Characterization of Cucurbit Yellow Stunting Disorder Virus Associated with Yellowing Disease of Watermelon in Saudi Arabia
H. Ahmad1, M. A. Al-Saleh1, I. M. Al-Shahwan1, M. T. Shakeel3, Y. E. Ibrahim1 and M. A. Amer1, 2, *
1 Plant Protection Department, College of Food and Agriculture Sciences, King Saud University, P. O. Box 2460, Riyadh 11451, Saudi Arabia. 2Viruses and Phytoplasma Research Department, Plant Pathology Research Institute, Agricultural Research Center, Egypt. 3Department of Plant Pathology, Bahauddin Zakariya University, Multan, Pakistan.
*Corresponding author’s E-mail: ruamerm@ksu.edu.sa
ABSTRACT
During the spring season of 2014, watermelon leaves were exhibiting yellowing, mottling and stunting symptoms in Western and Southwestern regions (Jeddah, Al-Lith, Jezan, Asfan, Ghonfada, Tofeel, Wadi Baish, Abu Arish), Saudi Arabia. One hundred thirty-nine samples were collected from symptomatic and asymptomatic watermelon plants. DAS-ELISA was performed to detect Cucurbit yellow stunting disorder virus (CYSDV) and other suspected cucurbit viruses. The obtained results revealed that out of 139 watermelon samples, nine and 98 samples were found to be positive with CYSDV and WmCSV, respectively and 22 samples were found to have mixed infection for both viruses. While no reaction was detected with, any of the other antisera tested against. Approximately 465 bp product were amplified by RT-PCR when CYSDV-Partial heat shock protein (HSP70) specific primer were used with ELISA positive symptomatic watermelon samples. Dot blot hybridization was also performed to detect CYSDV in the samples using CYSDV specific cDNA probe. Asymptomatic plant did not react when RT-PCR and dot blot hybridization assay were used. Partial HSP70 sequences of four CYSDV isolates were determined. When these HSP70 sequences were compared with other CYSDV isolates reported worldwide, the highest identity was between CYSDV KSA isolates (33-CYSDVKSA, 36-CYSDVKSA) and isolate from Portugal. Lowest identity was found between (34-CYSDVKSA, 35-CYSDVKSA), four Spanish isolates and one isolate from each USA and Lebanon. This is the first comprehensive study involving this virus in different regions in Saudi Arabia.
Key words: Cucurbit yellow stunting disorder virus, immunoblot and indirect enzyme-linked immunosorbent assay, reverse-transcription polymerase chain reaction, Partial heat shock protein.
https://doi.org/10.36899/JAPS.2020.5.0138
Published online June 25, 2020
INTRODUCTION
The cultivated species of Cucurbitaceae or gourd family are collectively called cucurbits. Among them, watermelon (Citrullus lanatus L.) is a valuable cash crop grown in temperate and tropical regions worldwide. It is placed in the genus Citrullus of the botanical family Cucurbitaceae. Watermelon is native to southern Africa, perhaps where Botswana is currently located (Robinson and Decker-Walters, 1997; Steven and Donald, 2012). CYSDV is a member of the genus Crinivirus (family Closteroviridae) (Martelli et al., 2000). Viral particles are flexible rods with lengths of 750–800 nm (Liu et al., 2000) that encapsidate two molecules of single-stranded RNA of positive polarity, known as RNA 1 and 2 (Aguilar et al., 2003). It is transmitted naturally by the whitefly B. tabaci in a semi-persistent manner. First detection of Cucurbit yellow stunting disorder virus (CYSDV) was in the United Arab Emirates in 1982 (Hassan and Duffus, 1991) and, since then, it has been reported in the Mediterranean region (Celix et al., 1996) including Egypt, Jordan, Spain, Turkey (Sese et al., 1994; Wisler et al., 1998; Cohen and Ben-Joseph, 2000, Papayiannis et al., 2005; El-Rahmany et al., 2014), Lebanon (Abou-Jawdah et al., 2000), Portugal (Louro et al., 2000), Morroco (Desbiez et al., 2000), USA (Kao et al., 2000; Kuo et al., 2007; Polston et al., 2008), China ( Liu et al., 2010) and Saudi Arabia (Rubio et al., 1999, 2001), where it has caused major economic damage to cucurbit crops. CYSDV has a narrow host range limited to species of Cucurbitaceae (Celix et al., 1996) in which it is believed to remain limited to phloem-associated cells. This phloem-limited virus causes serious damage to cucurbit crops with an approximated yield reduction of 30-50% (Hassan and Duffus, 1991; Celix et al., 1996; Karasev, 2000; Lopez-Sese and Gomez-Guillamon, 2000; Hourani and Abou-Jawdah, 2003; Wintermantel et al., 2009).
