EVALUATION OF THE RELATIONSHIP BETWEEN HETEROSIS AND PARENTAL GENETIC DISTANCES BASED ON RAPD AND STS MARKERS IN COMMON BUCKWHEAT (FAGOPYRUM ESCULENTUM M.)
H.Y. Li1*†, Y.Z. Zhang2†, L.Q. Deng2, T.X. Shi1, Q.Y. Lv1 and Q.F. Chen1*
1Research Center of Buckwheat Industry Technology, Guizhou Normal University, Guiyang, 550001, P.R. China
2School of Geography and Life Science, Bijie College, Bijie, 551700, P.R. China
† These authors contributed equally to this work
*Correspondence Author’s Email: lihongyouluod@163.com; cqf1966@163.com
ABSTRACT
Common buckwheat is a nutritionally dense minor grain crop. However, the self-incompatible (SI), shattering, and low yield limits its wide cultivation. In this study, the hybrid experiments were performed to a) identify self-compatible parental material with non-shattering hybrid offspring, b) investigate the heterosis, and c) evaluate the relationship between heterosis and parental genetic distances (GD) in common buckwheat. As a result, nineteen self-compatible wild common buckwheat pure lines were identified with non-shattering hybrid offspring, and 58 hybrids were produced by single crossing between these 19 pure lines and four common buckwheat cultivars. For five agronomic and one quality trait, the 58 hybrids showed significant mid- and high-parent heterosis in common buckwheat. Furthermore, the GD values among parental lines were successfully calculated based on the random-amplified polymorphic DNA (RAPD) and sequence tagged site (STS) molecular markers. Non-significant relationship was observed between heterosis and parental GDs for any trait under this study, suggesting that the parental GDs could not be reliable to predict the heterosis in common buckwheat. However, present results are still valuable for further research in heterosis and parents-pair selection for improvement in common buckwheat through cross-breeding program.
Key words: common buckwheat, heterosis, RAPD markers, STS markers, genetic distance.
Abbreviations: GD, genetic distance; SI, self-incompatible; PH, plant height; NB, number of primary branches per plant; NSP, number of seeds per plant; SYP, seed yield per plant; SW, 1000-seed weight; FC, total flavonoid content; CTAB, cetyltrimethylammonium bromide; RAPD, random-amplified polymorphic DNA; STS, sequence tagged site; PCR, Polymerase chain reaction; MPH, Mid-parent heterosis; BPH, Best-parent heterosis; QTL, quantitative trait loci
https://doi.org/10.36899/JAPS.2021.6.0370
Published online March 31, 2021
INTRODUCTION
Heterosis is a common natural phenomenon appeared in higher plants, in which the offsprings in form of hybrids from two genetically diverse individuals exhibit greater biomass, higher quality, faster development, or better fertility than its parents (Birchler et al., 2010; Fu et al., 2010). The discovery and utilization of heterosis has significant contribution to crop improvement in aspects of yield, nutritional quality, biotic and abiotic stress resistance (Fu et al., 2010; Zhou et al., 2012; Wei et al., 2015). The genetic basis of heterosis is very complicated. To date, it is still not well understood, and the performance of F1 hybrids is hardly to predict. Exhilaratingly, some studies have suggested that a positive relationship between heterosis and parental GD, which could be used as a good predictor of hybrid performance (Betrán et al., 2003; Reif et al., 2003; Krystkowiak et al., 2009). As the GD can be evaluated through the DNA-based molecular markers, some studies have investigated the effects of GD on heterosis by using different DNA molecular markers (Banerjee and Kole, 2010; Jagosz, 2011; Rajendran et al., 2014; Tian et al., 2017; Gupta et al., 2018; Pandey et al., 2018). Results of some of these studies further confirmed a positive relationship between heterosis and parental GD, which suggested that DNA-based molecular markers could be used to heterosis prediction though estimate the parental GD.
Common buckwheat (Fagopyrum esculentum Moench) is a nutritionally dense minor grain crop widely cultivated in Asia, Europe and North America (Yasui et al., 2016). Its grains contain high levels of starch, protein, fatty acid, flavonoids, dietary fiber and a variety of minerals, which are beneficial to human health (Comino et al., 2013). However, the produce of common buckwheat face a real problem of low yield, which can't attract more farmers to plant it. One major reason resulting low yield of common buckwheat is heteromorphic self-incompatiblity (SI) due to its flower structure having long style, low anthers and small pollen grains (Fig. 1A) (Mizuno and Yasui, 2019; Takeshima et al., 2019), suggesting the necessity of utilizing heterosis. Thus, breeding self-compatible common buckwheat to produce hybrids of economic importance may be a good strategy to increase yield. In nature, there exist a few wild common buckwheat which are self-compatible due their flower with low style and long anthers (Fig. 1B), which may be ideal materials for breeding self-compatible common buckwheat hybrid. To date, a few studies have performed the hybridization between self-compatible wild common buckwheat and cultivated common buckwheat and investigated the heterosis of hybrid (Woo et al., 1999; Mukasa et al., 2010). However, a large-scale hybridization between self-compatible wild common buckwheat and cultivated common buckwheat has not carried out to investigate the heterosis and evaluate the relationship between heterosis and parental GD. In addition, these self-compatible wild common buckwheat have a problem of strong shattering. Therefore, screening self-compatible wild common buckwheat with non-shattering hybrid offspring, evaluation of its heterosis and relationship between parental GDs and heterosis will be helpful to self-compatible common buckwheat hybrid breeding.

Figure 1 - Floral morphotypes of self-incompatible (A) and self-compatible (B) common buckwheat. 1 represents anthers, 2 represents style
The objectives of this study were to a) identify self-compatible wild common buckwheat with non-shattering hybrid offspring, b) investigate the heterosis, and c) evaluate the relationship between heterosis and parental GD. These findings could provide valuable information for parent-pair selection in future common buckwheat crossing.
MATERIALS AND METHODS
Screening self-compatible wild common buckwheat with non-shattering hybrid offspring: A total of 423 self-compatible wild common buckwheat (low style and long anther) pure lines and one common buckwheat cultivar “Fengtian1” (SI, long style and low anther, non-shattering, high yield) were used. All the breeding material was planted in the growth chamber at the Research Center of Buckwheat Industry Technology of Guizhou Normal University, China, during spring 2016. At the flowering stage, “Fentian1”, which as the female parent, were single crossed with 423 self-compatible wild pure lines, respectively. For each cross combination, 100 flowers were used for hybridization. The parents and obtained hybrids were planted during autumn 2016. At the flowering stage, the flower morphology of hybrids were observed and were compared with their parents to sure the true hybrid. Then, the shattering effect of these true hybrids were observed to identify the self-compatible wild common buckwheat pure lines with non-shattering hybrid offspring.
