
|
2.59a
|
1.60b
|
|
3
|
13.46
|
0.27b
|
10.13a
|
0.25b
|
|
4
|
14.12
|
2.43b
|
6.38a
|
2.49b
|
|
8
|
5.45
|
1.40c
|
11.14a
|
3.09b
|
|
14
|
22.06
|
0.54c
|
11.98a
|
3.52b
|
Table 8: Relative expression of pathogen-related protein gene PR8 in Gemmieza-11 wheat genotype at different days after inoculation (DAI) with P. striiformis at seedling stage in comparison with Yr15 resistance monogenic line.
|
Days after inoculation
|
Yr15 gene
|
Gemmieza-11
|
|
Control
|
Thymol
|
Carvacrol
|
|
0
|
3.00
|
1.16b
|
2.82a
|
1.34b
|
|
1
|
12.31
|
0.09c
|
5.33a
|
0.56b
|
|
3
|
7.97
|
4.13b
|
11.41a
|
4.99b
|
|
4
|
3.16
|
0.29c
|
2.91a
|
1.35b
|
|
8
|
3.83
|
0.24c
|
1.53b
|
2.42a
|
|
14
|
4.29
|
1.07b
|
3.19a
|
0.57c
|
Table 9: Relative expression of pathogen-related protein gene PR12 in Gemmieza-11 wheat genotype at different days after inoculation (DAI) with P. striiformis at seedling stage in comparison with Yr15 resistance monogenic line.
|
Days after inoculation
|
Yr15 gene
|
Gemmieza-11
|
|
Control
|
Thymol
|
Carvacrol
|
|
0
|
5.26
|
0.31c
|
3.01a
|
0.47b
|
|
1
|
7.18
|
1.53b
|
6.51a
|
0.38c
|
|
3
|
10.24
|
3.56b
|
7.08a
|
2.94c
|
|
4
|
11.89
|
3.89b
|
9.17a
|
4.11b
|
|
8
|
9.31
|
0.72c
|
5.66a
|
3.02b
|
|
14
|
32.04
|
5.40c
|
7.99b
|
9.65a
|
Conclusion: Stripe rust resistance-related genes were investigated in 19 commercial wheat cultivars using four DNA markers in this research. Genetic differences of various genes were observed in the genotypes evaluated. Stripe rust resistance in wheat genotypes may be improved by incorporating favorable alleles from this study. P. striiformis resistance in wheat was achieved using 100 mg/L thymol and carvacrol. According to gene expression data, low-molecular weight proteins, such as thymol and carvacrol, have a role in the signaling pathway of the resistance in the early stages of stripe rust disease via the interaction of multiple PR genes.
REFERENCES
- Ali, S., J. Rodriguez-Algaba, T. Thach, C.K. Sørensen, J.G. Hansen, P. Lassen, K. Nazari, D.P. Hodson, A.F. Justesen, and M.S. Hovmøller (2017). Yellow rust epidemics worldwide were caused by pathogen races from divergent genetic lineages. Front. Plant Sci. 8: 1057. doi: 10.3389/fpls.2017.0105.
- Afshari, F. (2008) Prevalent pathotypes of Puccinia striiformis sp. tritici in Iran. J Agric Sci Technol 10: 67-78.
- Banani, H., L. Olivieri, K. Santoro, A. Garibaldi, M.L. Gullino and D. Spadaro (2018). Thyme and savory essential oil efficacy and induction of resistance against Botrytis cinerea through priming of defense responses in apple. Foods 7: 11. doi:10.3390/foods7020011.
- Bariana, H.S., G.N. Brown, N.U. Ahmed, S. Khatkar, and R.L. Conner et al. (2002). Characterisation of Triticum vavilovii-derived stripe rust resistance using genetic, cytogenetic and molecular analyses and its marker-assisted selection. Theor Appl Genet 104:315-320.