Cucurbit crops infected by CYSDV, show severe yellowing symptoms that start as an interveinal mottle on the older leaves and intensify as leaves age (Abou-Jawdah et al., 2000). Chlorotic mottling, yellowing and stunting occur on cucumber (Louro et al., 2000) and yellowing and severe stunting on melon (Kao et al., 2000). Berdiales et al. (1999) reporting the natural infection have provided no description of symptoms on squash. Symptoms on cucurbit crops are said to be indistinguishable from those caused by Beet pseudoyellows virus (BPYV) and Cucurbit chlorotic yellows virus (CCYV) (Wisler et al., 1998). Although it produces symptoms similar to other members of the closteroviridae family, CYSDV can be distinguished by molecular techniques and serology (Rubio et al., 1999). Amplification of heat shock protein 70 (HSP70) homologous gene has allowed reverse-transcription polymerase chain reaction (RT-PCR) detection of CYSDV in plants (Celix et al., 1996). Closterovirus degenerate primers have also been used along with RT-PCR to generate, clone, and characterize cDNAs from CYSDV for use in detecting plant infections (Tian et al., 1996). Oligonucleotide primers have been designed based on the CYSDV, allowing the use of RT-PCR and hybridization assays for detection of CYSDV (Livieratos et al., 1998). The complete CYSDV CP gene has been cloned and purified and used to develop antiserum. As a result, reliable immunoblot and indirect enzyme-linked immunosorbent assay (ELISA) like tests have been developed for detecting CYSDV in infected plant extracts (Livieratos et al., 1999).
Recently, a yellowing disease were observed in many fields of watermelon in the western and southwestern regions of Saudi Arabia. A careful inspection of symptoms indicated they were similar to those previously described to be caused by CYSDV (Bananej et al., 2002). Since this virus is an important pathogen that infects cucurbits and produces yellowing disease, its etiology needs to be investigated and its presence needs to be confirmed. The objective of this work is to characterize the Saudi Arabian isolates of CYSDV using biological, serological, molecular methods and determine the phylogenetic relationship between these isolates and other isolates reported worldwide.
MATERIALS AND METHODS
Source of virus isolates and whitefly transmission: During the spring seasons of 2014 and 2015, a total of 139 samples from watermelon were collected from asymptomatic and symptomatic plants showing virus-like symptoms including yellowing, mosaic, mottling and stunting (Figure 1) from Western & Southwestern regions, Saudi Arabia (Figure 2). Thirty-eight samples were collected from Al-Lith, 28 from Wadi Baish, 18 from Jeddah, 19 from Tofeel, 13 from Abu Arish, 8 from Asfan, 10 from Ghonfada, and 5 samples were collected from Jezan region (Table 1). These samples were kept in plastic bags and preserved at 4oC for short term while for long-term storage -80oC freezer storage was used (Nelson and Bushe, 2006). To carry out transmission experimetns, whitefies were collected from several greenhouses and identified as B. tabaci by Shakeel et al. (2018). Healthy watermelon seedlings were grown in insect free environment until three leave stage was attained. A single watermelon seedling per pot per cage was maintained to restrict the whiteflies to the targetted leaves and each treatment was replicated six times. a total of 60 whiteflies were allowed to feed on CYSDV positive watermelon plant for 48 hours of acquisition access period (AAP) and later used to inoculate healthy watermelon seedling for an inoculation access period (IAP) of 48 hours. A set of five watermelon seedlings were inoculated by 60 aviruliferous whiteflies for a 48hrs IAP as a control. Inoculated plants were sprayed with Acetamiprid to kill the whiteflies and older leaves were cut off to avoid any hatching of whitefly eggs and unwanted virus transmission. These plants were kept in insect free cages and symptoms were recorded three weeks post inoculation and later were tested by RT-PCR for a period of three weeks to observe symptom development and testing by Reverse transcriptase chain reaction (RT-PCR).