Producing and planting of hybrids for heterosis analysis: Based above screening, 19 self-compatible wild common buckwheat pure lines with non-shattering hybrid offspring were obtained. The 19 self-compatible wild common buckwheat pure lines and four common buckwheat cultivars (“Pingqiao 2”, “Pinglu 1”, “Fengtian 1”, and “Weitian 1”) were used as parents. All parents were planted in pots of compost soil during spring 2017 in the growth chamber at the Research Center of Buckwheat Industry Technology of Guizhou Normal University, China. At the flowering stage, these four common buckwheat cultivars, which as the female parent, were single crossed with 19 self-compatible wild common buckwheat pure lines, respectively. To investigate agronomic and quality characteristics, all the 23 parents and hybrids were randomly grown in a field during autumn 2017 under the same cultivation conditions, with plant spacing of 10 cm and row width of 30 cm in Guiyang, Guizhou, China.
Investigation of agronomic and quality characteristics: Five agronomic and one quality characters namely plant height (PH), number of primary branches per plant (NBP), number of seeds per plant (NSP), seed yield per plant (SYP), 1000-seed weight (SW), and total flavonoid content (FC), respectively, were investigated. The measurement of total flavonoid content was performed as described previously (Lin et al., 2010).
DNA extraction, primers selected, PCR amplification, band analysis: The total genomic DNA from leaves of parents and hybrids were extracted by using cetyltrimethylammonium bromide (CTAB) method (Jiang, 2004). Then DNA was digested by RNaseA to eliminate the RNA, and the concentration and quality of DNA were analyzed by agarose gel electrophoresis (1.2%) and UV spectrophotometry. Based on the preliminary experimental results of 105 random-amplified polymorphic DNA (RAPD) primers and 50 sequence tagged site (STS) primer pairs, 7 RAPD primers and 15 STS primer pairs which displayed good polymorphism, clear band, and good repeatability between parents were selected for performing PCR analysis (Table S1 and Table S2). Polymerase chain reaction (PCR) amplification was performed in 20 μl reaction mixture. For RAPD amplification, the reaction mixture contained 10× PCR buffer (2.0 μl), 25.0 mM MgCl2 (1.5 μl), 10.0 mM dNTPs (0.4 μl), 4.0 μM primer (1.0 μl), 2.5 U/μl Taq DNA polymerase (0.2 μl), 40.0 ng/μl template DNA (3.0 μl), and ddH2O (11.9 μl). For STS amplification, the reaction mixture contained 10× PCR buffer (2.0 μl), 25.0 mM MgCl2 (1.2 μl), 10.0 mM dNTPs (0.4 μl), 10.0 μM forward primer (0.7 μl), 10.0 μM reverse primer (0.7 μl), 2.5 U/μl Taq DNA polymerase (0.3 μl), 40.0 ng/μl template DNA (3.0 μl), and ddH2O (11.7μl). The PCR reaction conditions were as follows: (i) pre-denaturation at 94°C for 10 min; (ii) 38 cycles of denaturation at 94°C for 1 min, annealing at (Tm-2) ℃ for 1 min, and extension at 72°C for 2 min; and (iii) extension at 72°C for 10 min. The amplified DNA fragments were separated by agarose gel electrophoresis (1.2%) and visualized by ethidium bromide staining. The gels were photographed by using a UV analyzer and a digital camera. For each sample, the PCR products presence of RAPD or STS amplification bands were recorded as 1 and the absence of a band was recorded as 0. Only the clearest and most highly repeatable bands were chosen.
Calculation of heterosis and parentalGD and evaluating the relationship between heterosis and parental GD: One way ANOVA was carried out to investigate the between all parents and hybirds for all tested traits. Heterosis was estimated as follows:
based on Jajosz's description (2011). For parental GD calculation, band profiles generated by RAPD or STS amplification were scored manually as 0 or 1, which represented absence or presence band, respectively. Then, the GD were calculated as follows:
,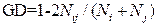
where Ni were the number of appeared bands for parent i, Nj were the number of appeared bands for parent j, and Nij were the number of shared bands for parent i and j (Nei and Li, 1979; Zhang et al., 2010). Correlation coefficients between heterosis and parental GD were estimated by using the GD values of 58 parental combinations and the MPH as well as HPH of the corresponding hybrids for yield and quality traits (Jagosz, 2011). SPSS11.5 software was used to analysis correlation coefficients.
RESULTS
Identification of self-compatible common buckwheat with non-shattering hybrid offspring. To identify the self-compatible wild common buckwheat with non-shattering hybrid offspring, 423 self-compatible wild common buckwheat pure lines were firstly tested by crossing with the common buckwheat cultivar “Fengtian1”. As a result, a total of 327 cross combinations successfully produced F1 hybrids, which had similar flower morphology with the self-compatible wild common buckwheat parent. Among them, although most F1 hybrids displayed strongly seed shattering, the seeds from 19 F1 hybrids were non-shattered. Therefore, these 19 corresponding self-compatible wild common buckwheat pure lines were selected as the one of parents for following study. The information of these 19 self-compatible wild common buckwheat pure lines with non-shattering hybrid offspring is listed in Table
Table 1 - Identified self-compatible wild common buckwheat with non-shattering hybrid.
|
No.
|
Name
|
Properties
|
Symbol
|
Native to
|
Origin
|
|
1
|
Tianzi 1
|
self-pollination
|
TZ 1
|
Guiyang
|
Chen qingfu
|
|
2
|
Tianzi 12
|
self-pollination
|
TZ 12
|
Guiyang
|
Chen qingfu
|
|
3
|
Tianzi 39
|
self-pollination
|
TZ 39
|
Guiyang
|
Chen qingfu
|
|
4
|
Tianzi 55
|
self-pollination
|
TZ 55
|
Guiyang
|
Chen qingfu
|
|
5
|
Tianzi 69
|
self-pollination
|
TZ 69
|
Guiyang
|
Chen qingfu
|
|
6
|
Tianzi 86
|
self-pollination
|
TZ 86
|
Guiyang
|
Chen qingfu
|
|
7
|
Tianzi95
|
self-pollination
|
TZ 95
|
Guiyang
|
Chen qingfu
|
|
8
|
Tianzi 118
|
self-pollination
|
TZ 118
|
Guiyang
|
Chen qingfu
|
|
9
|
Tianzi 141
|
self-pollination
|
TZ 141
|
Guiyang
|
Chen qingfu
|
|
10
|
Tianzi 159
|
self-pollination
|
TZ 159
|
Guiyang
|
Chen qingfu
|
|
11
|
Tianzi 171
|
self-pollination
|
TZ 171
|
Guiyang
|
Chen qingfu
|
|
12
|
Tianzi 187
|
self-pollination
|
TZ 187
|
Guiyang
|
Chen qingfu
|
|
13
|
Tianzi 193
|
self-pollination
|
TZ 193
|
Guiyang
|
Chen qingfu
|
|
14
|
Tianzi 208
|
self-pollination
|
TZ 208
|
Guiyang
|
Chen qingfu
|
|
15
|
Tianzi 219
|
self-pollination
|
TZ 219
|
Guiyang
|
Chen qingfu
|
|
16
|
Tianzi 228
|
self-pollination
|
TZ 228
|
Guiyang
|
Chen qingfu
|
|
17
|
Tianzi 237
|
self-pollination
|
TZ 237
|
Guiyang
|
Chen qingfu
|
|
18
|
Tianzi 253
|
self-pollination
|
TZ 253
|
Guiyang
|
Chen qingfu
|
|
19
|
Tianzi 268
|
self-pollination
|
TZ 268
|
Guiyang
|
Chen qingfu
|
F1 hybrid produce for heterosis analysis. To further evaluate the heterosis of common buckwheat, the previous obtained 19 self-compatible wild common buckwheat lines (as male parent) were single-cross with four common buckwheat cultivars “Pingqiao 2”, “Pinglu 1”, “Fengtian 1”, and “Weitian 1” (as female parent) to produce F1 hybrids. As result, a total of 58 cross combinations successfully produces non-shattering F1 hybrids (Table 3). Among these 19 self-compatible wild common buckwheat lines, 16, 14, 18, 10 lines could produce non-shattering F1 hybrids when they cross with “Pingqiao 2”, “Pinglu Tianqiao”, “Fengtian 1”, and “Weitian 1” (Table 3), respectively.
