TRANSFORMATION OF UPLAND COTTON WITH PINELLIA TERNATA AGGLUTININ GENE AGAINST SUCKING PESTS
S. Ahmed1, 2 and I. A. Nasir1
1Seed Biotechnology Lab, Centre of Excellence in Molecular Biology, University of the Punjab, Lahore
2The Superior College, Lahore
Corresponding author’s email: shafiq@cemb.edu.pk
ABSTRACT
Cotton plants are normally targeted by numerous challenges from which 75% are biotic. Among these biotic stresses, insects constitute major segment causing direct or indirect considerable yield loss. Bt cotton, corn and soybean are mostly resistant to chewing insects in the field, but their resistance to sap sucking pests has not yet achieved. Likewise Bt protein against chewing insect, there are some proteins characterized for their actions against sucking insects e.g. lectins. Major function of plant lectin is to provide anti-insect activity with other known and unknown function. Therefore, all necessary in-silico, in-vivo and plant transformation experimentation adopted to design a codon optimized (cotton targeted) Pinellia ternata agglutinin (pta) gene cassette under the influence of well-tested CaMV35S promoter in pCAMBIA2300 expression vector enabled by agrobacterium LB4404 plant infecting strain for stable delivery and integration of transgene in the recipient genome. Stable transformation of at least five different cotton plants (transformation efficiency 0.53%) took place successively, which was tested through PCR and Southern blotting in T0 and T1 generation. This work has provided genotypic bases and a candidate phenotype of commercial cotton crop resistant against sucking pests.
Keywords: Nuclear Transformation, Pinellia ternata agglutinin, Cotton
https://doi.org/10.36899/JAPS.2021.5.0339
Published online January 24, 2021
INTRODUCTION
There have been always efforts to strengthen crop culture through variety of methods (Tariq et al. 2020, Naheed et al. 2020). Plants face many stresses and diseases causing significant production loss and increase in cost of production. These stresses include many faces from living and non-living origins called biotic and abiotic factors, respectively. Biotic factors include fungi, insects, nematodes, viruses or any types of pathogens while abiotic factors include temperature, water, minerals and salt stresses. In all tropical regions of the world, virulent viruses are major cause in the decline of productivity of crop plants. Geminiviridae, family of viruses, have most of members of virulent or pathogenic nature posing biggest challenge to agriculture produce. Interestingly these disease-causing viruses transfer to the plant through one of most abundant and diverse set of set species known as insects (Loriato et al. 2020). Cotton crop plants are no exception when it comes to susceptibility to these sucking, chewing and virus delivering insects.
Cotton plants are generally targeted by numerous challenges from which 75% are biotic. As outline above regarding biotic stresses, insects constitute major segment among this class causing direct or indirect considerable yield loss. Bt cotton, corn and soybean are those Genetically Modified Organisms or GMOs which are spread all around the world. These GMOs are mostly resistant to chewing insects in the field, but their resistance to sap sucking pests has not yet achieved. Major sapsuckers include thrips, leafhoppers, whitefly, mired bugs and aphids are part of it. Among these pathogens, whitefly is a serious issue for growth of cotton in Pakistan (Khan et al. 2015). Whitefly (Bemisia tabaci) is an insect that harms field by sucking sap and then transmitting viral infections in plants. Whiteflies can carry different hosts that cause disease on many plants. They cause harm by sucking phloem sap bringing them about to early shrinking which causes premature defoliation and hindrance of growth that finally leads to yield loss. Also, honeydew secreted by aphids leave whitefly on the commercial parts of the plant lowering its market price and value (Navas-Castillo et al. 2011).
Learning from the case of controlling chewing insect, there is need of development of genetically modified cotton plants that can defy whitefly through transgenically expressed specific toxic metabolites for effective results. But no transgenic cotton plant is reported till now which has a strong and effective defence against whitefly. Likewise Bt protein against chewing insect, there are some proteins characterized for their actions against sucking insects e.g. lectins. The main role of lectin is still unclear, but the studies have shown that they are accumulated in the storage of vegetative tissues and also play role in natural plant defence mechanism as insecticidal protein when secreted in extracellular environment (Rüdiger and Gabius 2001, Macedo et al. 2015).
Considering the above given preamble on threat posed by the whitefly to cotton crop, challenges to IPM and available characterized nature of lectin protein against insect, this study was designed to test the hypothesis that transgenic cotton plants with Pinellia ternata agglutinin would show Mendelian inheritance regarding the transgene and induce tolerance to sucking pests.
