DNA BASED GENDER IDENTIFICATION OF MEAT SAMPLES
M. Iqbal1, W. Shehzad1*, M. Imran1, M. Y. Zahoor1, W. A. Khan2, M. I. Rashid3, K. Ashraf 3, H. Rehman4, M. S. Saleem1 and A. Nadeem1
1Institute of Biochemistry and Biotechnology, Faculty of Biosciences; 2Department of Wildlife and Ecology, Faculty of Fisheries and Wildlife; 3Department of Parasitology, Faculty of Veterinary Sciences; 4Department of Physiology, Faculty of Biosciences, University of Veterinary and Animal Sciences, Lahore, Pakistan
Corresponding author wasim.shehzad@uvas.edu.pk
The paper was presented in International Buffalo Congress 2019, February 18-20, Lahore, Pakistan
ABSTRACT
Meat is an important component of our diet. Male meat is exclusively preferred for consumption. In order to control the quality of suppliers the identification of the meat gender is important for meat processors. The DNA based methods have proved, to be efficient for gender identification. Females (bovine and ovine) are used for reproduction and are normally slaughtered only when older, which lead to increase in length and rigidity of muscle fibers resulting in tougher meat of lower quality. To enable detection of male bovine meat samples, a PCR-assay was designed in the present study. The Amelogenin gene located on X and Y-chromosome has proved to be a good marker for gender identification even in cooked meat samples. A pair of primers was designed based on AMEL-X and AMEL-Y genes which would be able to identify animal gender of bovine and ovine meat samples. The accuracy and specificity of the primers was assessed using DNA template extracted from bovine and ovine meat samples of known gender. Established method was applied to 180 bovine and ovine meat samples. PCR products of meat samples after electrophoresis showed two bands (250, 180-bp) for male DNA sample while only single band (250bp) for female DNA Sample. Result shows that the PCR assay based on the Amelogenin gene is reliable for gender determination in meat. This technique is also of practical utility in biodiversity assessment in non-invasive studies. Gender identification plays an important economic role in animal production industries where females are requisite for the dairy industry while males are preferred for beef production.
Keywords: Amelogenin, gender identification, non-invasive.
https://doi.org/10.36899/JAPS.2020.6.0157
Published online August 03,2020
INTRODUCTION
Consumers prefer low fat and high quality products (Todaro et al., 2004). Sensory attributes are not only having great importance for consumers, producers, and retailers but also to personals interested in meat quality and technology. Meat from males is of greater flavor quality, juiciness and general acceptability than meat from females (Rodrigues and Teixeira 2009). Meat from female animal is significantly less tender than male (Hanzelková et al., 2011).
Gender determination of meat and its products has gained great awareness in recent years (Branscheid and von Lengerken, 1998; Palanisamy et al., 2012). An excellent example exists in identification of beef, where male beef is considered to be of higher value than meat from heifer or cow (Pfeiffer and Brenig 2005) and therefore results in high price meat and meat products (Robertson and Gemmell 2006). At present, a variety of techniques have been developed for determining the gender of meat, mainly based on detecting hormone or DNA (Bai et al., 2010). In broad terms, hormone-based techniques comprise determination of immuno-chemicals using enzyme-linked immune-sorbent assay and mass-spectrometric detections by different chromatographic techniques like High Performance Liquid Chromatography (Lago et al., 2011) and Gas chromatography–mass spectrometry (Herrero et al., 2013). Most of these techniques provide precise and accurate results but have many disadvantages as variation in the composition of protein, even in the same source, minimize the chances of success of these procedures (Kadivar et al., 2013). Furthermore, these techniques require expensive equipment and cannot be performed on regular basis for gender identification (Joo et al., 2013).