Performance of parental genotypes and F1 hybrids. The 6 investigation traits, including PH, NBP, NSP, SYP, SW and FC, among parents and 58 F1 hybrids had large variations. In addition, the range among the F1 hybrids was clearly higher than their parents for various traits (Table 2 and Table S3). Among these 6 traits, NSP had the most amplitude both in parents (96.22) and F1 hybrids (162.65). The mean value of each trait (except flavonoid content) of F1 hybrids was also significantly higher than that of their parents. Among them, the NSP and SYP displayed the most amplitude, which was 3-fold and 4-fold higher than that of their parents, respectively (Table 2).
Table 2 - Means and ranges of agronomic and quality traits of parental lines and hybrids
|
Traits
|
Performance
|
| |
Parental lines
|
|
F1 hybrids
|
| |
Mean
|
Range
|
|
Mean
|
Range
|
|
Plant height (cm)
|
47.6
|
32.31-65.91
|
|
60.54**
|
36.43-81.81
|
|
Number of primary branches / plant
|
3.04
|
2.31-3.83
|
|
3.73*
|
2.4-5.0
|
|
Number of seeds / plant
|
34.83
|
7.89-104.11
|
|
108.76**
|
39.75-202.4
|
|
Seed yield / plant (g)
|
0.71
|
0.14-2.03
|
|
2.9**
|
0.82-6.37
|
|
1000 seed weight (g)
|
20.18
|
11.7-29.94
|
|
26.43*
|
16.26-39.44
|
|
Flavonoid content (%)
|
0.125
|
0.006-0.544
|
|
0.152
|
0.002-0.533
|
*Significant at P = 0.05, **Significant at P = 0.01
Heterosis among F1 hybrids for various traits: PH. The range of MPH in 58 tested crosses was -27.23% to 46.67%, which had a mean value of 18.48% (Table 3). Among these crosses, 53 crosses presented positive heterosis, accounting for 66.18% of the crosses. The top three crosses with highest MPH were PL × TZ228 (46.47%), PL × TZ39 (42.68%) and PL × TZ171 (41.45%), respectively. The mean value of HPH was 3.67%, ranging from -39.01 to 36.83%. A total of 35 crosses showed positive heterosis, in which hybrids PL × TZ228 (36.83%), PL × TZ171 (30.39%), PL × TZ95 (29.05%) were the top three crosses with highest MPH.
NBP. For NBH, the range of MPH and HPH was -7.16% to 68.28% and -7.94% to 71.83%, with a mean value of 18.5% and 19.05%, respectively (Table 3). A total of 86.2% and 91.4% of these 58 crosses exhibited positive heterosis for MPH and HPH, respectively. The top three crosses with highest MPH and HPH were FT × TZ69 (68.28%), FT × TZ253 (51.08%), PQ × TZ1 (49.48%) and FT × T Z69 (71.83%), FT × TZ253 (53.01%), WT × TZ208 (50.27%), respectively.
NSP. For NSP, the range of MPH was -18.08% to 372.4%, which had a mean value of 112.2% (Table 3). Among these crosses, 57 crosses presented positive heterosis, of which the MPH value of about 50% crosses were over 100%. The top three crosses with highest MPH were PQ × TZ141 (372.4%), PL × TZ1 (299.69%) and PQ × TZ208 (268.33%), respectively. The range of HPH was -46.21% to 227.87%, with a mean value of 40.05%. A total of 44 crosses presented positive heterosis, in which the top three crosses were PL × TZ1 (227.87%), PQ × TZ141 (172.19%), PQ × TZ1 (164.39%), respectively.
SYP. The range of MPH for SYP was -19.21% to 472.9%, and had a mean value of 175.35% (Table 3). All crosses presented positive heterosis except one cross (PQ × TZ69), of which the MPH value of about 80% crosses were over 100%. The top three crosses with highest MPH were PQ × TZ141 (472.19%), FT × TZ253 (388.12%) and PL × TZ171 (385.06%), respectively. The range of HPH of these 58 crosses was -50.3% - 294.39%, and had a mean value of 78.2%. Among these crosses, 52 crosses presented positive heterosis, in which the top three crosses were PL × TZ171 (294.39%), SP × TZ1 (224.3%), PQ × TZ141 (224.24%), respectively.
For SW, the range of MPH and HPH was -16.74% to 87.66% and -25.07% to 54.01%, with a mean value of 28.33% and 16.53%, respectively (Table 3). A total of 53 and 42 crosses exhibited positive heterosis for MPH and HPH, respectively. The top three crosses with highest MPH and HPH are WT × TZ1 (87.66%), FT × TZ253 (73.44%), FT × TZ1 (68.53%) and PL × TZ171 (54.01%), FT × TZ237 (53.28%), FT × TZ171 (52.29%), respectively.
For FC, the mean value of MPH was 266.46% with a range of -98.8% to 2438.1% (WT ×TZ228) (Table 3). Among these crosses, 38 had positive heterosis, accounting for 65.51% of the total number of crosses. The three crosses with highest MPH were WT × TZ228 (2438.1%), WT × TZ219 (1526.32%), WT × TZ55 (1504.26%), respectively. The mean value of HPH was 177.69% with a range of -99.36% to 2217.39%. Among these crosses, 32 had positive heterosis, accounting for 55.2% of the total number of crosses. The three crosses with highest MPH were also the WT × TZ228 (2217.39%), WT × TZ219 (1326.63%), WT × TZ55 (1246.43%), respectively.