MATERIALS AND METHODS
Codon Optimization and pta Gene Synthesis: The National Centre for Biotechnology Information GenBank database was used to retrieve pta gene (Gene Bank Accession number AY191305) sequence (Pruitt et al. 2007). Sequence optimization of pta gene was done by substituting codons prophesied often utilized in cotton with extra preferential codons devoid of modifying the amino acid sequence (Perlak et al. 1991, Owczarzy et al. 2008, Latif et al. 2015) by the means of codon usage database (https://eu.idtdna.com/CodonOpt). The pUC57 vector having selection drug tetracycline was used for chemical synthesis of codon-optimized gene sequence under CaMV35S promoter with Sma I and Pst I restriction sites.
Polymerase Chain Reaction (PCR): Designed primers were used with their expected product size and sequences as given in table 1. For confirmation of designed primers to check their gene orientation polymerase chain reaction was performed. Optimization of primer pair for pta gene were done by using gradient thermo-cycler by allocating PCR tubes at different temperature ranges. Reaction mixture of 20 µl were followed as given below in table 2.
Table 1: showing sequence of primers with their product size for pta gene amplification
|
Primer name
|
Primer sequence
|
Product size
|
Tm
|
|
mCEMB-Cot1-G3F
|
TCCAACGGCTTTAGCAAGTC
|
588
|
54.8℃
|
|
mCEMB-Cot1-G3R
|
TGCCATGTAGCAATTTGAGC
|
Table 2:PCR reaction mixture
|
Reagents
|
Volume
|
|
Plasmid DNA
|
2 µl
|
|
10X PCR buffer
|
1.5 µl
|
|
Primer-1 (Forward)
|
1.5 µl
|
|
Primer-2 (Reverse)
|
1.5 µl
|
|
25mM MgCl2
|
0.8 µl
|
|
2mM dNTPs
|
0.75 µl
|
|
Taq polymerase(5U/ µl)
|
0.5 µl
|
|
Sterilized Water
|
11.45 µl
|
|
Total volume
|
20 µl
|
Transformation of the plasmid in Top10 F’ competent cells: The transformation process was carried out following Latif et al. 2015.
Plasmid DNA isolation and Clone confirmation: Plasmid DNA was isolated done through Thermo Fisher Scientific gene JET Plasmid Miniprep kit with cat # K0502. For clone confirmation, Fast digest restriction endonucleases Sma I and Pst I were used to digest isolated plasmid DNA Reaction mixture of 15 µl concentration was prepared as mentioned below in table and incubation was done at 37℃ for 05 minutes.
Table 3: Restriction digestion of transformants for gene presence.
|
Reagents
|
Volume
|
|
Plasmid DNA (500ng)
|
4 µl
|
|
Sma I
|
1 µl
|
|
Pst I
|
1 µl
|
|
FD Green Buffer
|
2 µl
|
|
H2O
|
7 µl
|
|
Total volume
|
15 µl
|
Transformation and confirmation of Top10 F’ culture on plates: Heat shock method was used for the transformation of isolated plasmid. Sample of 100 µl and 200 µl concentration was spread on Tetracycline and Kanamycin LB agar selection plates after incubation for 90 minutes followed by overnight incubation at 37℃. Next morning, colonies (overnight grown) on petri plates confirmed the transformation and to test their positivity some colonies were selected.
Transformation and confirmation of Agrobacterium culture on plates: Next morning, 3 YEP media containing plates with Kanamycin and rifampicin (50 µg/ml) selection were taken and culture of transformed Agrobacterium cells of 50 µl, 100 µl and 150 µl concentration were spread on plates respectively with the help of sterilized spreader. To confirm the presence of gene of interest in plasmid construct, colonies were selected for performing colony PCR. Gene specific primers for pta gene whose sequences are given before in Top10F’ transformation section.
Transformation of Gossypium hirsutum
Screening of different varieties, delinting of seeds, sterilization and soaking of seeds: Five varieties of cotton were tested for its growth ability and vigor. Among these 5 varieties CA-12 variety of Gossypium hirsutum was selected due its good growth and best health conditions. Delinting of seeds, sterilization and soaking of seeds were done according to (Ahmed et al. 2019).
Culturing and harvesting of Agrobacterium with pta gene: For culturing of Agrobacterium YEP broth of 10ml concentration with 10 µl of rifampicin and Kanamycin were added in each 50ml falcon tube. These tubes were placed in incubator at 30℃ for one day. Next day, centrifugation of cultures were done at 4℃ with speed of 5000 rpm. MS broth of 10 ml concentration was used for dissolving pellet that was formed. The tubes containing vector pCAMBIA2300, pta gene were mixed for transformation purposes.
