EFFECT ON GENE EXPRESSION PROFILE OF HNRNPK KNOCKDOWN IN MOUSE GC-1SPG CELLS
H. Xu1,2, Q. Han1, Y. Wang1, N. Chen1, X. Cheng1,2, S. Yu1, C. Li 1,2, P. Zhang 1,2 and Y. Xu 1,2 *
1College of Life Science, Xinyang Normal University, Xinyang 464000, China;
2Institute for Conservation and Utilization of Agro-bioresources in Dabie Mountain, Xinyang Normal University, Xinyang 464000, China
*Corresponding author’s Email: xyj@xynu.edu.cn
ABSTRACT
HnRNPK is a multifunctional RNA binding protein. Our previous studies have found that it plays a key role in the survival of spermatogonia GC-1spg, but the mechanism is not clear. To reveal the functional mechanism of hnRNPK in spermatogonia, the expression profiles of hnRNPK knockdown and control groupGC-1spg cells were analyzed by RNA-seq. A total of 1453 differentially expressed genes were obtained, of which 604 genes were up-regulated and 849 genes were down-regulated in GC-1spg cells after hnRNPK knockdown. 24 genes were randomly selected from the differentially expressed genes for qRT-PCR verification. Pearson correlation analysis showed that r = 0.916 (P < 0.01), which indicated that there was a good correlation between the results of RNA-Seq and qRT-PCR. GO functional enrichment analysis showed that the biological process of differentially expressed genes was mainly related to cell proliferation, differentiation, apoptosis and other life activities, and then 17 genes related to proliferation and apoptosis were selected for further verification. The results showed that hnRNPK could affect the survival of GC-1spg cells by regulating proliferation and apoptosis-related genes. KEGG pathways enrichment analysis showed that GC-1spg cells mainly occurred cell-molecule interaction and activation of related signal transduction pathway after hnRNPK knockdown. The results will provide an important basis and clue for revealing the key targets and molecular regulation mechanism in the process of spermatogonia survival, and also provide new ideas to understand the problem of male sterility.
Keywords:Spermatogenesis; Spermatogonia; hnRNPK; RNAi; RNA-Seq
http://doi.org/10.36899/JAPS.2022.2.0452
Published first online August 13. 2021
Introduction
Mammalian spermatogenesis is a complex and precisely regulated developmental process, which is related to many types of germ cells, such as spermatogonia, spermatocytes and spermatozoa. It is well known that spermatogonia play crucial roles during this process. According to the morphological characteristics of spermatogonia, which can be divided into three types: A, In and B. Among them, As, Apr and Aal, are usually recognized as A spermatogonia, but only As are called spermatogonial stem cells (SSCs), while B spermatogonia are committed to differentiation,which finally divide to primary spermatocytes (Hermann et al. 2010; Wen et al. 2019; Lord et al. 2020). As constantly updates itself to produce more As, some As and then divide into Apr, through division and divides into Aal, and then differentiates into A1, A2, A3, A4, In and B. B spermatogonia further differentiate into spermatocytes, then form spermatids, and finally produce sperm (Nakagawa et al. 2007; Ni et al. 2019). Spermatogonia are very important and complex in the whole stage of spermatogenesis, and revealing the key regulators and mechanisms in the process of proliferation and survival of spermatogonia has always been a research hotspot in reproductive biology.
Heterogeneous ribonucleoproteins (hnRNP) are a family of RNA binding proteins with at least 20 protein members, which are successively named A to U. Among them, hnRNPK is a widely studied member, which can interact with RNA, DNA and proteins, participate in the regulation of transcription and translation, regulate RNA splicing and cell signal transduction (Han et al. 2010; Xu et al. 2019). It is a multifunctional protein, which is closely related to the development of reproductive system (Lee et al. 2004; Xu et al. 2015), hematopoiesis (Ostareck-Lederer et al. 2012), nervous system development (Hutchins et al. 2013), myoblast differentiation (Xu et al. 2018b) and tumorigenesis (Gallardo et al. 2015). However, there are few studies on the relationship between hnRNPK and spermatogenesis. We analyzed the spatio-temporal expression profile of hnRNPK in pigs and rats, and found that the expression of hnRNPK in pig and rat testes were significantly higher than that in other tissues, and had a high expression abundance in the early stage of spermatogenesis, suggesting that hnRNPK may play an important role in the spermatogonia stage (Xu et al. 2018a). Lee et al. have reported that Rbm can interact with hnRNPK and another sex-specific splicing activator Tra2α to regulate the specific splicing of RNA during spermatogenesis, in which hnRNPK may act as an anchored protein, providing a platform for Rbm and Tra2α to promote the interaction between protein complexes (Lee et al. 2004). GC-1spg cell line is a widely used spermatogonia model,which was immortalized by pSV3-neo transfection into 10-day-old male mice and the characteristics of the cell line were between type B spermatogonia and leptotene spermatocytes (Hofmann et al. 1992). Therefore, we carried out a preliminary study on the function of hnRNPK in GC-1spg cells through RNA interference. The results showed that the number of GC-1spg cells decreased significantly, and there were some floating cells after knocking down hnRNPK (Xu et al. 2017), but the mechanism is not clear. In this study, high-throughput transcriptome sequencing was used to analyze the gene expression profiles in hnRNPK knockdown and control GC-1spg cells, to screen differentially expressed genes, and to enrich their functions and pathways, so as to further explore the possible downstream target genes and pathways regulated by hnRNPK, and provide a basis for revealing the key targets and molecular mechanisms in the process of spermatogonia survival. The research results will provide a new perspective for solving the problem of male sterility.