Table 3 - Mid- and high-parent heterosis of the measured agronomic and quality trait
|
No.
|
Hybrids
|
PH (%)
|
NBP (%)
|
NSP (%)
|
SYP (%)
|
SW (%)
|
FC (%)
|
| |
|
MPH
|
HPH
|
MPH
|
HPH
|
MPH
|
HPH
|
MPH
|
HPH
|
MPH
|
HPH
|
MPH
|
HPH
|
|
1
|
PQ × TZ1
|
31.73
|
10.16
|
49.48
|
48.81
|
258.99
|
164.39
|
363.11
|
189.09
|
39.83
|
5.48
|
-55.23
|
-77.21
|
|
2
|
PQ × TZ12
|
14.08
|
-4.91
|
15.01
|
8.75
|
138.30
|
42.82
|
131.09
|
35.15
|
-2.13
|
-6.04
|
114.21
|
12.83
|
|
3
|
PQ × TZ39
|
15.59
|
-7.08
|
35.78
|
26.49
|
203.86
|
98.02
|
290.77
|
130.91
|
42.61
|
12.57
|
-90.24
|
-93.55
|
|
4
|
PQ × TZ69
|
-27.23
|
-39.01
|
0.32
|
-6.55
|
-18.08
|
-46.21
|
-19.21
|
-50.30
|
4.83
|
-10.74
|
76.04
|
-7.73
|
|
5
|
PQ × TZ86
|
7.33
|
-11.55
|
29.17
|
13.99
|
138.12
|
50.62
|
162.24
|
55.76
|
38.52
|
23.04
|
129.23
|
24.17
|
|
6
|
PQ × TZ95
|
17.03
|
-0.55
|
30.29
|
19.05
|
150.46
|
38.52
|
238.55
|
83.64
|
35.70
|
20.74
|
315.00
|
137.14
|
|
7
|
PQ × TZ118
|
12.49
|
11.17
|
25.79
|
19.05
|
126.91
|
45.24
|
199.06
|
92.12
|
28.91
|
27.57
|
-67.83
|
-82.71
|
|
8
|
PQ × TZ141
|
13.83
|
12.02
|
18.78
|
11.49
|
372.40
|
172.19
|
472.19
|
224.24
|
22.58
|
12.83
|
72.89
|
-10.87
|
|
9
|
PQ × TZ159
|
3.41
|
-14.01
|
7.21
|
6.25
|
59.97
|
12.68
|
91.60
|
38.18
|
24.69
|
22.17
|
-75.93
|
-87.58
|
|
10
|
PQ × TZ171
|
24.55
|
3.08
|
20.30
|
19.05
|
139.98
|
68.87
|
212.07
|
119.39
|
34.50
|
29.09
|
-90.24
|
-93.55
|
|
11
|
PQ × TZ187
|
16.32
|
-3.45
|
18.01
|
9.23
|
129.89
|
56.90
|
158.37
|
82.42
|
3.75
|
-3.38
|
975.00
|
760.00
|
|
12
|
PQ × TZ193
|
-1.37
|
-4.14
|
12.81
|
11.66
|
98.88
|
93.04
|
147.32
|
131.05
|
23.69
|
22.25
|
-41.69
|
-69.82
|
|
13
|
PQ × TZ208
|
13.44
|
2.85
|
9.85
|
6.25
|
268.33
|
121.32
|
293.51
|
120.61
|
14.23
|
0.52
|
-25.41
|
-61.11
|
|
14
|
PQ × TZ237
|
11.94
|
-9.59
|
7.64
|
0.60
|
121.05
|
31.46
|
178.31
|
59.39
|
51.93
|
33.30
|
534.15
|
319.35
|
|
15
|
PQ × TZ253
|
22.62
|
16.16
|
23.56
|
11.61
|
83.07
|
18.79
|
139.46
|
61.82
|
14.58
|
8.01
|
83.10
|
6.56
|
|
16
|
PQ × TZ268
|
23.64
|
9.43
|
14.09
|
0.00
|
47.91
|
8.32
|
106.79
|
66.06
|
28.64
|
13.73
|
-3.23
|
-48.28
|
|
17
|
PL × TZ1
|
30.14
|
21.40
|
18.87
|
17.65
|
299.69
|
227.87
|
368.92
|
224.30
|
15.33
|
-11.24
|
-38.28
|
-67.10
|
|
18
|
PL × TZ12
|
16.94
|
8.69
|
15.48
|
9.81
|
93.17
|
22.57
|
107.41
|
30.84
|
-2.94
|
-4.15
|
-49.78
|
-70.05
|
|
19
|
PL × TZ39
|
42.68
|
27.16
|
32.70
|
22.94
|
31.04
|
-7.56
|
21.17
|
-22.43
|
13.97
|
-8.06
|
359.70
|
327.78
|
|
20
|
PL × TZ69
|
-0.09
|
-6.55
|
18.41
|
9.71
|
1.66
|
-27.60
|
42.07
|
-3.74
|
37.47
|
19.95
|
-22.63
|
-54.59
|
|
21
|
PL × TZ95
|
35.81
|
29.05
|
1.29
|
-7.94
|
124.97
|
28.65
|
205.79
|
72.90
|
21.02
|
10.51
|
296.23
|
200.00
|
|
22
|
PL × TZ118
|
19.76
|
7.51
|
29.38
|
21.76
|
36.35
|
-5.96
|
75.32
|
26.17
|
27.91
|
25.58
|
128.40
|
45.11
|
|
23
|
PL× TZ141
|
17.82
|
6.17
|
27.25
|
20.10
|
168.76
|
62.11
|
304.65
|
143.93
|
44.15
|
36.31
|
8.94
|
-39.44
|
|
24
|
PL × TZ159
|
24.91
|
15.75
|
40.60
|
38.53
|
170.97
|
110.51
|
213.33
|
163.55
|
10.12
|
9.20
|
-48.00
|
-71.02
|
|
25
|
PL × TZ171
|
41.45
|
30.39
|
37.52
|
35.29
|
196.75
|
130.24
|
385.06
|
294.39
|
55.99
|
54.01
|
58.21
|
47.22
|
|
26
|
PL × TZ187
|
28.16
|
18.53
|
23.00
|
13.24
|
133.72
|
74.63
|
166.29
|
117.76
|
5.54
|
-4.28
|
1342.86
|
741.67
|
|
27
|
PL × TZ219
|
33.98
|
21.53
|
19.75
|
15.00
|
76.95
|
40.26
|
179.49
|
103.74
|
22.36
|
11.20
|
580.00
|
419.44
|
|
28
|
PL × TZ228
|
46.47
|
36.83
|
29.45
|
27.35
|
246.46
|
136.34
|
379.71
|
209.35
|
24.66
|
13.92
|
588.14
|
463.89
|
|
29
|
PL × TZ237
|
28.18
|
14.85
|
4.11
|
-3.24
|
173.79
|
72.00
|
118.32
|
33.64
|
-16.74
|
-25.07
|
210.45
|
188.89
|
|
30
|
PL × TZ253
|
31.22
|
22.59
|
39.12
|
25.00
|
126.51
|
58.93
|
160.61
|
100.93
|
3.48
|
-5.04
|
15.46
|
-8.20
|
|
31
|
FT × TZ1
|
27.94
|
6.41
|
15.92
|
26.35
|
136.11
|
57.94
|
249.18
|
109.85
|
68.53
|
34.82
|
-28.95
|
-63.24
|
|
32
|
FT × TZ12
|
14.30
|
-5.24
|
-7.16
|
6.87
|
30.03
|
-25.76
|
67.10
|
-4.93
|
15.94
|
11.39
|
109.71