Embryo cut method: After selecting germinated embryos, these embryos were cut at their plemule site with the aid of sterile sharp blade and then added to Agrobacterium tumefaciens containing MS broth medium. The Agrobacterium tumefaciens contain our gene of interest i.e. pta. After that this media was incubated for 2 hours with continuous shaking at 30℃. After taking out embryos from broth media, they were desiccated by placing them on sterile filter paper. Embryos were then placed on MS media containing plates. Shifting of plantlets into test tubes and shifting of cotton transgenic plants into soil was done as described earlier by (Puspito et al. 2015).
Molecular Analysis
Genomic DNA extraction and PCR Confirmation of transgenic plants: Transformed plants were subjected to DNA extraction (Puspito et al. 2015) for their molecular analysis. PCR was performed to detect the incorporated pta gene. DNA isolated from non-transgenic plants were taken as template for negative control while plasmid of pCAMBIA2300 was used as positive control. Gene specific primers for pta gene whose sequence are in Top10F’ transformation section.
Transgenic plants confirmation through southern blot analysis
Southern blotting: DNA was isolated form transgenic as well as non-transgenic cotton plants for analysis of southern blot which was defined earlier (Ahmed et al. 2019).
Probe labelling: For pta gene detection, probe was labelled by using Biotin DecaLabel DNA labelling Kit (Cat# K0652). Probe (labelled DNA) was assessed by comparing it with control probe (provided in kit) spotted on nitrocellulose membrane in sequential dilutions and lastly detection was done by means of Biotin Chromogenic Detection Kit (Cat# K0651).
RESULTS
Codon Optimization: Codon Adaptation Index (CAI) or codon usage frequency distribution of pta gene was adjusted lengthwise at level 0.95 to the gene sequence. A CAI of 1.0 is measured to be faultless in the anticipated expression organism, as well as a CAI of > 0.8 is considered as decent, in terms of great expression level of gene (Figure 1A). Frequency of Optimal Codons (FOP) was increased up to 74%. However, 26% of expressive codons could not be optimized as per molecular preferences of cotton cells mainly due fear of loss of function (Figure 1B). GC content was adjusted at average 38.52% for all stretch of codons.
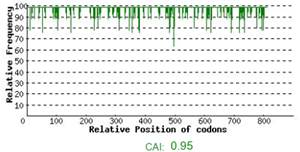 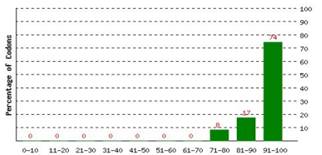

Figure 1: (A) Codon usage frequency distribution along the gene sequence length. A CAI of 1.0 is measured to be faultless in the anticipated expression organism, as well as a CAI of > 0.8 is considered as decent, in terms of great expression level of gene. (B) The codons percentage distribution in computed codon quality groups. The 100 value is set as highest usage frequency for the codon for a particular amino acid in the chosen expression organism. The perfect range of %age of GC content is among percent. (C) The %GC content peaks in a 60 base pairs window have been detached.
Integration of pta Gene Cassette Assemblyin pUC57: Recombinant pUC57 vector harbouring pta gene cassette was propagated and for this was transformed into E. coli Top10F’ competent cells. Miniprep Kit (GeneJET ™ Cat# K0503) was used for Plasmid DNA was isolation from selected colonies. It was found that restriction digestion resulted in two distinct fragments; one at ~ 2.7kb depicting pUC57 vector while the one at ~1.79kb depicted pta Gene Cassette Assembly (figure 2).
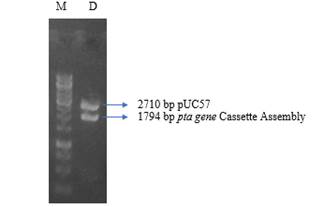
Figure 2: Restriction digestion of the plasmid DNA to reveal pta gene assembly in pUC57 vector. M = 1kb DNA Ladder, D = Digestion of pUC57 Plasmid isolated from E. coli Top10F’ cells grown on selection media.
Cloning of pta gene assembly cassette in plant binary vector, pCAMBIA2300: Plant binary vector, pCAMBIA2300 was propagated in E.coli cells. Further, the pta gene cassette was directionally cloned in pCAMBIA2300. Randomly six transformed colonies were selected, cultured and subjected to examination for the presence of pta Gene Cassette Assembly in pCAMBIA2300 vector. Mini-prep from five cultures yielded good quality plasmid (figure 3A). PCR examination was done on isolated five plasmids resulting all positive (figure 3B). PTA specific primers were applied via colony PCR for confirmation purpose. An amplified band of ~595 bp was observed on agarose gel. Plasmid from non-transformed E. coli cells was taken as negative control, while PCR confirmed pUC57 vector was used as positive control to validate results. Finally, restriction digestion with Sma I and Pst I restriction enzyme excised exact fragment size corresponding to ~1794 bp of pta Gene Cassette Assembly (figure 3C). This experiment confirmed the presence of pta Gene Cassette Assembly in pCAMBIA2300 vector rendering it applicable for further Agrobacterium transformation.