Molecular tests provide improved diagnostic capability in many fields of biology, including ecology, evolutionary and conservation biology (Musliji et al., 2015) and have the advantages of being less time consuming in certain situations (use of noninvasive samples instead of laborious tracking down of the animal), require smaller samples and easier to perform (Asma et al., 2017). Mostly male gender is identified by using the SRY gene (sex-determining region Y) which is a Y chromosome-specific sequence (Mara et al., 2004). The disadvantage of this method is that if there is no amplification, it cannot be assumed that the individual is female or that there was a mistake in the experimental process (Langen et al., 2010). A gene present in both, males and females, should be amplified in the same tube as internal control. The amelogenin (AMEL) genes, located on both X and Y chromosomes of mammals, usually shows length polymorphism between AMEL-X and AMEL-Y (Palanisamy et al., 2012). PCR assays based on AMEL have facilitated gender identification in many mammalian species (Hrovatin and Kunej 2018; Li et al., 2018). In present study PCR assay is designed to identify gender of meat and its products. This developed method will be helpful for authentication of the meat source male or female.
MATERIALS AND METHODS
Sample selection: One hundred and eighty meat samples were collected from different butcher shops, markets and restaurants in and around Lahore to investigate the gender of meat and its products. In addition to these unknown meat samples, male and female reference samples were also collected from animals slaughtered at Eid al-Adha to confirm the reliability of proposed assay. These samples were kept at -20⁰C until further use.
DNA extraction of meat tissue samples: Samples were taken from interior muscle tissue using a sterile surgical blade. All the samples were homogenized and DNA was extracted from 100 mg of sample. The DNA was extracted from meat samples using DNeasy Blood and Tissue kit (QIAGEN, Germany) according to manufacturer instructions. Quantification of genomic DNA was performed using Nanodrop spectrophotometer 2000 (Thermo Fisher Scientific, Waltham, MA, USA).
Primer designing: The Primer pair for gender identification was designed using Primer3 software v0.4.0 (Untergasser et al., 2012) targeting conserved regions of Amelogenin gene for X and Y chromosome. The primers were designated as AMEL-F 5’- CAGCCAAACCTCCCTCTGCC -3’and AMEL-R: 5’- TTGGTCTTGTCTGTTGCTGGCCA-3’ with complete homology among X and Y chromosome (Bajwa et al., 2020). The primers were expected to amplify PCR product of 250bp with the AMELX gene as target sequence and 180bp with AMELY gene, respectively.
PCR amplification and thermocycling protocols: Strict precautions against PCR contamination were taken, including the use of aerosol-resistant pipette tips for handling the solutions. Polymerase chain reaction (PCR) was performed in 20μl of reaction mixture containing 50–100ng of genomic DNA, 10μl premix solution 2XTOPsimpleTM DyeMIX-n Taq containing 0.4mM of each dNTP, nTaq buffer with 3mM Mg+2, 5pmoles of each primer, 0.2units/μl nTaq DNA Polymerase (Enynomics, USA) andnuclease-free water to make a final volume. Mock reactions with water were also run parallel to monitor any contamination during amplification process. The PCR reactions were performed using the thermocycler (BIO-RAD) PCR System (PE Applied Biosystems, USA). Amplicons were obtained using an initial denaturation of 95◦C for 5 min; followed by 45 cycles of denaturation at 94◦C for 30 s, primer annealing at 62◦Cfor 30 s, and polymerase extension at 72◦C for 30 s; and a final extension step at 72◦C for 7 min. Electrophoresis of PCR product was performed at 90 V on 2% agarose gel (Thermo Fisher Scientific, USA)using 1x TAE buffer (8.0). Fifty base pair gene ruler DNA ladder (Thermo Scientific, USA) was used as size marker.PCR products were visualized under UV trans-illuminator.
RESULTS
Amplification by PCR using pair of primer based on amelogenin gene yielded expected 180bp and 250bp PCR products with 100ng genomic DNA per reaction (Fig. 1). Females were expected to yield one band of 250bp where as male’s two bands of 180bp and 250bp. Amplification was specific as non-specific amplification was not observed.