|
15.51
|
|
33
|
FT × TZ39
|
17.03
|
-6.41
|
34.14
|
36.97
|
9.34
|
-33.48
|
33.05
|
-23.65
|
36.06
|
14.46
|
-44.00
|
-54.84
|
|
34
|
FT × TZ55
|
14.15
|
-12.47
|
35.97
|
48.53
|
71.67
|
3.42
|
144.17
|
44.33
|
48.26
|
39.44
|
848.94
|
696.43
|
|
35
|
FT × TZ69
|
20.61
|
0.55
|
68.28
|
71.83
|
73.20
|
5.98
|
158.09
|
53.20
|
52.23
|
39.23
|
107.96
|
13.53
|
|
36
|
FT × TZ86
|
24.12
|
1.73
|
36.69
|
42.06
|
55.53
|
-7.51
|
117.95
|
25.62
|
41.76
|
35.79
|
-55.40
|
-74.17
|
|
37
|
FT × TZ95
|
17.54
|
-0.68
|
23.38
|
23.38
|
59.18
|
-14.38
|
91.71
|
2.46
|
31.12
|
25.85
|
329.21
|
172.86
|
|
38
|
FT × TZ118
|
-8.66
|
-10.34
|
4.67
|
8.65
|
46.58
|
-12.04
|
96.00
|
20.69
|
28.51
|
19.89
|
-7.89
|
-47.37
|
|
39
|
FT × TZ141
|
-14.17
|
-16.10
|
7.05
|
24.05
|
43.45
|
-20.47
|
108.89
|
15.76
|
46.45
|
45.85
|
72.43
|
-8.70
|
|
40
|
FT × TZ159
|
6.60
|
-11.84
|
-5.15
|
2.96
|
15.43
|
-25.43
|
63.77
|
11.33
|
43.28
|
34.93
|
-17.72
|
-56.37
|
|
41
|
FT × TZ171
|
23.98
|
2.06
|
0.30
|
8.73
|
92.11
|
24.00
|
208.15
|
104.93
|
58.47
|
52.29
|
856.00
|
670.97
|
|
42
|
FT × TZ187
|
28.37
|
5.98
|
33.92
|
35.82
|
91.74
|
20.86
|
114.76
|
43.35
|
0.30
|
-13.17
|
548.00
|
326.32
|
|
43
|
FT × TZ193
|
4.78
|
2.51
|
2.04
|
12.72
|
19.33
|
4.94
|
19.08
|
15.27
|
-0.44
|
-9.00
|
136.84
|
26.32
|
|
44
|
FT × TZ208
|
26.78
|
14.24
|
-4.46
|
1.35
|
95.62
|
11.90
|
254.26
|
94.58
|
46.67
|
39.08
|
177.47
|
50.00
|
|
45
|
FT × TZ228
|
15.63
|
8.84
|
-5.78
|
2.14
|
74.31
|
3.90
|
102.56
|
16.75
|
15.59
|
11.03
|
471.43
|
421.74
|
|
46
|
FT × TZ237
|
3.39
|
-16.93
|
-0.68
|
1.75
|
20.65
|
-31.51
|
87.67
|
4.93
|
62.18
|
53.28
|
180.00
|
125.81
|
|
47
|
FT × TZ253
|
13.00
|
6.36
|
51.08
|
53.01
|
157.27
|
55.99
|
388.12
|
213.79
|
73.44
|
51.81
|
72.50
|
13.11
|
|
48
|
FT × TZ268
|
9.08
|
-4.03
|
7.91
|
12.99
|
84.45
|
22.84
|
181.85
|
110.34
|
35.19
|
11.62
|
51.22
|
-14.48
|
|
49
|
WT × TZ1
|
40.46
|
13.03
|
-4.80
|
9.69
|
165.07
|
86.62
|
357.64
|
178.72
|
87.66
|
41.58
|
-96.45
|
-98.16
|
|
50
|
WT × TZ55
|
36.98
|
2.06
|
29.80
|
33.61
|
132.54
|
45.05
|
178.22
|
66.49
|
14.16
|
-0.26
|
1504.26
|
1246.43
|
|
51
|
WT × TZ95
|
26.70
|
3.50
|
11.87
|
18.93
|
126.07
|
23.38
|
171.29
|
45.74
|
20.63
|
7.35
|
455.06
|
252.86
|
|
52
|
WT ×TZ159
|
14.15
|
-8.63
|
-2.42
|
12.00
|
98.04
|
33.95
|
172.03
|
88.83
|
36.66
|
33.93
|
-98.80
|
-99.36
|
|
53
|
WT ×TZ171
|
5.59
|
-15.85
|
14.59
|
31.36
|
174.61
|
85.57
|
243.53
|
132.98
|
24.06
|
19.10
|
232.00
|
167.74
|
|
54
|
WT ×TZ187
|
11.07
|
-11.24
|
10.84
|
19.40
|
69.73
|
11.59
|
103.13
|
38.30
|
9.08
|
1.58
|
284.00
|
152.63
|
|
55
|
WT ×TZ208
|
13.21
|
-1.68
|
33.76
|
50.27
|
47.46
|
-13.44
|
146.15
|
36.17
|
44.45
|
27.14
|
183.79
|
53.42
|
|
56
|
WT ×TZ219
|
33.11
|
24.11
|
18.21
|
32.62
|
185.27
|
95.81
|
186.92
|
80.85
|
3.31
|
-8.48
|
1526.32
|
1326.63
|
|
57
|
WT ×TZ228
|
21.55
|
10.04
|
13.37
|
29.97
|
132.00
|
42.92
|
210.50
|
80.85
|
21.66
|
8.35
|
2438.10
|
2217.39
|
|
58
|
WT ×TZ268
|
27.78
|
8.48
|
-5.14
|
-3.61
|
0.55
|
-29.57
|
21.53
|
-6.91
|
8.86
|
-3.77
|
-60.98
|
-77.93
|
Note: PH, plant height; NBP, number of primary branches per plant; NSP, number of seeds per plant; SYP, seed yield per plant; SW, 1000-seed weight (SW); FC, total flavonoid content; MPH, Mid-parent heterosis; HPH, Better-parent heterosis.
Parental GD. The GDs of parent-pairs were estimated by using RAPD and STS markers. As a result, GDRAPD and GDSTS of the parent-pairs ranged from 0.103 to 3.235 and from 0.275 to 1.110, with an average of 0.563 and 0.587, respectively (Table 4). For GDRAPD, the three parent-pairs with highest or lowest GD were TZ187 and TZ159 (3.235), TZ187 and TZ1 (1.973) and TZ187 and TZ39 (1.742) or TZ86 and TZ69 (0.103), TZ86 and TZ39 (0.11) and TZ69 and TZ39 (0.129), respectively (Table 4). For GDSTS, the SP and TZ159 (1.110), TZ159 and TZ69 (1.059) and PQ and TZ159 (1.042) had the most GD, and TZ55 and TZ12 (0.275), TZ95 and TZ86 (0.276) and TZ69 and TZ55 (0.0.293) had the lowest GD, respectively (Table 4).