Materials and methods
Cell culture: GC-1spg cells were cultured in DMEM medium supplemented with 10% fetal bovine serum, 1 mM sodium pyruvate, 2 mM L-glutamine and 100 mM non-essential amino acids, and cultured in an incubator containing 5% CO2 at 37 ℃.
siRNA screening and cell transfection: Three pairs of siRNA were designed and synthesized according to the coding region of mouse hnRNPK gene and verified by previous screening experiments (Xu et al. 2017). Finally, the siRNA-hnRNPK with better interference effect (interference efficiency > 80%), was selected for follow-up experiments. The cells were inoculated in a six-well cell plate, waiting for the density to reach 40-60%, ready for transfection. The final concentration of siRNA was added to 100 nM, siRNA-Mate12 μL, mixed gently, and incubated at room temperature for 15 min, to form siRNA-transfection reagent complex. At the end of incubation, the complex was added drop by drop into each well, gently mixed, and RNA was extracted after 48 h.
Cell viability assay: For the cell viability assay, cells were seeded in 96-well plates with 6000 cells per well and measured at 3 different times (transfection for 0 h, 24 h, 48 h) by MTT assay according to the manufacturer’s protocol. Briefly, we added 20 μL MTT (4mg/mL) to each well and incubated them at 37 ℃ for 4 h. Then, we discarded the supernatants and dissolved the remains with 150 μL DMSO per well, and the absorbance was measured at 490 nm using a SpectraMax M5 unit (Molecular Devices). The graph was plotted using GraphPad Prism 8 software.
Transcriptome sequencing: The cell transfection experiment was repeated three times, and the cells were collected for RNA extraction to obtain RNA samples from knockdown group (siRNA-hnRNPK) and control group (siRNA-NC). After passing through the quality inspection, the RNA-Seq library was constructed and sequenced, and the work was provided by Suzhou Jisai Gene sequencing Technology Co., Ltd. Through the paired-end sequencing of Solexa RNA, a large number of sample data of 2 to 100 bp were obtained. After the quality preprocessing of the original data, the low-quality fragments and joint sequences were removed, and the preprocessed Reads was compared to the reference genome using TopHat alignment software. Through Cufflinks software, the expression level of each gene was estimated by fragments per kilobase of transcript per million mapped reads (FPKM) algorithm, and then the differentially expressed genes between the two groups were analyzed according to the screening criteria of log2Fold Change ≥ 1, q-value < 0.05. Using GO seq software, the calculated P-value is corrected by Bonferroni, and the threshold value of corrected P-value ≤ 0.05. The GO term satisfying this condition is defined as GO term significantly enriched in differentially expressed genes. The main biological functions of differentially expressed genes can be revealed by GO functional significance enrichment analysis. Pathway enrichment analysis was carried out by using Biocarta and KEGG databases to screen differential genes in different pathways, so as to reveal the cellular signal pathways that these differentially expressed genes may participate in.
RNA isolation, cDNA synthesis and qRT-PCR analysis: Total-RNA extractions were carried out from hnRNPK knockdown and control group GC-1spg cells using TRIzol (Takara, Japan). The first strand cDNA was synthesized by reverse transcription of 1-5 μg total RNA using M-MLV Reverse Transcriptase and Oligo (dT)15 (Invitrogen, USA). qRT-PCR was performed on the ABI 7300 (ABI, USA) using the GoTaq® qPCR Master Mix (Promega, USA). Gene-specific primers were used to determine the relative expression levels of the detected genes according to the standard curve method. The primer sequences are listed in Supplemental Table 1. Gene expression levels were quantified relatively to the expression of the mouse Gapdh gene, by employing an optimized comparative Ct (ΔΔCt) value method. The expression level was calculated as 2^ (−ΔΔCt) to compare the relative expression, and SPSS17.0 software was used for statistical analysis, one-way ANOVA was conducted to identify differentially expressed genes, and P < 0.05 was considered as significant.