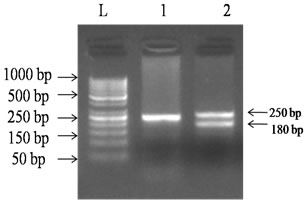
Fig .1 PCR assays using AMEL-X and AMEL-Y primers for gender identification. Targeted 250bp products were amplified from X- chromosome and 180bp products were amplified from Y- chromosome. Lane 1 contains female bovine DNA and Lane 2 contains male bovine DNA. L = 50bp ladder.
Blind tests were also performed to verify the accuracy and reliability of method. Gender determination by purposed method was in concurrence with actual gender of standard DNA samples extracted from meat tissues. Results are presented in Table.1 which indicates that gender identification method by PCR is 100% (10/10) accurate and reproducible.
Table 1.Comparisons of results of gender identified by PCR method and gender of standard sample collected from different slaughter house.
|
Meat samples
|
Gender of standard samples
|
Gender identified by PCR
|
|
1
2
3
4
5
6
7
8
9
10
|
M
M
F
M
F
F
M
M
M
F
|
M
M
F
M
F
F
M
M
M
F
|
|
*F = Female
*M = Male
|
|
|
Good bands of male and female genomic DNA were observed by decreasing the DNA concentration up to 0.5ng. No results were observed at concentration below 0.3ng per PCR reaction (Fig.2 A and B).
Fig. 2. PCR reaction to check the sensitivity of developed test with decreasing concentrations of DNA template. A. Lane 1 to 8 contains female bovine DNA at 100, 50, 20, 10, 2, 1, 0.5, 0.3 ng per reaction of template DNA, respectively. B. Lane 1 to 8contains male bovine DNA at 100, 50, 20, 10, 2, 1, 0.5, 0.3 ng per reaction of template DNA, respectively. L lanes contain 50bp ladder.
After assuring the specificity, sensitivity and accuracy of developed test, assay was applied for screening of 180 bovine and ovine meat samples to identify the male and female gender. Out of 180 of total meat samples 96 were of bovine (cattle and buffalo) and 84 were of ovine (sheep and goat). The screening of these samples by developed test showed that 37 (38.54 %) were identified as male and 59 (61.45%) as female out of 96 bovine. Screening of ovine meat samples results in 31 (36.9 %) female and 53 (63.09 %) male as indicated in Fig.3.

Fig.3 Screening results of gender identification, showing male and female proportions on x-axis and number of meat samples on y-axis.
Discussion and conclusions: The assay proposed here for gender identification to assure male bovine meat was applied to uncooked and cooked meat samples and the identification results were in perfect concurrence with anatomical gender of the meat sample. Amelogenin gene (AMEL) was selected as a target gene for gender identification. This gene is found on X-chromosome (AMELX) and as well as Ychromosome (AMELY) (Iwase et al., 2003). Amelogenin is used for gender discrimination in mammals on the basis of length polymorphism between AMELX and AMELY gene. This polymorphism is due to 63bp deletion in exon 5 of AMELY gene as compare to AMELX homologue. This deletion enables Amelogenin gene to discriminate between male and female gender by giving banding pattern of different sizes (Gibson 1999; Palanisamy et al., 2012). Due to this polymorphism Amelogenin gene is used for sex identification in different mammals like humans (Gibbon et al., 2009), horses (Hasegaw et al., 2000), cattle (Palanisamy et al., 2012; Weikard et al., 2006){Weikard, 2006 #97;Palanisamy, 2012 #157} and goats or sheep (Phua et al., 2003). We observed similar findings as reported by (Gokulakrishnan et al., 2012). However, the two articles are differing in terms of sample size i.e., 16 in case of Gokulakrishnan et al. (2012) and 180 in the present study. The primer set suggested in the given study targeted only cattle, while in this study; we have targeted bovines and ovines species, using a single meta-barcode primer pair. Further, we have exploited both cooked and uncooked samples in our study for the validation of the technique. Moreover, the present study also validates the proposed technique both for the samples of known and unknown genders.