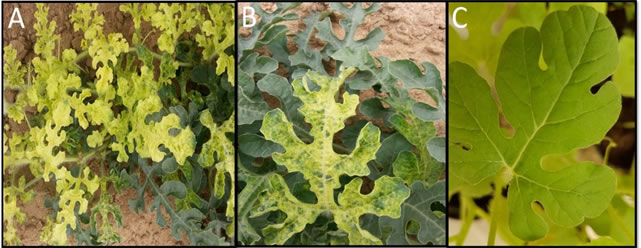

Figure 1: (A, B): natural symptoms observed in watermelon plants showing yellowing and mottling symptoms and C: uninfected watermelon as a negative control.
ELISA tests: Commercial ELISA kits for cucurbit viruses (AC diagnostic, USA) were used to serologically identify the viruses infecting watermelon. These polyclonal antisera including: Cucumber mosaic virus (CMV), Cucumber green mottle mosaic virus (CGMMV), Watermelon mosaic virus (WMV), Zucchini yellow mosaic virus (ZYMV), Squash mosaic virus (SqMV), Watermelon chlorotic stunt virus (WmCSV) Cucurbit yellow stunting disorder virus CYSDV as described by Clark and Adams (1977).
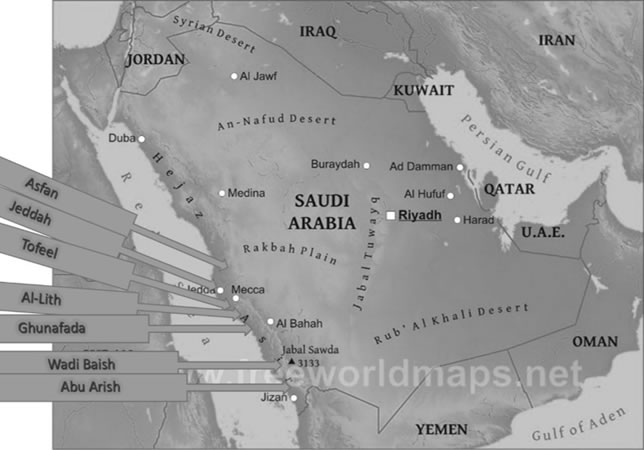
Figure 2. Different locations visited during the field survey in western and southwestern regions of Saudi Arabia.
Total RNA extraction and RTPCR: For CYSDV identification, total RNA was extracted from four selected DAS-ELISA positive leaf samples representing the four main regions (Jeddah, Al-Leith, Tofeel, Wadi Baish) using the SV Total RNA Isolation System (Promega Co., USA). RT-PCR was carried out based on the results of DAS-ELISA test using specific primers (CYSDV1:5′-acg agg atc cag aga cgg taa gta tgt c-3′ and CYSDV2:5′-gtt gga att ctt ggg cat gtg aca tag ag-3′, Livieratos et al., 1998). targeting heat shock protein 70 (HSP70) to amplify a 465bp region of CYSDV genome. The targeted region was amplified using MyTaqTM One-Step RT-PCR Kit (Bioline, United Kingdom) in a thermal cycler (Eppendorf, Germany) according to manufacturer’s protocol and the amplified products were visualized as described by Sambrook and Russell (2001) and photographed using DNA documentation gel analysis (IN GENIUS, Syngene Bio Imaging, UK). DNA Ladder 100 bp RTU (Genedirex) was used to determine the size of PCR amplified DNA products.