Table 4 - GD of parental lines estimated by RAPD (GDRAPD) and STS (GDSTS) markers
|
Parent
|
TZ1
|
TZ12
|
TZ39
|
TZ55
|
TZ69
|
TZ86
|
TZ95
|
TZ118
|
TZ141
|
TZ159
|
TZ1171
|
TZ187
|
TZ193
|
TZ208
|
TZ219
|
TZ228
|
TZ237
|
TZ253
|
TZ268
|
PQ
|
SP
|
FT
|
WT
|
|
GDRAPD
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ1
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ12
|
0.255
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ39
|
0.326
|
0.200
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ55
|
0.325
|
0.173
|
0.218
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ69
|
0.405
|
0.195
|
0.129
|
0.190
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ86
|
0.305
|
0.131
|
0.110
|
0.147
|
0.103
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ95
|
0.385
|
0.246
|
0.271
|
0.243
|
0.249
|
0.199
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ118
|
0.503
|
0.349
|
0.296
|
0.348
|
0.321
|
0.274
|
0.223
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ141
|
0.472
|
0.559
|
0.526
|
0.564
|
0.636
|
0.580
|
0.575
|
0.580
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ159
|
0.781
|
0.688
|
0.611
|
0.744
|
0.744
|
0.641
|
0.641
|
0.681
|
0.815
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ171
|
0.561
|
0.516
|
0.447
|
0.524
|
0.524
|
0.433
|
0.468
|
0.433
|
0.505
|
0.703
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ187
|
1.973
|
1.560
|
1.742
|
1.533
|
1.380
|
1.513
|
1.498
|
1.360
|
1.695
|
3.235
|
1.402
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ193
|
0.462
|
0.366
|
0.366
|
0.366
|
0.366
|
0.291
|
0.405
|
0.376
|
0.501
|
0.758
|
0.235
|
1.502
|
0
|
|
|
|
|
|
|
|
|
|
|
|
TZ208
|
0.464
|
0.423
|
0.453
|
0.311
|
0.520
|
0.468
|
0.606
|
0.535
|
0.606
|
0.671
|
0.495
|
1.684
|
0.366
|
0
|
|
|
|
|
|
|
|
|
|
|
TZ219
|
0.348
|
0.339
|
0.366
|
0.366
|
0.395
|
0.318
|
0.347
|
0.347
|
0.500
|
0.654
|
0.325
|
1.350
|
0.308
|
0.319
|
0
|
|
|
|
|
|
|
|
|
|
TZ228
|
0.372
|
0.335
|
0.392
|
0.393
|
0.424
|
0.344
|
0.405
|
0.344
|
0.374
|
0.640
|
0.392
|
1.318
|
0.305
|
0.364
|
0.223
|
0
|
|
|
|
|
|
|
|
|
TZ237
|
0.373
|
0.280
|
0.280
|
0.306
|
0.306
|
0.233
|
0.401
|
0.287
|
0.393
|
0.822
|
0.427
|
1.257
|
0.334
|
0.395
|
0.249
|
0.245
|
0
|
|
|
|
|
|
|
|
TZ253
|
0.493
|
0.421
|
0.392
|
0.424
|
0.456
|
0.374
|
0.374
|
0.315
|
0.436
|
0.727
|
0.325
|
1.471
|
0.277
|
0.426
|
0.220
|
0.363
|
0.218
|
0
|
|
|
|
|
|
|
TZ268
|
0.513
|
0.473
|
0.359
|
0.476
|
0.386
|
0.367
|
0.488
|
0.456
|
0.590
|
0.650
|
0.409
|
1.369
|
0.248
|
0.478
|
0.357
|
0.446
|
0.384
|
0.415
|
0
|
|
|
|
|
|
PQ
|
0.631
|
0.622
|
0.484
|
0.594
|
0.523
|
0.538
|
0.612
|
0.575
|
0.612
|
1.045
|
0.500
|
1.068
|
0.528
|
0.725
|
0.564
|
0.651
|
0.461
|
0.571
|
0.446
|
0
|
|
|
|
|
SP
|
0.820
|
0.811
|
0.657
|
0.916
|
0.783
|
0.722
|
0.806
|
0.763
|
0.683
|
0.893
|
0.572
|
1.543
|
0.712
|
0.840
|
0.673
|
0.681
|
0.855
|
0.808
|
0.655
|
0.604
|
0
|
|
|
|
FT
|
0.560
|
0.461
|
0.464
|
0.462
|
0.462
|
0.472
|
0.601
|
0.472
|
0.637
|
0.780
|
0.490
|
1.166
|
0.375
|
0.462
|
0.403
|
0.526
|
0.344
|
0.431
|
0.423
|
0.344
|
0.862
|
0
|
|
|
WT
|
0.597
|
0.517
|
0.483
|
0.598
|
0.489
|
0.435
|
0.469
|
0.403
|
0.746
|
0.739
|
0.429
|
1.438
|
0.494
|
0.568
|
0.425
|
0.537
|
0.428
|
0.499
|
0.445
|
0.329
|
0.570
|
0.310
|
0
|
|
GDSTS
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ1
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ12
|
0.427
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ39
|
0.654
|
0.433
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ55
|
0.485
|
0.275
|
0.469
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ69
|
0.645
|
0.334
|
0.423
|
0.293
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ86
|
0.611
|
0.304
|
0.482
|
0.397
|
0.311
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ95
|
0.569
|
0.363
|
0.508
|
0.406
|
0.367
|
0.276
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ118
|
0.811
|
0.592
|
0.509
|
0.486
|
0.572
|
0.544
|
0.528
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ141
|
0.852
|
0.500
|
0.518
|
0.578
|
0.408
|
0.431
|
0.410
|
0.692
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ159
|
0.984
|
0.833
|
0.876
|
0.953
|
1.059
|
0.809
|
0.886
|
0.842
|
0.923
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ171
|
0.831
|
0.734
|
0.733
|
0.7667
|
0.861
|
0.749
|
0.872
|
0.612
|
0.832
|
0.717
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ187
|
0.709
|
0.507
|
0.589
|
0.503
|
0.565
|
0.562
|
0.511
|
0.553
|
0.578
|
0.892
|
0.693
|
0
|
|
|
|
|
|
|
|
|
|
|
|
|
TZ193
|
0.772
|
0.565
|
0.678
|
0.538
|
0.600
|
0.578
|
0.644
|
0.612
|
0.674
|
0.871
|
0.682
|
0.336
|
0
|
|
|
|
|
|
|
|
|
|
|
|
TZ208
|
0.727
|
0.442
|
0.557
|
0.481
|
0.429
|
0.432
|
0.493
|
0.548
|
0.450
|
0.910
|
0.735
|
0.442
|
0.536
|
0
|
|
|
|
|
|
|
|
|
|
|
TZ219
|
0.615
|
0.564
|
0.601
|
0.578
|
0.619
|
0.595
|
0.620
|
0.542
|
0.651
|
0.875
|
0.636
|
0.455
|
0.571
|
0.400
|
0
|
|
|
|
|
|
|
|
|
|
TZ228
|
0.629
|
0.445
|
0.641
|
0.443
|
0.505
|
0.485
|
0.493
|
0.508
|
0.521
|
0.781
|
0.693
|
0.523
|
0.646
|
0.481
|
0.381
|
0
|
|
|
|
|
|
|
|
|
TZ237
|
0.634
|
0.426
|
0.548
|
0.449
|
0.373
|
0.411
|
0.452
|
0.587
|
0.479
|
0.809
|
0.703
|
0.414
|
0.485
|
0.415
|
0.392
|
0.347
|
0
|
|
|
|
|
|
|
|
TZ253
|
0.673
|
0.462
|
0.660
|
0.442
|
0.503
|
0.467
|
0.458
|
0.526
|
0.450
|
0.853
|
0.785
|
0.521
|
0.598
|
0.412
|
0.474
|
0.318
|
0.432
|
0
|
|
|
|
|
|
|
TZ268
|
0.609
|
0.440
|
0.570
|
0.478
|
0.479
|
0.517
|
0.605
|
0.632
|
0.614
|
1.006
|
0.696
|
0.478
|
0.535
|
0.476
|
0.533
|
0.521
|
0.404
|
0.455
|
0
|
|
|
|
|
|
PQ
|
0.790
|
0.646
|
0.693
|
0.612
|
0.676
|
0.566
|
0.653
|
0.675
|
0.727
|
1.042
|
0.717
|
0.547
|
0.583
|
0.545
|
0.460
|
0.547
|
0.509
|
0.525
|
0.545
|
0
|
|
|
|
|
SP
|
0.795
|
0.705
|
0.756
|
0.602
|
0.733
|
0.704
|
0.708
|
0.709
|
0.761
|
1.110
|
0.851
|
0.501
|
0.704
|
0.559
|
0.573
|
0.582
|
0.620
|
0.520
|
0.579
|
0.451
|
0
|
|
|
|
FT
|
0.645
|
0.576
|
0.705
|
0.553
|
0.575
|
0.557
|
0.565
|
0.598
|
0.729
|
0.898
|
0.714
|
0.459
|
0.495
|
0.514
|
0.508
|
0.573
|
0.521
|
0.495
|
0.450
|