- Burt, S.A., S.J.M. Adolfse, D.S.A. Ahad, M.H.G. Tersteeg-Zijderveld, B.G.M. Jongerius-Gortemaker, J.A. Post, H. Brüggemann, and R.R. Santos (2016). Cinnamaldehyde, carvacrol and organic acids affect gene expression of selected oxidative stress and inflammation markers in IPEC-J2 cells exposed to Salmonella typhimurium. Phytother. Res. 30: 1988–2000.
- Bux, H., A. Rasheed, M.A. Siyal, A.G. Kazi, and A.A. Napar et al. (2012). An overview of stripe rust of wheat (Puccinia striiformis sp. tritici) in Pakistan. Arch Phytopathol Plant Protect 45: 2278-2289.
- Cao, Z.J., M.N. Wang, and J.X. Jing (2001). Progress in the inheritance of resistance to stripe rust in wheat. Triticeae Crops 21: 80-83.
- Chen, X. (2020). Pathogens which threaten food security Puccinia striiformis the wheat stripe rust pathogen. Food Security 12: 239–251. https://doi.org/10.1007/ s12571- 020-01016-z.
- Chen, X.M. (2005). Epidemiology and control of stripe rust (Puccinia striiformis sp. tritici) on wheat. Canadian Journal of Plant Pathology 27: 314-337.
- Chen, X., M.A. Soria, G. Yan, J. Sun, and J. Dubcovsky (2003). Development of sequence tagged site and cleaved amplified polymorphic sequence markers for wheat stripe rust resistance gene Crop Sci 43: 2058-2064.
- Dai, L., D. Wang, X. Xie, C. Zhang, X. Wang, Y. Xu, Y. Wang, and Zhang (2016). The novel gene vpPR4-1 from Vitis pseudoreticulata increases powdery mildew resistance in transgenic Vitis viniferaL. front. Plant Sci 7: 695.
- Elsharkawy, M.M., M. Shimizu, H. Takahashi, and M. Hyakumachi (2012) Induction of systemic resistance against Cucumber mosaic virus by Penicillium simplicissimum GP17-2 in Arabidopsis and tobacco. Plant Pathol. 61:964–976.
- Esmail, S.M., A.A. Aboulila, and D. Abd El-Moneim (2020). Variation in several pathogenesis - related (PR) protein genes in wheat (Triticum aestivum) involved in defense against Puccinia striiformis sp. tritici. Physiological and Molecular Plant Pathology 112: 101545.
- FAO (2017). FAOSTAT. Rome: FAO available at: http://faostat.fao.org.Accessed 8 Nov 2019.
- Farrakh, S., M. Wang, and X. Chen (2018). Pathogenesis-related protein genes involved in race-specific allstage resistance and non-race specific high-temperature adult plant resistance to Puccinia striiformis sp. tritici in wheat. Journal of Integrative Agriculture 17(11): 2478-2491.
- German S.E., and J.A. Kolmer (1992). Effect of gene Lr34 on the enhancement of resistance to leaf rust of wheat. Theor Appl Genet 84: 97-105.
- Giovagnoni, G., B. Rossi, B. Tugnoli, F. Ghiselli, A. Bonetti, A. Piva, and E. Grilli (2020). Thymol and carvacrol downregulate the expression of Salmonella typhimurium virulence genes during an in vitro infection on Caco-2 cells. Microorganisms 8: 862 doi:10.3390/microorganisms8060862.
- Green, T.R., and C.A. Ryan (1972). Wound-induced proteinase inhibitor in plant leaves: a possible defense mechanism against insects. Science 175: 776-777
- Inamuco, J., A. Veenendaal, S. Burt, J. Post, J. Bokhoven, H. Haagsman, and E. Veldhuizen (2011). Sub-lethal levels of carvacrol reduce Salmonella typhimurium motility and invasion of porcine epithelial cells. Vet. Microbiol. 157: 200–207.