Molecular hybridization for detection of CYSDV: The probe for CYSDV was synthesized using PCR DIG Probe Synthesis Kit (Roche) through PCR amplification. Selected samples were prepared for dot blotting on nitrocellulose membrane according to Podleckis et al., 1993; Pallas et al., 1998; Ruiz et al., 2002. Prehybridization, hybridization, and colorimetric detection were carried out using a hybridization oven (Amersham Biosciences, Piscataway, NJ, USA) following the protocol recommended by Boehringer Mannheim (Germany). The results were documented by photographing the wet filter.
Nucleotide sequence and phylogenetic analysis for CYSDV-HSP70 gene: Selected four RT-PCR products for CYSDV-HSP70 infecting watermelon representative to four regions (Jeddah, Al-Leith, Tofeel, Wadi Baish) were purified using AxyPrep DNA Gel Extraction Kit from Axygen Biosciences (USA) according to the manufacturer's instructions and sequenced in both directions at BGI Tech Solutions, Hong Kong, China. The obtained nucleotide sequences for Saudi isolates of HSP70 gene for CYSDV were subjected to the BLASTn and were compared to the seven published sequences retrieved form GenBank. Multiple sequence, phylogenetic relationship reconstructed for these isolates were analyzed using Lasergene DNASTAR, V5-05 Software.
RESULTS
Virus Isolates and Serological Identification: Under field conditions, naturally infected watermelon plants showed typical yellowing and mottling symptoms on leaves, collected from different locations. To identify the causal virus associated with those symptoms developed on watermelon plants grown under field conditions; samples of symptomatic plants were collected from Western & Southwestern regions, extracted and serologically tested using DAS-ELISA. Results of DAS-ELISA showed that, CYSDV antiserum reacted positively with 9 tested extracts out of 139. Results also revealed that out of 139 watermelon samples and 98 were WmCSV positive, 22 samples had mixed infection of both viruses (Table 1). While no reaction was detected with, any of the other antisera tested against CMV, CGMMV, WMV, ZYMV and SqMV. ELISA is considered positive when the absorbance of the tested sample is at least double the absorbance of the control. When the transmission was done using B. tabaci biotype B whitefly, inoculated plants showed typical virus symptoms and were tested positive by DAS-ELISA for CYSDV positive three weeks post inoculation.
Table 1. Distribution of CYSDV affecting watermelon in different locations using DAS-ELISA test.
|
Location
|
No. of Samples
|
WmCSV
|
CYSDV
|
Mixed Infection
(WmCSV and CYSDV)
|
|
Al-Lith
|
38
|
25
|
4
|
6
|
|
Wadi Baish
|
28
|
23
|
1
|
1
|
|
Jeddah
|
18
|
8
|
3
|
5
|
|
Tofeel
|
19
|
14
|
1
|
3
|
|
Abu Arish
|
13
|
12
|
0
|
1
|
|
Asfan
|
8
|
6
|
0
|
2
|
|
Ghonfada
|
10
|
6
|
0
|
3
|
|
Jezan
|
5
|
4
|
0
|
1
|
| |
139
|
98
|
9
|
22
|
RT- PCR identification: ELISA positive samples from four locations (Jeddah, Al-Leith, Tofeel, Wadi Baish). were successfully amplified by the RT-PCR reaction (Figure 3). The 465 bp fragments of the RT-PCR product obtained by 1% agarose gel electrophoresis using specific primers was corresponding to the expected size of the HSP70 gene of CYSDV. The obtained results revealed that no RT-PCR products were amplified from total RNA extracted from uninfected watermelon sample.