Verification of differentially expressed genes: Through qRT-PCR technology, 24 differentially expressed genes were randomly selected for verification (Supplemental Table 1). The relative differential fold changes between RNA-Seq and qRT-PCR between the two groups were analyzed by Pearson correlation analysis with SPSS17.0 software to verify whether the results of RNA-Seq were reliable.
Results
GC-1spg cell proliferation is impeded after hnRNPK knockdown and RNA quality detection: The number of GC-1spg cells decreased significantly after knocking down the expression of hnRNPK, and there were also some floating cells (Xu et al. 2017).To verify this result, MTT assay was conducted. Compared with the control, the obtained MTT assay data also showed that the number of living GC-1spg cells with hnRNPK gene knockdown decreased significantly, indicating that hnRNPKknockdown impeded GC-1spg cell proliferation (P<0.05) (Figure 1).
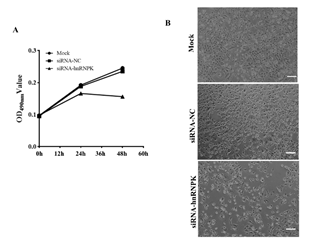
Figure 1. GC-1spg cell proliferation is impeded after hnRNPK knockdown. (A) MTT detection for GC-1spg cell proliferation after hnRNPK knockdown. (B) GC-1spg cell phenotypic change after hnRNPK knockdown, Bar = 50 μm.
The mixed RNA fromGC-1spg cells obtained by three independent siRNA transfection experiments was detected and analyzed by Agilent 2000, it was found that the total RNA of knockdown and control groups was intact and non-degraded, RNA content > 5 ug, RIN > 9. The quality of RNA samples met the requirements of Illumina sequencing, and the detailed quality detection information was in Supplemental Table 2.
Sequencing data quality control and reference genome alignment: After pre-processing and removing low-quality fragments and joint sequences, a total of 140 million effective sequencing data were obtained, with a data volume of 14 G and an average length of 97.40 bp. The amount of data sequenced by the two groups was more than 5 G, which could meet the needs of subsequent data analysis (Supplemental Table 3). After data comparison, 23337 genes, 30455 transcripts and 24759 proteins were obtained on 22 chromosomes of chr1-19, X, Y and MT. The proportion of reads on the reference genome of the two groups is about 93%, of which the proportion of pairwise reads alignment is about 91%, a single reads alignment is about 91%, and more than 87% of the reads is aligned to the exons of known or predicted genes (Supplemental Table 4).
Gene expression analysis: The high-throughput sequencing results showed that there were total 23337 genes in mice, and 13886 genes from the libraries of knockdown group and control group, including 13206 in knockdown group and 13083 in control group. Among them, the FPKM value is 0.1-3.75 belongs to a low-expression gene, 3.75-15 is a moderate-expression gene, and more than 15 is a high-expression gene (Mortazavi et al. 2008). The FPKM values of genes in the two groups were mainly distributed in the range of 3.57-60, accounting for 57% (Supplemental Figure 1). There were 65 and 38 genes with FPKM values greater than 1000 in siRNA-NC and siRNA-hnRNPK, of which 35 genes were same, including Eef1a1, Cxcl10, Rpl19, Eif6, Snhg6, Hspa8 and other genes.
Analysis of differential gene expression: Furthermore, the difference of gene expression between the knockdown group and the control group was analyzed, and the selection standard was as follows: log2Fold Change ≥ 1, q-value < 0.05. The results showed that among the 1453 differentially expressed genes, 604 genes were up-regulated in knockdown group, accounting for 41.57%, and 849 genes were down-regulated, accounting for 58.43% (Supplemental Table 5). Among the differentially expressed genes, 1316 (90.57%) genes encoded proteins, and only 137 (9.43%) belonged to non-coding RNA or pseudogenes and unknown types. There were 32 and 51 highly expressed differential genes in siRNA-hnRNPK and siRNA-NC groups respectively, and 16 in both groups (Supplemental Figure 2). Among them, Rplp0, Bre, Dbi and other highly expressed genes were significantly down-regulated after hnRNPK knockdown, while Ccl2, Rsad2, Cxcl10 and other highly expressed genes were significantly up-regulated.