Figure 3: Detection and confirmation of pta Gene Cassette Assembly in pCAMBIA2300 through PCR and restriction digestion. Figure 5A: Plasmid DNA isolated from transformed E.coli Top10F’ cells (L is 1 kb DNA ladder, lane 1 to 6: plasmid DNA samples; figure 5B: amplification of pta Gene Cassette Assembly with gene specific primers(L is 1 kb plus DNA ladder, lane 1 to 5 are samples, lane 6 & 7 are control negative and control positive respectively); figure 5C: Restriction digestion of plasmid DNA for pta Gene Cassette Assembly in pCAMBIA2300 vector (L is 1 kb plus DNA ladder, lane 1 to 5 are digested plasmid with Sma I and Pst I restriction enzyme while lane 6 is undigested control negative plasmid).
Transformation of pCAMBIA_PGCA construct into Agrobacterium tumefaciens: Recombinant pCAMBIA_PGCA construct was successfully transferred to Agrobacterium tumefaciens (strain LBA4404) via electroporation. Positively electroporated Agrobacterium cells were analyzed for transgene cassette presence through PCR with specific primers. It was observed that a fragment of ~588 bp was amplified in 9 out of 10 screened colonies as shown in figure below.
Transformation of Gossypium hirsutum var. GS-1:For transformation of GS-1 variety of Gossypium hirsutum, four set experiments were done over the period of two months. Mature embryos were taken as explants. Approximately, 2235 healthy embryos were used for transformation by pCAMBIA_PGCA recombinant Agrobacterium clone through co-cultivation and tissue culture as described in material & methods section.
Healthy embryos were cutat their epicotyl end and exposed with Agrobacterium containing pCAMBIA_PGCA construct. To develop the roots, plantlets were placed in MS tubes for more four to six weeks. Plants were moved to soil pots after formation of roots (Figure 5 a-f).


Figure 5: Schematic overview of Agrobacterium mediated cotton transformation (a) Germinated seeds of desired cotton variety (b, c) Agrobacterium inoculated cotton embryos (d) Agrobacterium treated embryos growing on MS medium (e) Plants in MS tubes with developing roots and shoots (f) Putative transgenic plants shifted to soil for acclimatization.
Molecular Analysis of Putative Transgenic Cotton Plants (T0 Generation)
Conventional PCR with pta gene specific primers was performed to assure presence of pCAMBIA_PGCA gene cassette in T0 generation of cotton plants before acclimatization in soil.
Plant Genomic DNA Isolation and PCR:The DNA was isolated from 15 randomly selected plants according to protocol mentioned in chapter 3. DNA quality was checked by resolving it on 1% concentration of Agarose gel electrophoresis (Figure 6A). PCR was performed on extracted DNA from transformed cotton plants transformed with pCAMBIA_PGCA construct by specific primers (designed from pta gene sequence). Primers conditions were optimized with gradient PCR. Same profile was run and reagent concentration used as mentioned in previous chapter. Amplified fragments when resolved on agarose gel, revealed amplification of 595 bp pta gene fragment in 12 plants (P-11, P-34, P-46, P-48, P-52, P-59, P-62, P-9R, P-10R, P-11R, P-88R and P-109) (Figure 6B). While no amplification was detected in plant number P-51, P-54, P-60 and P-1R. Control -ve and control +ve samples validated results.
  
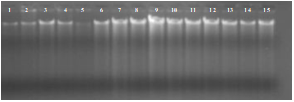 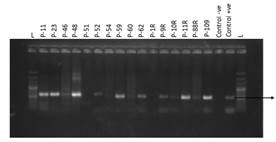
Figure 6: (A) Genomic DNA extracted from transformed cotton plants, Lane 1-15: DNA samples. (B) Molecular (PCR) detection of pta gene construct in transformed cotton plants. L represents 1kb Plus Ladder. Plant number P-11, P-34, P-46, P-48, P-52, P-59, P-62, P-9R, P-10R, P-11R, P-88R and P-109 showed a band of 595 bp as result of specific (designed from pta gene sequence) primers amplification.
Analysis of transgenic T1 progeny
Molecular Analysis of transgenic cotton plants of T1 progeny: Seeds from transgenic cotton plants (T0) were allowed to develop as in normal conditions and sowed again in contained field to obtain T1 generation. For each of twelve transgenic cotton lines (T1), three transgenic cotton plants were selected randomly for molecular screening including PCR and Southern blot to reveal transgene integration in transgenic cotton plants.