A pair of primer was developed by targeting the Y-chromosome sequence which amplifies the PCR product from male source only (Mara et al., 2004). Disadvantage of this assay was inability to differentiate between female identification and failures of PCR (Schön et al., 2009). Another primer pair was used to discriminate between male and female DNA as amplification control (Fontanesi et al., 2008). As a result, double PCR reactions were required for single sample. In order to minimize the expenses of gender identification there should be a single pair of primer (Langen et al., 2010). Identification of gender was performed to ensure the quality of meat for food processors (Abdulmawjood et al., 2012). Analysis of hormone in meat and meat products for gender determination by Gas chromatography–mass spectrometry is good technique, but not comparable with PCR based procedures which are cost effective and rapid more over hormones are heat sensitive (Gokulakrishnan et al., 2015). Gender identification of meat is important to control meat quality and protecting the consumer’s interests. In order to achieve the target of study PCR assay was developed using pair of primers which yielded PCR products of 250bp and 180bp in case of male DNA and only single band of 250bp in case of female DNA. The pair of primers developed for present assay is gender specific as their sequences located in highly conserved regions and also it serves as internal PCR control to discriminate between female and male DNA source with accuracy in single PCR reaction per sample. In sensitivity test no PCR product was observed at DNA concentration 0.3ng/ul. But when number of cycles was increased to 50, results were positive which shows that developed test is highly specific and sensitive. The primers designed for this study can also be used for identification of gender of other animals which is helpful in biodiversity assessment as well as conservation studies. Precise gender determination is very important in food authentication, captivity breeding, wildlife management and evolutionary studies (Centeno-Cuadros et al., 2017).
Out of 180 meat samples only 68 were identified as male and 112 were identified as female. These results indicate high proportion of female meat, these findings are contrary to the statement of meat supplying personnel as they sell the meat by assuring us that the meat is from male source according to “The Punjab Animals Slaughter Control Act, 1963”. This law prohibits the slaughtering of female animals used for reproduction and dairy purposes. Now days, consumers are health conscious and demand high quality food products. They prefer low fat meat which is required to sustain its flavor and juiciness (Sarriés and Beriain 2006). The gender of animal had evident effect on quality of the meat, particularly on tenderness, water content, intramuscular fat deposition (Choat et al., 2006; Zhang et al., 2010). In female animals, lipid profile of the meat found to be less favorable to consumer health (Araujo et al., 2017).The results of present study show high rate of fraudulence in meat supply which indicate poor meat trade. There is need for establishment of quality control unit in slaughter houses to assure superior quality meat for consumer. The technique developed in present study can be helpful to increase the volume of meat export which depends upon authentication of good meat quality. Furthermore, justification of food fraud, quality control and fair trade increase consumer’s interest toward food in terms of health and economics.
Acknowledgment: The study was funded by National Research Program for Universities by Higher Education Commission, Pakistan [Project Number: 6790].
REFERENCES
- Abdulmawjood, A., C. Krischek, Wicke, and G. Klein (2012). Determination of pig sex in meat and meat products using multiplex real time-PCR. Meat Sci. 91: 272-276.
- Araujo, T., E. Pereira, I. Mizubuti, Campos, Pereira, E. Heinzen, H. Magalhaes, L. Bezerra, L. Pinheiro da Silva, and R. Oliveira (2017). Effects of quantitative feed restriction and sex on carcass traits, meat quality and meat lipid profile of Morada Nova lambs. J. Anim. Sci. Biotechnol.8: 38-46.
- Asma Ul, H., M.A. Awan, Mehmood, T. Sultana, Q. Shahzad, M.S. Ansari, B.A. Rakha, S.M. Saqlan Naqvi, and S. Akhter, (2017). Sperm sexing in Nili-Ravi buffalo through modified swim up: Validation using SYBR(R) green real-time PCR. Anim. Reprod. Sci. 182: 69-76.