Figure 2.The reliability of RNA-Seq expression results were identified by qRT-PCR.
In addition, among the differentially expressed genes, such as Nanog, Itgb1, SFN, Ptges, Eaf2 are proliferation related genes, and Casp3, Xiap, Spa17, Rap1GAP1, Sirt1 are apoptosis related genes. Comprehensive analysis of their FPKM values and fold changes has found that the changes of the above genes were very significant after hnRNPK knockdown.
Verification of differentially expressed genes: In order to verify the reliability of RNA-seq results, 24 genes were randomly selected from the differentially expressed genes for further analysis (Supplemental Table 2). Because the gene expression obtained by RNA-Seq and qRT-PCR are based on different calculation methods and standards, the two results cannot be compared directly. Therefore, we compared the differential fold changes obtained by RNA-Seq and qRT-PCR between the two groups. The results showed that the expression patterns of 24 randomly selected differential genes were same as those obtained by RNA-Seq (Figure 2). Furthermore, Pearson correlation analysis of the differential fold changes from the two methods was carried out by SPSS17.0 software, obtained r = 0.916, P < 0.01, the results showed that there was a very significant correlation between RNA-Seq and qRT-PCR, indicating that the results of RNA-Seq were reliable.
GO analysis of differentially expressed genes: GO analysis mainly clustered and analyzed the molecular functions, cellular components and biological processes of differentially expressed genes in order to screen the significantly enriched biological functions. The molecular functions of differentially expressed genes after hnRNPK knockdown in GC-1spg cells are mainly involved in nucleotide binding, protein binding, ion binding, receptor activity, receptor binding and regulatory activity. Cell components are mainly related to cellular content, cell surface, membrane, extracellular matrix, vesicle and envelope; biological processes are mainly related to development, transcriptional regulation, cell proliferation and differentiation, cell migration, immune response, signal transduction and metabolism, apoptosis and other life activities (Figure 3).

Figure 3. GO analysis of differentially expressed genes from siRNA-hnRNPK and siRNA-NC groups.
Pathway analysis of differentially expressed genes: Pathway analysis of differentially expressed genes was carried out by Biocarta and KEGG databases, and Biocarta analysis showed that these differentially expressed genes were involved in cell-surface molecular pathways, cell differentiation pathways and cytokine-mediated hematopoiesis, and KEGG results suggested that the main enrichment pathways were ligand-receptor interaction, cytokine receptor interaction, cell molecular adhesion, hematopoietic cell lines and calcium signal transduction (Figure 4). In short, the Pathway enrichment analysis of the two databases showed that cell-molecule, molecular-molecular interaction and activation of related signal transduction pathways are mainly occurred in hnRNPK knockdown GC-1spg cells.
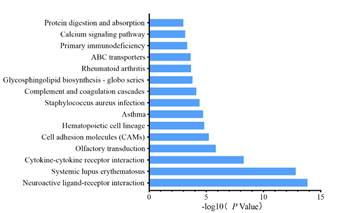
Figure 4.KEGG analysis of differentially expressed genes from siRNA-hnRNPK and siRNA-NC groups.
Further analysis of proliferation and apoptosis related genes: Further analysis of differentially expressed genes showed that there were significant changes in proliferation and apoptosis related genes in hnRNPK knockdown GC-1spg cells. Therefore, we selected Cdkn2a, Itgb1, Nanog, Igfbp6, Lst1, Oaz1 and Icmt proliferation genes regulate positively; E2f7, Spry1, Ptpn6, Bcam, Fpgt, Nos2 proliferation genes regulate negatively and Casp3, Xiap, Rap1GAP1, Sirt1 apoptosis related genes for further qRT-PCR analysis. The results showed that after hnRNPK knockdown, the expression of proliferation genes regulate positively was down-regulated, while the expression of proliferation genes regulate negatively was up-regulated, the expression of anti-apoptosis gene was down-regulated, and the expression of pro-apoptotic genes was up-regulated. It is suggested that hnRNPK may affect the survival of GC-1spg cells by regulating proliferation and apoptosis-related genes (Figure 5).
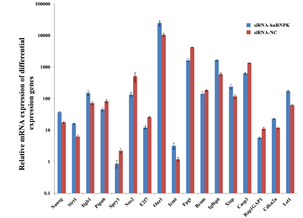
Figure 5.Expression of proliferation and apoptosis related gene verified by qRT-PCR.