0.465
|
0.465
|
0
|
|
|
WT
|
0.623
|
0.543
|
0.666
|
0.534
|
0.554
|
0.551
|
0.597
|
0.498
|
0.740
|
0.986
|
0.709
|
0.451
|
0.507
|
0.531
|
0.546
|
0.556
|
0.592
|
0.597
|
0.489
|
0.416
|
0.509
|
0.410
|
0
|
The correlation of parental GD with offspring’s heterosis. The correlations between parental GDs and offspring’s heterosis were listed in Table 5. The correlation between the RAPD measured GD and the MPH for PH, NSP, SYP, and FC was positive, whereas that for NBP and SW was negative. In addition, PH, SYP, and NSP showed positive BPH correlation, while negative BPH correlation were observed for NBP and SW. Notably, none of these correlations were significant except the MPH for SW was significantly negatively correlated with the parental GD (r=-0.294*, P<0.05). For the correlation based on STS markers, the MPH of NSP, SYP and SW were positively correlated with parental GD, while the MPH of PH and FC were negatively correlated with parental GD. The BPH of PH, NSP, SYP and SW were positively correlated with parental GD, while the BPH of FC was negatively correlated with parental GD. Furthermore, no significant correlation was found except MPH of FC was significantly negatively correlated with the parental GD (r=-0.256*, P=0.05).
Table 5 - Correlations between GD determined using RAPD or STS markers and heterosis of different agronomic and quality traits
| |
MPH correlations
|
|
BPH correlations
|
| |
PH
|
NBP
|
NSP
|
SYP
|
SW
|
FC
|
|
PH
|
NBP
|
NSP
|
SYP
|
SW
|
FC
|
|
GDRAPD
|
0.137
|
- 0.103
|
0.109
|
0.007
|
- 0.294*
|
0.041
|
|
0.229
|
- 0.1
|
0.109
|
0.029
|
- 0.188
|
-0.006
|
|
GDSTS
|
-0.037
|
0.009
|
0.151
|
0.134
|
0.061
|
- 0.256*
|
|
0.018
|
0
|
0.183
|
0.180
|
0.174
|
-0.216
|
*Significant at P = 0.05, n=58
DISSCUSSION
Discovery of heterosis in higher plant had contribute significantly to improve crop yield and quality (Fu et al., 2010). Common buckwheat is a considerable self-incompatible (SI) and nutritionally dense minor grain crop, and its low yield limits its cultivation. Utilizing heterosis to breed self-compatible buckwheat hybrids may be an effective method to solve its low-yield problem. In nature, there are a few of self-compatible wild common buckwheat genotypes, which makes the breeding of self-compatible common buckwheat hybrids possible. Furthermore, it also has a considerable shattering problem. Therefore, it is very important to identify a self-compatible wild common buckwheat with non-shattering hybrid offspring. In this study, we obtained 19 self-compatible lines with non-shattering hybrid offsprings through screening 423 self-compatible wild common buckwheat pure lines crossed with one common buckwheat cultivar “Fengtian1”, which provided extremely valuable material for the breeding of self-compatible common buckwheat hybrids. Then these 19 self-compatible lines were crossed with 4 cultivars. Obvious heterosis, for five agronomic traits and one quality trait, was observed in hybrid offspring, suggesting that heterosis could be used in common buckwheat breeding.
In general, the degree of heterosis depends on the degree of the genetic differences and complementary traits of parents (Zhang et al., 2010). However, the genetic basis of heterosis is very complicated and hardly to predict. DNA molecular markers have been used to study the heterosis prediction due to its simplicity. To date, there are two different views on the heterosis predication based on the GD from the molecular marker. Some researchers think that the GD of parents based on molecular markers can be used to predict heterosis due to a significant correlation between parental GD and offspring’s heterosis (Zhang et al., 2000; Cai et al., 2005; Yao et al., 2016). In contrast, other researchers believe that the GD based on molecular markers can’t be reliable to predict heterosis because there is no significant correlation between parent GD and offspring’s heterosis (Zhang et al., 2006; Zhao et al., 2009; Qian et al., 2009; Luo et al., 2016; Tian et al., 2017). In our study, we explored the relationship between parental GD and heterosis in common buckwheat based on RAPD and STS markers. The GDs among the tested lines were successfully detected by both RAPD and STS markers. However, no significant positive correlations existed between parental GD and heterosis, which was alike to the results reported in the previous studies (Zhang et al., 2006; Zhao et al., 2009; Qian et al., 2009; Luo et al., 2016; Tian et al., 2017), suggesting that the parental GD could not be reliable to predict the heterosis of common buckwheat at least in this study. This might be attributed to the following two reasons. Firstly, the number of RAPD or STS markers was too small to cover all target traits (quantitative traits). Therefore, more markers, such as SSR or SNP, need to be used in further studies. Secondly, this uncorrelation might be a result that RAPD or STS markers were not linked to the target genes controlling target traits. Thus, further studies such as quantitative trait loci (QTL) mapping and development SSR or SNP markers that link to agronomic traits are required.
In conclusion, self-compatible wild common buckwheat with non-shattering hybrid offspring existed in nature. Common buckwheat has significant heterosis. Although no significant correlations between heterosis and parental GD were observed, this study will still be helpful for parent selection in future self-compatible common buckwheat hybrid breeding.
ACKNOWLEDGEMENTS: This research was financially supported by the National Natural Science Foundation of China-Project of Karst Science Research Center of Guizhou Provincial People’s Government (U1812401), the Natural Sciences Foundation of China (31471562 and 31701494), and the Science and Technology Foundation of Guizhou Province (QianKeHeJiChu [2019]1235).
REFERENCES
- Banerjee, P. and P. Kole (2010). Heterosis, inbreeding depression and their relationship with genetic divergence in sesame (Sesamum indicum ). Acta. Agronomica. Hungarica. 58: 313-321.