Conventional PCR: Leaves of the subject T1 transgenic cottonplants was taken, complete genomic DNA isolated and then was subjected to PCR using internal detection primers. PCR, through the pCAMBIA_PGCA specific primers, resulted in amplification of ~595 bp fragment in 5 transgenic cotton lines (P-48, P-109, P-10R, P-59, P-9R) (figure 7). While no amplification was detected in seven cotton lines that were positive at T0 stage along with non-transgenic plants (figure 7).
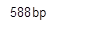 
Figure 7: Molecular (PCR) detection of pta gene in transformed cotton plants. Ladder: 1kb Plus Ladder; P-48, P-109, P-10R, P-59, P-9R are positive for pCAMBIA_PGCA; while the same was note detected in T1 generation of P-52, P-34, P-88R, P-11R, P-62, P-46 and P-11. Control +ve and control -ve validated the results.
Southern Blotting:Confirmation of pCAMBIA_PGCA transgenic integration in transgenic cotton lines (T1 generation) was done by Southern blot analysis. PCR positive transgenic cotton lines (P-48, P-59, P-9R, P-10R and P-109R) were found positive for transgene integration as they develop signals in chromogenic reaction when probed with DIG-labelled PTA specific probe (Figure 8)
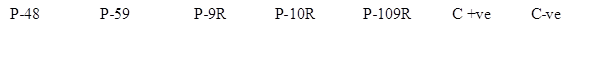 
Figure 8: Southern blot analysis of the PTA gene in transformed cotton plants. As expected after PCR analysis, confirmed pCAMBIA_PGCA positive plants P-48, P-59, P-9R, P-10R and P-109R positively blotted the chromogenic probe specific to pta gene region as shown in the figure. Control positive and control negative samples appeared as expected thus confirming the validity of experiment.
DISCUSSION
Cotton in subcontinent, once the source of "the costliest and handsomest” clothing fiber with superior advantage over wool and source of inspiration and class for the indigenous rulers, invading warriors and great travellers of the likes of Marco Polo and Vasco Da Gama, has taken a serious plunge into disaster in recent times. This crop is facing perhaps the biggest challenge in history in its native and most ancient 5000 BC old habitat Mehrgarh and Indus Valley civilizations currently identified as rich planes of Punjab and Sindh province in Pakistan (Ghosh and Ghosh 1995). In year 2019 alone, Pakistani cotton growing sector sustained a loss of 6 million bales worth approx. 3 billion USD denting already struggling national economic recession (Chohan et al. 2020). Experts blame abiotic and biotic factors, including irregular weather patterns and sucking/chewing/virus pests respectively, for these agronomically catastrophic outcomes. Among biotic stresses, whitefly is the prime one hitting the crop with its extensive sap-sucking and mediating infection of Cotton Leaf Curl Virus (CLCuV).
A natural necessity arose during last five years to develop a cotton plant that would be resistant or at least to some satisfactory degree tolerant against whitefly infestation. Conventional breeding techniques require extensive time to develop desired trait in plants, which consequently are over-run by extent of the demand. Therefore, genetic transformation of non-transgenic cotton plants was done through plant infection bacteria called Agrobacterium tumefaciens. There could be number of options in the process of selecting candidate genes from Cry or Lectins or other pest-toxic gene families but, considering its efficiency, ability to defy multiple biotic stresses and short sequence, agglutinin gene from Pinellia ternata or simple called Pinellia ternata agglutinin (pta) was chosen and transformed. It is important to continuously search for novel enzymes and transcript for plant improvement (Shah et al. 2019). Agglutinin proteins work on the principle of hemagglutination of cells in different biological systems. As discussed previously in the case of lectin proteins, the agglutinins use their mannose binding sites to develop clumps through bonding carbohydrate units employing specific recognition system (Chandra et al. 1999, Shridhar et al. 2009).
Transgenic expression of Allium sativum lectin or ASAL has reported to fight off cotton leafworm in tobacco plants (Sadeghi et al. 2008). PTA, the subject of this study, had also shown antifungal, insecticidal effects against anti‐pinewood nematode, Myzus persicae or peach potato when transformed and transgenically expressed in tobacco and other plants (Yao et al. 2003, Gaofu et al. 2008, Ling et al. 2010). However, there was no study or examination of PTA effects on whitefly when transformed in cotton plants.