Discussion and conclusion
Transcriptome sequencing is an important tool for in-depth study of transcriptome, which is to sequence the reverse transcribed cDNA of RNA in tissues or cells by high-throughput sequencing technology (Stark et al. 2019). In order to reveal the genes and signal pathway regulated by hnRNPK during spermatogonia survival, we used RNA-seq to analyze the expression profiles of hnRNPK knockdown and control group GC1-spg cells. 1453 differentially expressed genes were screened, and the GO function of these genes were further enriched. It was found that 23 genes regulated cell proliferation and 15 genes were related to apoptosis. Among the 15 genes involved in apoptosis, Xiap (X-linked inhibitor of apoptosis protein) is an anti-apoptotic gene with low expression in knockdown group, which can regulate apoptosis by directly inhibiting the activity of caspases (Pop et al. 2009; Hanahan et al. 2011; Nielsen et al. 2017). However, hnRNPK can regulate the expression of Xiap gene at the transcriptional level (Xiao et al. 2013; Chen et al. 2017). After the decreased expression of hnRNPK in GC-1spg spermatogonia, it may inhibit the transcriptional activity of Xiap, thus down-regulate the expression of Xiap, weaken the inhibition of casp3 and casp7 enzyme activity, and then induce the apoptosis of GC-1spg cells. The highly expressed casp2 and casp3 encode two important signal transduction proteins in the apoptosis pathway. Studies have shown that casp3 gene can induce apoptosis in insect Sf9 cells. After removing casp3 from the extract of apoptotic cells, the ability to induce apoptosis was lost, and after the addition of purified casp3, the ability to induce apoptosis was restored (Porter et al. 1999; Wright et al. 1999). Previous studies have also shown that casp3 is expressed in both mammalian spermatogonia and spermatocytes. Apoptosis of spermatogenic cells depends on the increase of casp3 expression and activity, which can promote spermatogonia apoptosis (Kim et al. 2001; Omezzine et al. 2003; Pasha et al. 2016). Casp2 may also be involved in apoptosis, but it must be activated by casp3 (Li et al. 1997). In addition to the high expression of these pro-apoptotic genes, we also found that many genes needed for anti-apoptosis or survival were down-regulated. For example, Rap1GAP1 is the upstream negative regulator ofRap1, while Rap1 mediates the activation of p38a and promotes cell apoptosis (Gutierrez-Uzquiza et al. 2010; Zhang et al. 2015). The protein encoded by the down-regulated gene Sirt1, is a member of the histone deacetylase family, which plays an anti-apoptotic role through p53/bax and NF-κB/PGC-1α pathways (Vaziri et al. 2001; Yeung et al. 2004; Morigi et al. 2018), and may play an important role in spermatogenesis and germ cell development (Coussens et al. 2008; Tatone et al. 2018). Therefore, we speculate that the up-regulated pro-apoptotic genes and down-regulated anti-apoptotic genes will be important candidates for further study of the effect of hnRNPK on germ cell apoptosis.
Among the 23 genes involved in regulating cell proliferation, 13 positively regulatory genes were down-regulated in hnRNPK knockdown GC-1spg cells, which were Itgb1, Nanog, Fabp4, Fbxw4, Crip2, Figf, Fgf10, Il7, Oaz1, Il11, Tnfsf13, Fntb and Icmt; and the other 10 genes were up-regulated, which were Ptpn6, Csf1, Cd274, Icosl, E2f7, Irs2, Spry1, Tgfbr3, Serpine1 and Nos2 (Figure 6). Further analysis of these proliferation regulatory genes, we found that fold changes > 4 are Nanog, Itgb1 and Ptpn6.
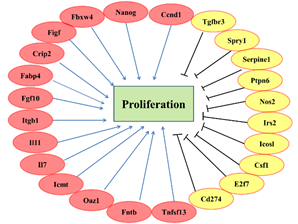
Figure 6.The differential expressed genes that may be involved in GC-1spg proliferation regulation.
Previous studies have shown that the most down-regulated gene Nanog, is expressed in both embryonic stem cells and embryonic germ cells, and when embryonic stem cells differentiate or male germ cells enter the mitotic arrest phase, the expression is down-regulated (Chambers et al. 2003; Hart et al. 2004; Yamaguchi et al. 2005). Transcription factors T and FOXD3 in embryonic stem cells can up-regulate the expression of Nanog, promote cells to enter S phase and promote cell proliferation (Pan et al. 2006; Evans et al. 2012). Our study found that the expression of Nanog was significantly down-regulated, and transcription factor T and FOXD family members FOXD1 and FOXD2 were also down-regulated in hnRNPK knockdown GC-1spg cells. These results suggest that Nanog may promote spermatogonia proliferation and self-renewal by interacting with a variety of transcription factors (Saunders et al. 2013).