- Koebner, R.M., and R.W. Summers (2003). 21st century wheat breeding: Plot selection or plate detection? Trends Biotechnol 21: 59-63.
- Lagudah, E.S., S.G. Krattinger, S. Herrera-Foessel, R.P. Singh, and J. Huerta-Espino et al. (2009). Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor Appl Genet 119: 889-898.
- Livak, K.J., and T.D. Schmittgen (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25(4):402-8. doi: 10.1006/meth.2001.1262. PMID: 11846609.
- Metraux, J.P., L. Streit, and T. Staub (1988). A pathogenesis-related protein in cucumber is a chitinase Physiol. Mol. Plant Pathol. 33: 1-9.
- Mukhtar, S., M.A. Khan, B.A. Paddar, A. Anjum, G. Zaffar, S.A. Mir, S. Naseer, M.A. Bhat and Kamaluddin (2015). Molecular characterization of wheat germplasm for stripe rust resistance genes (Yr5, Yr10, Yr15 & Yr18) and identification of candidate lines for stripe rust breeding in Kashmir. Indian Journal of Biotechnology 14: 241-248.
- Murphy L.R., D. Santra, K. Kidwell, G. Yan, and X. Chen (2009). Linkage maps of wheat stripe rust resistance genes Yr5 and Yr15 for use in marker-assisted selection. Crop Sci 49: 1786-1790.
- Murray, M.G., and F. Thompson (1980). Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8(19): 4321-4325. doi: 10.1093/nar/8.19.4321. PMID: 7433111; PMCID: PMC324241.
- Návarová, H., F. Bernsdorff, A.C. Döring, and J. Zeier (2012). Pipecolic acid, an endogenous Mediator of defense amplification and priming, is a critical regulator of inducible plant immunity. Plant Cell 24: 5123-5141.
- Omara, R.I., G.A. El-Kot, F.M. Fadel, K.A.A. Abdelaal, and E.M. Saleh (2019). Efficacy of certain bioagents on patho-physiological characters of wheat plants under wheat leaf rust stress. Physiol Mol Plant Pathol 106:102–108
- Omara, R.I., S.M. Kamel, Y.M. Hafez, and S.Z. Morsy (2015) Role of non-traditional control treatments in inducing resistance against wheat leaf rust caused by Puccinia triticina. Egyp J Biol Pest Cont 25(2):335–344
- Omara, R.I., Shahin, A.A., Ahmed, S.M., Mostafa, Y.S., Alamri, S.A., Hashem, M., and M.M Elsharkawy (2021). Wheat resistance to stripe and leaf rusts conferred by introgression of slow rusting resistance genes. J. Fungi 7: 622 https://doi.org/10.3390/jof7080622.
- Parlevliet, J.E., and H.J. Kuiper (1977). Partial resistance of barley to leaf rust. Puccinia hordei IV Effect of cultivars and development stage on infection frequency. Euphytica 26: 249–255.
- Parlevliet, J.E. (1975). Partial resistance of barley to leaf rust. Puccinia hordei I effect of cultivars and development stage on latent period. Euphytica 24: 21–27.
- Peterson, R.F., A.B. Compbell, and A.E. Hamah (1948). A diagrammatic scale for estimating rust intensity on leaves and stems of cereal. Can. J. Res. 60: 496-500.
- Pourkhorshid, Z., A. Dadkhodaie, A. Niazi, B. Heidari, and E. Ebrahimi (2014). Identification of wheat stripe rust resistance genes in Iranian wheat cultivars using molecular markers. Annual Research & Review in Biology 4(17): 2766-2778.
- Roelfs, A.P., R.P. Singh, and E.E. Saari (1992). Rust diseases of wheat: Concepts and Methods of Diseases Management. Mexico, D. F.CIMMYT, 81.pp.
- Sadravi, M. (2008). Important diseases of crop plants. Mashhad. Mashhad University Publications, Iran. Persian.