For better understanding effects of the foreign gene, non-transgenic variety of Gossypium hirsutum called GS-1 was chosen. This variety is common among non-GM cotton growers of the southern Punjab region. From the applied aspects of this study, this variety had disadvantages over superior GM varieties. Better performing plants could have been achieved if PTA was transformed into varieties that were already made tolerant to chewing pest or weedicides like the variety CEMB-33 but in that case the current study would not have yielded exact functional characterization of pta gene alone in non-transgenic cotton plants. Also, it was important to set grounds to develop decisive strategy against whitefly which preys on other important sap rich vegetable crops in local farming conditions. Therefore, extensive and all necessary in-silico, in-vivo and contained field experimentation adopted to design a codon optimized (cotton targeted) pta gene cassette under the influence of well-tested CaMV35S promoter in pCAMBIA2300 expression vector enabled by agrobacterium LB4404 plant infecting strain for stable delivery and integration of transgene in the recipient genome (few results are still being analysed). Transgenic plants among the experimented lot were distinguished and identified differently at different stages. The selection began at tissue culture stage when MS media was supplemented with Kanamycin which only let the transformed plant grew. Rest of them couldn’t survive. After sufficient growth and acclimatization, DNA from leave tissues of putative plants were examined for presence of our transgene i.e. PTA through specific primers. This standardized procedure has been reported by multiple studies (Rao et al. 2011, Ahmed et al. 2019).
However, the gene delivery process remained as less efficient, tedious and industrious as reported by previous reports on cotton transformation. During the length of this study, which involved a total of 2235 embryos were used, 194 of them germinated successfully in MS media and 5 of them found positive in both T0 and T1 generation (transformation efficiency 0.53%). This pattern is no different from earlier studies (Puspito et al. 2015) and (Rao et al. 2011) who tried and transformed two different cotton varieties (VH-289 and CIM-482) respectively. However, fewer studies in literature has reported higher efficiency of gene transformation in cotton plants i.e. up to 60%, which is far from pattern recorded in this study. It is pertinent to mention that this efficiency is calculated considering all the plants which survived on MS media selection not the ones which got successfully acclimatized and subsequently found positive through PCR and southern blotting. The reason of low transformation may stem from the cotton variety, transformation method or tissue culturing techniques (Finer and McMullen 1990, MAJEED et al. 2000, Satyavathi et al. 2002, Chhalgri et al. 2020).
Compared to control negative cotton plants, whitefly population was found significantly retarded on the pta transformed plants due to strong transgenic expression (results of in-planta bioassay are under analysis and premature for reporting, yet). This observation was quite in line with an earlier study where aphid population was found controlled significantly in GNA transformed tobacco plants (Hilder et al. 1995). Evaluation of T1 progeny, (plant no. P-48, P-59, P-9R, P-10R and P-109R). Same pattern of Mendelian inheritance was reported in previous report (Khan et al. 2010, Rao et al. 2011, Ahmed et al. 2019).
This research employed single gene, but as the technological advancements are taking place in gene synthesis, plant artificial chromosome development and novel vector construction, scientists are more and more inclined toward multi-gene transformation approaches. The multi-gene or stacked gene construct may seem convincing in fighting pests through targeting the same problem with multiple dots but, in experience this strategy, is full of technical pitfalls (Hamilton et al. 1996, Houben et al. 2008). For example, one way for transgenes stacking, that is crossing of transgenic homozygous plant requires tedious, non-specific and lengthy process of classical breeding (De Cosa et al. 2001, Quesada-Vargas et al. 2005). Nonetheless, breeding proves advantageous to consolidate acquired trait in the advance generations of transgenic lines. Based on the experience and current technological standing, it is advisable to transform single gene construct and efforts should be made to achieve uniformity in expression in all progeny of the selected transformed event for better field/agronomic performance through crossing.
Pinellia ternata has been consumed (cooked/uncooked) as a traditional medicinal plant in China to treat respiratory disorder e.g. cough, for thousands of years (Shuhong and Zhengsong 2003, Shih et al. 2018). Determination of the exact active phytochemicals in folk medicinal supplements is growing field of knowledge and there is still a lot to uncover. It may be safe to assume that the PTA protein, sourced from already consumed plant, would impart no non-target effects on worms, bees or animals who consume secondary products derived from cotton. However, it is imperative to call for bio-safety studies of the PTA transformed cotton plants on all relevant biological models to have scientifically proven argument of safety. Biosafety always remains a challenge for newly developed transgenic entity casting the shadow of uncertainty, but this should not hinder the research and development of ever new strategies employing different presumably safe genes to achieve maximum crop protection against biotic stresses.
In conclusion, there has been nuclear transformation of pta gene in cotton plants achieved. The resultant plants may be enrolled in breeding programs with high yielding cotton varieties for further improvement in production character. Agronomic performance is a function of better crop protection and decreased cost of production. Immunity against whitefly serves both causes at the same time by reducing the need of pesticide sprays, labor requirement, loss to crop vigour and overall stress to plant immune system; and perhaps the indirect blocking leaf curl virus. Once applied and agronomically developed into stable variety, these transformed cotton plants may bring relief to severely struck cotton crop in the country.