The integrin protein family includes two subtypes of alpha and beta, which mainly mediate cell migration, cell adhesion and immune response and other life activities (Brakebusch et al. 2002; Grose et al. 2002; Kaemmerer et al. 2015). Itgb1, which belongs to beta subtype, is an important adhesion molecule, which can promote the secretion of matrix metalloproteinases (MMP) to degrade extracellular matrix, release homogenous cell bondage and promote cell migration. Studies have shown that Itgb1 promotes cancer cell proliferation by up-regulating the cell cycle regulatory protein Ccnd1(Sales et al. 2014; Song et al. 2014). These reports are consistent with our results that the expression of Itgb1 was significantly down-regulated in hnRNPK knockdown GC-1spg cells, and the expression of Mmp family genes Mmp17, Mmp10, Mmp13 and Mmp11 were also significantly down-regulated. These results suggest that hnRNPK may regulate spermatogonia proliferation through Itgb as a key factor.
To sum up, hnRNPK is a key regulator in the process of spermatogonia survival. By analyzing the gene expression profile of hnRNPK knockdown and control GC-1spg cells through RNA-Seq, screening differential genes and enriching their functions and pathways, we found that some factors related to cell proliferation and apoptosis may play irreplaceable roles, but the detailed mechanism needs to be further studied. Therefore, the revelation of the function and mechanism of hnRNPK in spermatogonia will help to explain the complex biological process of animal spermatogenesis and promote its wide application in the fields of animal reproduction, health and medicine.
Author contribution: Conceptualization, H. Xu and Y. Xu; experiments, H. Xu, Q. Han, Y. Wang, N. Chen; data curation and software, S. Yu and C. Li; writing-original draft preparation, H. Xu and Q. Han; writing-review and editing, H. Xu, X. Cheng, P. Zhang and Y. Xu.
Acknowledgments: This work was financially supported by the National Natural Science Foundation of China (31972537), and the Nanhu Scholars Program for Young Scholars of XYNU.
Conflicts of Interest: The Authors Declare No Conflict of Interest.
ReferenceS
- Brakebusch, C., S. Fillatreau, A. J. Potocnik, G. Bungartz, P. Wilhelm, M. Svensson, P. Kearney, H. Korner, D. Gray and R. Fassler (2002). Beta1 integrin is not essential for hematopoiesis but is necessary for the T cell-dependent IgM antibody response. Immunity. 16(3): 465-477.
- Chambers, I., D. Colby, M. Robertson, J. Nichols, S. Lee S. Tweedie and A. Smith (2003). Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell. 113(5): 643-655.
- Chen X., P. Gu, R. Xie, J. Han, H. Liu, B. Wang, W. Xie, W. Xie, G. Zhong, C. Chen, S. Xie, N. Jiang, T. Lin and J. Huang (2017). Heterogeneous nuclear ribonucleoprotein K is associated with poor prognosis and regulates proliferation and apoptosis in bladder cancer. J Cell Mol Med. 21(7): 1266-1279.
- Coussens M., J. Maresh, R. Yanagimachi, G. Maeda and R. Allsopp (2008). Sirt1 deficiency attenuates spermatogenesis and germ cell function. PLoS One. 3(2): e1571.
- Evans A., T. Faial, M. Gilchrist, T. Down, L. Vallier, R. Pedersen, F. Wardle and J. Smith (2012). Genomic targets of Brachyury (T) in differentiating mouse embryonic stem cells. PLoS One. 7(3): e33346.
- Gallardo M., H. Lee, X. Zhang, C. Bueso-Ramos, L. Pageon, M. McArthur, A. Multani, A. Nazha, T. Manshouri, J. Parker-Thornburg, I. Rapado, A. Quintas-Cardama, S. Kornblau, J. Martinez-Lopez and S. Post (2015). hnRNP K Is a Haploinsufficient Tumor Suppressor that Regulates Proliferation and Differentiation Programs in Hematologic Malignancies. Cancer Cell. 28(4): 486-499.
- Grose R., C. Hutter, W. Bloch, I. Thorey, F. Watt, R. Fassler, C. Brakebusch and S. Werner (2002). A crucial role of beta 1 integrins for keratinocyte migration in vitro and during cutaneous wound repair. Development. 129(9): 2303-2315.