- Bai, W.L., R.H. Yin, J. Zhao, C. Li, Z.J. Ma, R.L. Yin, G.B. Luo, and Z.H. Zhao (2010). A PCR assay for sex determination of yak (Bos grunniens) meat by amplification of the male-specific SRY gene. Food Control. 21: 726-731.
- Bajwa, A.A., S. Islam, M. Imran, K. Ashraf, A. Khan, M.F. Khan, Y. Zahoor, W.A Khan, and W. Shehzad (2020). DNA-Based Gender Identification of Punjab Urial (Ovis vignei punjabiensis) using Non-Invasive Sampling. Pakistan J. Zool. vol. 52,.
- Branscheid, W., and G. von Lengerken (1998). Die Erfassung der Schlachtkörperzusammensetzung und die Einstufung in Handelsklassen. W., editor. Qualitat von Fleisch und Fleischwaren. Frankfurt am Main (Germany): Deutscher Fachverl, 97-163.
- Centeno-Cuadros, A., I. Abbasi, and R. Nathan (2017). Sex determination in the wild: a field application of loop-mediated isothermal amplification successfully determines sex across three raptor species. Mol. Ecol. Resour. 17: 153-160.
- Choat, W., J. Paterson, B. Rainey, M. King, G. Smith, K. Belk, and Lipsey (2006). The effect of cattle sex on carcass characteristics and longissimus muscle palatability. J. Anim. Sci. 84: 1820-1826.
- Fontanesi, L., E. Scotti, and V. Russo (2008). Differences of the porcine amelogenin X and Y chromosome genes (AMELX and AMELY) and their application for sex determination in pigs. Mol. Reprod. Dev. 75: 1662-1668.
- Gibbon, V., M. Paximadis,. G. Štrkalj, P. Ruff and C. Penny (2009). Novel methods of molecular sex identification from skeletal tissue using the amelogenin gene. Forensic. Sci. Int. 3: 74-79.
- Gibson, C.W (1999). Regulation of amelogenin gene expression. Crit. Rev. Gene Exp. 9: 45-57.
- Gokulakrishnan, P., R. Kumar, B. Sharma, S. Mendiratta, D. Sharma, and O. Malav (2012). Sex determination of cattle meat by polymerase chain reaction amplification of the amelogenin AMELXAMELY gene. Vet. World. 5: 526-529.
- Gokulakrishnan, P., R.R. Kumar, B.D. Sharma, S.K. Mendiratta, O. Malav, and D. Sharma (2015). Determination of sex origin of meat and meat products on the DNA basis: a review. Crit. Rev. Food. Sci. Nutr. 55:1303-1314.
- Hanzelkova, S., J. Simeonovova, D. Hampel, A. Dufek, and J. Šubrt (2011). The effect of breed, sex and aging time on tenderness of beef meat. Acta. Veterinaria Brno. 80: 191-196.
- Hasegaw, T., F. Sato, N. Ishida, Y. Fukushima, and H. Mukoyama (2000). Sex determination by simultaneous amplification of equine SRY and amelogenin genes. J. Vet. Med. Sci. 62: 1109-1110.
- Herrero, B., L.J. Royo, F.C. Lago, J.M. Vieites, and M. Espiñeira (2013). Authentication of male beef by multiplex fast real-time PCR. Food. Addi. Cont. A. 30: 218-225.
- Hrovatin, K., and T. Kunej, (2018). Genetic sex determination assays in 53 mammalian species: Literature analysis and guidelines for reporting standardization. Ecol. Evol. 8: 1009-1018.
- Iwase, M., Y. Satta, Y.Hirai, H. Hirai, H. Imai, And N. Takahata (2003). The amelogenin loci span an ancient pseudoautosomal boundary in diverse mammalian species. Proc. Nat. Acad. Sci. 100: 5258-5263.
- Joo, S.T., G.D. Kim, Y.H. Hwang, and Y.C. Ryu (2013). Control of fresh meat quality through manipulation of muscle fiber characteristics. Meat Sci. 95: 828-836.