References:
- Ahmed, M., S. Akhtar, M. Fanglu, M. Hasan, A. Shahid, X. Yanang, M. Sarwar, A. Rao, T. Husnain and X. Wang (2019). Sucrose Synthase (SuSy) Gene Expression: An Indicator for Cotton Fiber Initiation and Early Development. Russian J. Plant Physio. 66(1): 41-49.
- Chandra, N. R., G. Ramachandraiah, K. Bachhawat, T. K. Dam, A. Surolia and M. Vijayan (1999). Crystal structure of a dimeric mannose-specific agglutinin from garlic: quaternary association and carbohydrate specificity. J. mol. bio. 285(3): 1157-1168.
- Chhalgri, M. A., M. T. Khan, G. S. Nizamani, S. Yasmeen, I. A. Khan, M. M. Aslam, A. A. Rajpar, Tayyaba, F. Nizamani, M. R. Nizamani, R. Iqbal, M. J. Panhwar and M. A. Siddiqui (2020). Effect of Plant Growth Hormones on Shoot and Root Regeneration in Rose under In Vitro Conditions. Adv. Life Sci. 8(1): 93-97.
- Chohan, S., R. Perveen, M. Abid, M. N. Tahir and M. Sajid (2020). Cotton Diseases and Their Management. Cotton Production and Uses, Springer: 239-270.
- De Cosa, B., W. Moar, S.-B. Lee, M. Miller and H. Daniell (2001). Overexpression of the Bt cry 2Aa2 operon in chloroplasts leads to formation of insecticidal crystals. Nat. biotech. 19(1): 71-74.
- Finer, J. J. and M. D. McMullen (1990). Transformation of cotton (Gossypium hirsutum L.) via particle bombardment. Plant cell rep. 8(10): 586-589.
- Gaofu, Q., M. Shiqing, Z. Fayin, Y. Zhiniu and Z. Xiuyun (2008). In vitro assessment of plant lectins with anti-pinwood nematode activity. Journal of invertebrate pathology 98(1): 40-45.
- Ghosh, G. and S. Ghosh (1995). Indian textiles: past and present, APH Publishing.
- Hamilton, C. M., A. Frary, C. Lewis and S. D. Tanksley (1996). Stable transfer of intact high molecular weight DNA into plant chromosomes. Proc. Nat. Acad. Sci. 93(18): 9975-9979.
- Hilder, V., K. Powell, A. Gatehouse, J. Gatehouse, L. Gatehouse, Y. Shi, W. Hamilton, A. Merryweather, C. Newell and J. Timans (1995). Expression of snowdrop lectin in transgenic tobacco plants results in added protection against aphids. Trans. Rea. 4(1): 18-25.
- Houben, A., R. K. Dawe, J. Jiang and I. Schubert (2008). Engineered plant minichromosomes: a bottom-up success? The Plant Cell 20(1): 8-10.
- Khan, M. A. U., A. A. Shahid, A. Q. Rao, K. S. Bajwa, T. R. Samiullah, A. Muzaffar, I. A. Nasir and T. Husnain (2015). Molecular and biochemical characterization of cotton epicuticular wax in defense against cotton leaf curl disease. Iranian J. Biotech. 13(4): 3.
- Khan, N. U., K. B. Marwat, G. Hassan, S. B. Farhatullah, K. Makhdoom, W. Ahmad and H. U. Khan (2010). Genetic variation and heritability for cotton seed, fiber and oil traits in Gossypium hirsutum L. Pakistan J. Bot 42(1): 615-625.
- Latif, A., A. Q. Rao, M. A. U. Khan, N. Shahid, K. S. Bajwa, M. A. Ashraf, M. A. Abbas, M. Azam, A. A. Shahid and I. A. Nasir (2015). Herbicide-resistant cotton (Gossypium hirsutum) plants: an alternative way of manual weed removal. BMC Re. Not. 8(1): 453.
- Ling, L.-J., Y.-Z. Yang and Y.-R. Bi (2010). "Expression and characterization of two domains of Pinellia ternata agglutinin (PTA), a plant agglutinin from Pinellia ternata with antifungal activity." World J. Micro. Biotech. 26(3): 545-554.
- Loriato, V. A., L. G. Martins, N. C. Euclydes, P. A. Reis, C. E. Duarte and E. P. Fontes (2020). Engineering resistance against geminiviruses: a review of suppressed natural defenses and the use of RNAi and the CRISPR/Cas system. Plant Sci. 11: 410.
- Macedo, M. L. R., C. F. Oliveira and C. T. Oliveira (2015). Insecticidal activity of plant lectins and potential application in crop protection. Molecules. 20(2): 2014-2033.