- Gutierrez-Uzquiza A., M. Arechederra, I. Molina, R. Banos, V. Maia, M. Benito, C. Guerrero and A. Porras (2010). C3G down-regulates p38 MAPK activity in response to stress by Rap-1 independent mechanisms: involvement in cell death. Cell Signal. 22(3): 533-542.
- Han S., Y. Tang and R. Smith (2010). Functional diversity of the hnRNPs: past, present and perspectives. Biochem J. 430(3): 379-392.
- Hanahan D. and R. Weinberg (2011). Hallmarks of cancer: the next generation. Cell. 144(5): 646-674.
- Hart A., L. Hartley, M. Ibrahim and L. Robb (2004). Identification, cloning and expression analysis of the pluripotency promoting Nanog genes in mouse and human. Dev Dyn. 230(1): 187-198.
- Hermann B., M. Sukhwani, M. Hansel and K. Orwig (2010). Spermatogonial stem cells in higher primates: are there differences from those in rodents? Reproduction. 139(3): 479-493.
- Hofmann M., S. Narisawa, R. Hess and J. Millan (1992). Immortalization of germ cells and somatic testicular cells using the SV40 large T antigen. Exp Cell Res. 201(2): 417-435.
- Hutchins E. and B. Szaro (2013). c-Jun N-terminal kinase phosphorylation of heterogeneous nuclear ribonucleoprotein K regulates vertebrate axon outgrowth via a posttranscriptional mechanism.
- J Neurosci. 33(37): 14666-14680.
- Kaemmerer E., P. Kuhn, U. Schneider, T. Clahsen, M Jeon., C. Klaus, J. Andruszkow, M Harer., S. Ernst, A. Schippers, N. Wagner and N. Gassler (2015). Beta-7 integrin controls enterocyte migration in the small intestine. World J Gastroenterol. 21(6): 1759-1764.
- Kim J., S. Ghosh, A. Weil and B. Zirkin (2001). Caspase-3 and caspase-activated deoxyribonuclease are associated with testicular germ cell apoptosis resulting from reduced intratesticular testosterone. Endocrinology. 142(9): 3809-3816.
- Lee J., E. Kim, S. Jang and K. Rhee (2004). Tra2α and hnRNP K might be functional partners of Rbm for regulation of RNA processes during spermatogenesis. Animal Cells and Systems. 8(1): 65-70.
- Li H., L. Bergeron, V. Cryns, M. Pasternack, H. Zhu, L. Shi, A. Greenberg and J. Yuan (1997). Activation of caspase-2 in apoptosis. J Biol Chem. 272(34): 21010-21017.
- Lord T. and B. Nixon (2020). Metabolic Changes Accompanying Spermatogonial Stem Cell Differentiation. Dev Cell. 52(4): 399-411.
- Morigi M., L. Perico and A. Benigni (2018). Sirtuins in Renal Health and Disease. J Am Soc Nephrol. 29(7): 1799-1809.
- Mortazavi A., B. Williams, K. McCue, L. Schaeffer and B. Wold (2008). Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 5(7): 621-628.
- Nakagawa T., Y. Nabeshima and S. Yoshida (2007). Functional identification of the actual and potential stem cell compartments in mouse spermatogenesis. Dev Cell. 12(2): 195-206.
- Ni F., S. Hao and W. Yang (2019). Multiple signaling pathways in Sertoli cells: recent findings in spermatogenesis. Cell Death Dis. 10(8): 541.
- Nielsen O. and E. LaCasse (2017). How genetic testing can lead to targeted management of XIAP deficiency-related inflammatory bowel disease. Genet Med. 19(2): 133-143.
- Omezzine A., S. Chater, C. Mauduit, A. Florin, E. Tabone, F. Chuzel, R. Bars and M. Benahmed (2003). Long-term apoptotic cell death process with increased expression and activation of caspase-3 and -6 in adult rat germ cells exposed in utero to flutamide. Endocrinology. 144(2): 648-661.
- Ostareck-Lederer A. and D. Ostareck (2012). Precision mechanics with multifunctional tools: how hnRNP K and hnRNPs E1/E2 contribute to post-transcriptional control of gene expression in hematopoiesis. Curr Protein Pept Sci. 13(4): 391-400.
- Pan G., J. Li, Y. Zhou, H. Zheng and D. Pei (2006). A negative feedback loop of transcription factors that controls stem cell pluripotency and self-renewal. FASEB J. 20(10): 1730-1732.
- Pasha H., N. Rezk, S. Selim and D. Abd El Motteleb (2016). Therapeutic effect of spermatogonial stem cell on testicular damage caused by lead in rats. Gene. 592(1): 148-153.