- Betrán, F.J., M. Ribaut, D. Beck and D. Gonzalez de Leon (2003). Genetic diversity, specific combining ability, and heterosis in tropical maize under stress and non-stress environments. Crop Sci. 43: 797-806.
- Birchler, J.A., H. Yao, S. Chudalayandi, D. Vaiman and R.A. Veitia (2010). Heterosis. Plant Cell. 22: 2105-2112.
- Cai, J. and W. Lan (2005). Using of AFLP marker to predict the hybrid yield and yield heterosis in rice. Chin. Agric. Sci. Bull. 21: 39-43.
- Comino, I., M. de Lourdes Moreno, A. Real, A. Rodríguez-Herrera, F. Barro and C. Sousa (2013). The gluten-free diet: testing alternative cereals tolerated by celiac patients. Nutrients. 5: 4250-4268.
- Fu, D.H., M. Xiao, A. Hayward, Y. Fu, G. Liu, G.J. Jiang and H.H. Zhang (2010). Utilization of crop heterosis: a review. Euphytica 197: 161-173.
- Gupta, S.K., T. Nepolean, C.G. Shaikh, K. Kedarnath Rai, C.T. Hash, , R.R. Roma Rani Das and A. Rathore (2018). Phenotypic and molecular diversity-based prediction of heterosis in pearl millet (Pennisetum glaucum (R.) Br.). The Crop J. 6: 271-281.
- Jagosz, B., (2011). The relationship between heterosis and genetic distances based on RAPD and AFLP markers in carrot. Plant Breed. 130: 574-579.
- Jiang, J. (2004). Principle and Technology of Molecular Biology Experimentation. Haerbin: Press of Northeast Forest University. 28-29.
- Krystkowiak, K., T. Adamski, M. Surma and Z. Kaczmarek (2009). Relationship between phenotypic and genetic diversity of parental genotypes and the specific combining ability and heterosis effects in wheat (Triticum aestivum). Euphytica 165: 419-434.
- Lin, Y. X., K.F. Huang and Q.F. Chen (2010). Comparison of two methods for measuring flavonoid content of Fagopyrum megaspartanium J. Anhui Agric. Sci. 11: 1210-1224.
- Luo, X., C. Ma, B. Yi, J. Tu, J. Shen and T. Fu (2016). Genetic distance revealed by genomic single nucleotide polymorphisms and their relationships with harvest index heterotic traits in rapeseed (Brassica napus). Euphytica 209: 41-47.
- Mizuno, N., and Y. Yasui (2019). Gene flow signature in the S-allele region of cultivated buckwheat. BMC Plant Biol. 19: 125.
- Mukasa Y., T. Suzuki T and Y. Honda (2010). A methodology for heterosis breeding of common buckwheat involving the use of the self-compatibility gene derived from Fagopyrum homotropicum. Euphytica 172: 207-214.
- Nei M., and W. Li (1979). Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. U. S. A. 76: 5269-5273.
- Pandey, S.K., T. Dasgupta, A. Rathore and A. Vemula (2018). Relationship of parental genetic distance with heterosis and specific combining ability in Sesame (Sesamum indicum ) based on phenotypic and molecular marker analysis. Biochem. Genet. 56: 1-22.
- Qian, W., Q. Li, J. Noack, O. Sass, J. Meng, M. Frauen and C. Jung (2009). Heterotic patterns in rapeseed (Brassica napus): II. Crosses between European winter and Chinese semi-winter lines. Plant Breed. 128: 466-470.
- Rajendran, A., A. Muthiah, J. Joel, P. Shanmugasundaram and D. Raju (2014). Heterotic grouping and patterning of quality protein maize inbreds based on genetic and molecular marker studies. Turk. J. Biol. 38: 10-20.
- Reif, J.C., A.E Melchinger, X.C. Xia, M.L. Warburton, D.A. Hoisington, S.K. Vasal, G. Srinivasan, M. Bohn and M. Frisch (2003). Genetic distance based on simple sequence repeats and heterosis in tropical maize populations. Crop Sci. 43: 1275-1282.
- Takeshima, R., T. Nishio, S. Komatsu, N. Kurauchi and K. Matsui (2019). Identification of a gene encoding polygalacturonase expressed specifically in short styles in distylous common buckwheat (Fagopyrum esculentum). Heredity. 123: 492-502.
- Tian, H.Y., S.A. Channa and S.W. Hu (2017). Relationships between genetic distance, combining ability and heterosis in rapeseed (Brassica napus). Euphytica 213: 1.
- Wei, X.Y., B. Wang, Q. Peng, F. Wei, K.J. Mao, X.G. Zhang, P. Sun, Z.H. Liu and J.H. Tang (2015). Heterotic loci for various morphological traits of maize detected using a single segment substitution lines test-cross population. Mol. Breed. 35: 94.
- Woo, S.H., T. Adachi, S.K. Jong and C.G. Campbell (1999). Inheritance of self-compatibility and flower morphology in an inter-specific buckwheat hybrid. Can. J. Plant Sci. 79: 483-490.
- Yao, J.X., H.G. Li, J. Ye and L.L. Shi (2016). Relationship between parental genetic distance and offspring’s heterosis for early growth traits in Liriodendron: implication for parent pair selection in cross breeding. New Forest. 47: 163-177.
- Yasui, Y., H. Hirakawa, M. Ueno, K. Matsui, T. Katsube-Tanaka, S.J.Yang, J. Aii, S. Sato and M. Mori (2016). Assembly of the draft genome of buckwheat and its applications in identifying agronomically useful genes. DNA Res. 23: 215-224.
- Zhang, P.J., H.W. Cai, H.C. Li, L.S. Yang, Y.S. Bai, X.M. Hu and C.W. Xu (2000). RAPD molecular markers of rice genetic distance and its relationship with heterosis. J. Anhui Agric. Univ. 28: 697-700.
- Zhang, T., L. Han, J.D. Xu, K.F. Jiang, X.J. Wu, X.D. Wang and J.K. Zheng (2006). Correlation between genetic distance and yield heterosis of hybrid aromatic rice. Sci. Agric. Sin. 39: 831-835.
- Zhang, T., X.L. Ni, K.F. Jiang, H.F. Deng, Q. He, Q.H. Yang, Y. Li, X.Q. Wan, Y.J. Cao and J.K. Zheng (2010). Relationship between heterosis and parental genetic distance based on molecular markers for functional genes related to yield traits in rice. Rice Sci. 17: 288-295.
- Zhao, Q.Y., Z. Zhu, Y.D. Zhang, L. Zhao, T. Chen, Q.F. Zhang and C.L. Wang (2009). Analysis on correlation between heterosis and genetic distance based on simple sequence repeat markers in japonica rice. China J. Rice Sci. 23: 141-147.
- Zhou, G., Y. Chen, W. Yao, C. Zhang, W. Xie, J. Hua, Y. Xing, J. Xiao and Q. Zhang (2012). Genetic composition of yield heterosis in an elite rice hybrid. Proc. Natl. Acad. Sci. U. S. A. 109: 15847- 15852.
|