- MAJEED, A., T. HUSNAIN and S. Riazuddin (2000). Transformation of virus-resistant genotype of Gossypium hirsutum L. with pesticidal gene. Plant Biotech. 17(2): 105-110.
- Naheed, R., M. Arfan, F. Farhat, S. Ijaz and H. Khalid (2020). Acclimatization of drought tolerance with Somaclonal variants of sugarcane (Saccharum officinarum L.). Adv. Life Sci. 8(1): 57-62.
- Navas-Castillo, J., E. Fiallo-Olivé and S. Sánchez-Campos (2011). Emerging virus diseases transmitted by whiteflies. Ann. Rev. Phytopath. 49: 219-248.
- Owczarzy, R., A. V. Tataurov, Y. Wu, J. A. Manthey, K. A. McQuisten, H. G. Almabrazi, K. F. Pedersen, Y. Lin, J. Garretson and N. O. McEntaggart (2008). DT SciTools: a suite for analysis and design of nucleic acid oligomers. Nucleic Acid. Re. 36(suppl_2): W163-W169.
- Perlak, F. J., R. L. Fuchs, D. A. Dean, S. L. McPherson and D. A. Fischhoff (1991). Modification of the coding sequence enhances plant expression of insect control protein genes. Proc. Nat. Acad. Sci. 88(8): 3324-3328.
- Pruitt, K. D., T. Tatusova and D. R. Maglott (2007). NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acid. Re. 35(suppl_1): D61-D65.
- Puspito, A. N., A. Q. Rao, M. N. Hafeez, M. S. Iqbal, K. S. Bajwa, Q. Ali, B. Rashid, M. A. Abbas, A. Latif and A. A. Shahid (2015). Transformation and Evaluation of Cry1Ac+ Cry2A and GTGene in Gossypium hirsutum L. Frontiers Plant Sci. 6: 943.
- Quesada-Vargas, T., O. N. Ruiz and H. Daniell (2005). Characterization of heterologous multigene operons in transgenic chloroplasts. Transcription, processing, and translation. Plant Physio. 138(3): 1746-1762.
- Rao, A. Q., M. Irfan, Z. Saleem, I. A. Nasir, S. Riazuddin and T. Husnain (2011). Overexpression of the phytochrome B gene from Arabidopsis thaliana increases plant growth and yield of cotton (Gossypium hirsutum). J. Zhejiang Uni. SCI. B. 12(4): 326-334.
- Rüdiger, H. and H.-J. Gabius (2001). Plant lectins: occurrence, biochemistry, functions and applications. Glyco. J. 18(8): 589-613.
- Sadeghi, A., G. Smagghe, S. Broeders, J.-P. Hernalsteens, H. De Greve, W. J. Peumans and E. J. Van Damme (2008). Ectopically expressed leaf and bulb lectins from garlic (Allium sativum L.) protect transgenic tobacco plants against cotton leafworm (Spodoptera littoralis). Trans. Res. 17(1): 9.
- Satyavathi, V., V. Prasad, B. G. Lakshmi and G. L. Sita (2002). High efficiency transformation protocol for three Indian cotton varieties via Agrobacterium tumefaciens. Plant Sci. 162(2): 215-223.
- Shah, S. R. U., M. Rashid, M. A. Jaffar, M. Aslam, H. Shah and G. Rasool (2019). Evolution of Phosphoenolpyruvate carboxylase encoding transcripts in Chickpea (Cicer arietinum L.). Adv. Life Sci. 6(4): 152-158.
- Shih, T. T., H. L. Lee, S. C. Chen, C. Y. Kang, R. S. Shen and Y. A. Su (2018). Rapid analysis of traditional Chinese medicine Pinellia ternata by microchip electrophoresis with electrochemical detection. J. Sep. Sci. 41(3): 740-746.
- Shridhar, S., D. Chattopadhyay and G. Yadav (2009). PLecDom: a program for identification and analysis of plant lectin domains. Nucleic Acids Res. 37(suppl_2): W452-W458.
- Shuhong, W. and P. Zhengsong (2003). Survey of Pinellia ternata and Its Adulterants. J.Chinese Med. Mat. 11.
- Tariq, A., H. Afrasiab and F. Farhat (2020). In vitro Micropropagation of Citrullus colocynthis (L.) Schrad: an endangered medicinal plant. Adv. Life Sci. 8(1): 52-56.
- Yao, J., Y. Pang, H. Qi, B. Wan, X. Zhao, W. Kong, X. Sun and K. Tang (2003). Transgenic tobacco expressing Pinellia ternata agglutinin confers enhanced resistance to aphids. Transgenic Res. 12(6): 715-722.
|