- Salman, E.K., E.S. Badr, K.E. Ghoniem, A.A. Aboulila, and A.A. Emeran (2021). Role of silymarin induced rice immunity against blast pathogen Magnaporthe oryzae through regulation of resistance genes expression. Physiological and Molecular Plant Pathology 115: 101678.
- Scholtz, J.J., and B. Visser (2013). Reference gene selection for qPCR gene expression analysis of rust-infected wheat. Physiological and Molecular Plant Pathology 81: 22-25.
- Sels, J., J. Mathys, B.M. De Coninck, B.P. Cammue, and M.F. De Bolle (2008). Plant pathogenesis- related (PR) proteins: a focus on PR peptides. Plant Physiol. Biochem 46: 941-950
- Sharma, I. (2012). Disease resistance in wheat. Punjab Agricultural University, India.
- Simons, M.D., J.W. Martens, R.I.H. McKenzie, I. Nishiyama, K. Sadanaga, J. Sebesta and H. Thomas (1978). Oats: A standardized system of nomenclature for genes and chromosomes and catalogue of genes governing characters U.S., Dept. Agric., Handbook. 509: 1-40.
- Soković, M.D., J. Vukojević, P.D. Marin, D.D. Brkić, V. Vajs, and L.J.L.D. van Griensven (2009). Chemical composition of essential oils of Thymus and Mentha species and their antifungal activities. Molecules 14: 238–249.
- Spadaro, D., and M.L. Gullino (2014). Use of Essential Oils to Control Postharvest Rots on Pome and Stone Fruit. In Post-Harvest Pathology; Prusky, D., Gullino, M.L., Eds.; Springer International Publishing: Cham, Switzerland 101–110.
- Tervet, I. and R. C. Cassel (1951). The use of cyclone separation in race identification of cereal rusts. Phytopathology 41.282-285.
- Terras, F.R., K. Eggermont, V. Kovaleva, N.V. Raikhel, R.W. Osborn, A. Kester, S.B. Rees, J. Vanderleyden, B.P. Cammue, and W.F. Broekaert (1995). Small cysteine-rich antifungal proteins from radish: their role in host defense. Plant Cell 7: 573-588
- Van Loon, L.C. (1982). Regulation of changes in proteins and enzymes associated with active Defense against virus infection R.K.S. Wood (Ed.), Active Defense Mechanisms in Plants, Plenum Press, New York, pp. 247-273
- Wang L.F., J.X. Ma, R.H. Zhou, X.M. Wang, and J.Z. Jia (2002). Molecular tagging of the yellow rust resistance gene Yr10 in common wheat, P.I.178383 (Triticum aestivum ), Euphytica 124: 71-73.
- William, H.M., R. Trethowan, and E.M. Crosby-Galvan (2007). Wheat breeding assisted by markers: CIMMYT’s experience, Euphytica 157: 307-319.
- Yan, G.P., X.M. Chen, R.F. Line, and C.R. Wellings (2003). Resistance gene-analog polymorphism markers co-segregating with the Yr5 gene for resistance to wheat stripe rust. Theor Appl Genet 106:636-643.
- Yu, G., H. Ma, Z. Xu, L. Ren, and M. Zhou, et al. (2004). Cloning a DNA marker associated to wheat scab resistance. J Appl Genet 45: 17-25.
- Zhang, H., C. Wang, Y. Cheng, X. Chen, Q. Han, L. Huang, G. Wei, Z. Kang (2012). Histological and cytological characterization of adult plant resistance to wheat stripe rust. Plant Cell Reporter 31: 2121–2137.
- Zhou, X.L., D.J. Han, X.M. Chen, H.L. Gou, and S.J. Guo, et al. (2014). Characterization and molecular mapping of stripe rust resistance gene Yr61 in winter wheat cultivar Pindong 34. Theor Appl Genet 127: 935-945.
|