- Pop C. and G. Salvesen (2009). Human caspases: activation, specificity, and regulation. J Biol Chem. 284(33): 21777-21781.
- Porter A. and R. Janicke (1999). Emerging roles of caspase-3 in apoptosis. Cell Death Differ. 6(2): 99-104.
- Sales K., F. Giudice, R. Castilho, F. Salles, C. Squarize, A. Abrahao and D. Pinto Jr (2014). Cyclin D1-induced proliferation is independent of beta-catenin in head and neck cancer. Oral Dis. 20(3): e42-48.
- Saunders A., F. Faiola and J. Wang (2013). Concise review: pursuing self-renewal and pluripotency with the stem cell factor Nanog. Stem Cells. 31(7): 1227-1236.
- Song J., J. Zhang, J. Wang, Z. Cao, J. Wang, X. Guo and W. Dong (2014). beta1 integrin modulates tumor growth and apoptosis of human colorectal cancer. Oncol Rep. 32(1): 302-308.
- Stark R., M. Grzelak and J. Hadfield (2019). RNA sequencing: the teenage years. Nat Rev Genet. 20(11):631–656.
- Tatone C., G. Di Emidio, A. Barbonetti, G. Carta, A. Luciano, S. Falone and F. Amicarelli (2018). Sirtuins in gamete biology and reproductive physiology: emerging roles and therapeutic potential in female and male infertility. Hum Reprod Update. 24(3): 267-289.
- Vaziri H., S. Dessain, E. Ng Eaton, S. Imai, R. Frye, T. Pandita, L. Guarente and R. Weinberg (2001). hSIR2(SIRT1) functions as an NAD-dependent p53 deacetylase. Cell. 107(2): 149-159.
- Wen L. and F. Tang (2019). Human Germline Cell Development: from the Perspective of Single-Cell Sequencing. Mol Cell. 76(2):320-328.
- Wright M., D. Han, L. Carter, S. Fields, S. Schwartz and D. Hockenbery (1999). Caspase-3 inhibits growth in Saccharomyces cerevisiae without causing cell death. FEBS Lett. 446(1): 9-14.
- Xiao Z., H. Ko, E. Goh, B. Wang and E. Ren (2013). hnRNP K suppresses apoptosis independent of p53 status by maintaining high levels of endogenous caspase inhibitors. Carcinogenesis. 34(7): 1458-1467.
- Xu H., R. Li, S. Wang, K. Zhang, P. Zhang, L. Chen and Y. Xu (2017). Screening of siRNA targeted for hnRNPK gene and it effecton the spermatogonium survival. Journal of Xinyang Normal University(Natural Science Edition). 30(4): 561-566.
- Xu H., Y. Xu, X. Liang and W. Zeng (2015). The biological function of heterogeneous nuclear ribonucleoprotein K (hnRNP K) and its roles in spermatogenisis. Journal of Agricultural Biotechnology. 23(5): 661-670.
- Xu H., P. Zhang, R. Li, Wu W., S. Wang and Y. Xu (2018a). Expression analysis of multifunctional RNA-binding protein hnRNP K during development of mammalian testis. Pol J Vet Sci. 21(2): 343-351.
- Xu Y., R. Li, K. Zhang, W. Wu, S. Wang, P. Zhang and H. Xu (2018b). The multifunctional RNA-binding protein hnRNPK is critical for the proliferation and differentiation of myoblasts. BMB Rep. 51(7): 350-355.
- Xu Y., W. Wu, Q. Han, Y. Wang, C. Li, P. Zhang and H. Xu (2019). New Insights into the Interplay between Non-Coding RNAs and RNA-Binding Protein HnRNPK in Regulating Cellular Functions. Cells. 8(1):62.
- Yamaguchi S., H. Kimura, M. Tada, N. Nakatsuji and T. Tada (2005). Nanog expression in mouse germ cell development. Gene Expr Patterns. 5(5): 639-646.
- Yeung F., J. Hoberg, C. Ramsey, M. Keller, D. Jones, R. Frye and M. Mayo (2004). Modulation of NF-kappaB-dependent transcription and cell survival by the SIRT1 deacetylase. EMBO J. 23(12): 2369-2380.
- Zhang Z., W. Zhang, S. Huang, Q. Sun, Y. Wang, Y. Hu, N. Sun, Y. Zhang, Z. Jiang, N. Minato, J. Pin, L. Su and J. Liu (2015). GABAB receptor promotes its own surface expression by recruiting a Rap1-dependent signaling cascade. J Cell Sci. 128(12): 2302-2313.
|