- Kadivar, A., H. Hassanpour, P. Mirshokraei, M. Azari, K. Gholamhosseini, and A. Karami (2013). Detection and quantification of cell-free fetal DNA in ovine maternal plasma; use it to predict fetal sex. Theriogenology 79: 995-1000.
- Lago, F.C., B. Herrero, M. Madriñán, J.M. Vieites, and Espiñeira (2011). Authentication of species in meat products by genetic techniques. Euro. Food Res, Tech. 232: 509-515.
- Langen, M., U. Peters, U. Korner, C. Gissel, Stanislawski, and G. Klein (2010). Semiquantitative detection of male pork tissue in meat and meat products by PCR. Meat. Sci. 86: 821-824.
- Li, Q., Y. Wang, L. Tan, J. Leng, Q. Lu, S. Tian, S. Shao, C., Duan, Li, and H. Mao (2018). Effects of age on slaughter performance and meat quality of Binlangjang male buffalo. Saudi. J. Biol. Sci. 25: 248-252.
- Mara, L., S. Pilichi, A. Sanna, C. Accardo, B. Chessa, F. Chessa, M. Dattena, G. Bomboi, and P. Cappai (2004). Sexing of in vitro produced ovine embryos by duplex PCR. Mol. Reprod. Dev. 69: 35-42.
- Musliji, Z., M. Svetozarevic, T. Nestorovski, B. Tanaskovska, Z.and Popovski (2015). Different approaches in the identification of meat origin based on protein profiling and simple PCR. Int. Sci. 70: 27-31.
- Palanisamy, G., R. Kumar, B. Sharma, S. Mendiratta, D. Sharma, and O. Malav (2012). Sex determination of cattle meat by polymerase chain reaction amplification of the amelogenin (AMELX/AMELY) gene 1. Vet. world. 5:526-529.
- Pfeiffer, I., and B. Brenig (2005). X- and Y-chromosome specific variants of the amelogenin gene allow sex determination in sheep (Ovis aries) and European red deer (Cervus elaphus). BMC genetics 6: 16-21.
- Phua, A.C.Y., R.B. Abdullah, and Z. Mohamed (2003). A PCR-based sex determination method for possible application in caprine gender selection by simultaneous amplification of the Sry and Aml-X genes. J. Reprod. Dev. 49: 307-311.
- Robertson, B.C., and N.J. Gemmell (2006). PCR-based sexing in conservation biology: Wrong answers from an accurate methodology? Cons. Gen. 7: 267-271.
- Rodrigues, S., and A. Teixeira (2009). Effect of sex and carcass weight on sensory quality of goat meat of Cabrito Transmontano1. J. Anim. Sci. 87: 711-715.
- Sarriés, M.V., and M.J. Beriain (2006). Colour and texture characteristics in meat of male and female foals. Meat Sci. 74: 738-745.
- Schön, S., T. Wilke, and R. Erdmann (2009). Molecular biological sex determination of processed porcine in fresh meat and meat preparations. Fleischwirtschaft 89: 48-48.
- Todaro, M., A. Corrao, M.L. Alicata, R. Schinelli, P. Giaccone, and A. Priolo (2004). Effects of litter size and sex on meat quality traits of kid meat. Small Rum. Res. 54: 191-196.
- Untergasser, A., I. Cutcutache, T. Koressaar, J. Ye, B.C. Faircloth, M., Remm, and S.G. Rozen (2012). Primer3—new capabilities and interfaces. Nucleic Acids Res. 40: e115-e115.
- Weikard, R., C. Pitra, and C., Kühn (2006). Amelogenin cross-amplification in the family Bovidae and its application for sex determination. Mol. Reprod. Dev. 73: 1333-1337.
- Zhang, Y.Y., L.S. Zan, H.B. Wang, Y.P. Xin, C. Adoligbe, and J. Ujjan (2010). Effect of sex on meat quality characteristics of Qinchuan cattle. Afr. J.Biotechno. 9: 4504-4509.
|