Molecular hybridization for detection of CYSDV: Out of one hundred and thirty-nine samples tested with nucleic acid hybridization assay against CYSDV, thirty-four samples were found to be positive for CYSDV. Figure 4 Showed the results of molecular hybridization using dot-blotting assay of selected watermelon samples collected from different four locations using CYSDV DIG cDNA probe from Al-Leith (Row: A), Jeddah (Row: B), Tofeel (Row: C) and Wadi Baish (Row: D) locations. RT-PCR product DNA as positive control (P). No hybridization reaction was observed with healthy watermelon samples (N).
Nucleotide Sequence and Phylogenetic Analysis for CYSDV-HSP70 gene: Partial sequences of HSP70 gene for four Saudi Arabian CYSDV isolates were determined and submitted to GenBank under the accession numbers KY123231, KY123232, KY123233 and KY123234 for 33-CYSDVKSA, 34-CYSDVKSA, 35-CYSDVKSA and 36-CYSDVKSA isolates respectively (Table 2). The obtained results showed that all isolates including Saudi Arabian isolates fell within two distinct groups. These results showed that group A contains the seven different isolates obtained from GenBank while group B contains the Saudi isolates (Figure 5). The nucleotide sequence identity among the four Saudi Arabian isolates was 100%. While between these isolates and other isolates from GenBank aligned, HSP70 ranged between 89.8 and 90.3%. The highest similarity (90.3%) between isolates 33-CYSDVKSA, 36-CYSDVKSA and isolate (AF287474) from Portugal, while the lowest similarity (89.8%) was found between isolates 34-CYSDVKSA, 35-CYSDVKSA and four Spanish isolates (AJ223619, AY242078, AJ439690, U67170) and one isolate from each USA (FJ492808) and Lebanon (KC633820).
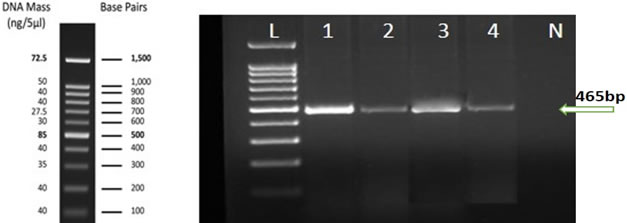
Figure 3. Agarose gel (1%) electrophoresis analysis shows amplified RT-PCR product (465 bp) for CYSDV using specific primers for HSP70 gene from selected symptomatic watermelon collected from Al-leith (Lane 1), Wadi Baish (Lane 2), Jeddah (Lane 3) and Tofeel (Lane 4) locations. No RT-PCR amplification was observed in uninfected sample tissue (lane N). Lane L: 100 bp DNA Ladder RTU Genedirex.

Figure 4. Dot blot hybridization with CYSDV (B) DIG-cDNA probe. Selected samples of watermelon collected from Al-leith (A), Jeddah (B), Tofeel (C) and Wadi Baish (D), locations. Nitrocellulose membranes indicating positive (purple) & negative (Colorless) results. No hybridization reaction was observed with uninfected watermelon samples (D: 9). PCR product DNA as positive control (D: 10).

Figure 5. Phylogenetic tree, based on HSP70 sequences of eleven isolates, shows two clads A &B. Clad A having seven isolates while clad B having four isolates.
Table 2. Percentage identity based on HSP70 sequences of eleven CYSDV isolates after aligning them by cluster W method.
|
Accession No:
|
Country
|
Host
|
Saudi Isolate
|
|
KY123231
|
KY123232
|
KY123233
|
KY123234
|
|
AJ223619
|
Spain: Almeria
|
Cucumis melo
|
90.0
|
89.8
|
89.8
|
90.0
|
|
AF287474
|
Portugal
|
C. sativus
|
90.3
|
90.0
|
90.0
|
90.3
|
|
AY242078
|
Spain
|
Melon
|
90.0
|
89.8
|
89.8
|
90.0
|
|
AJ439690
|
Spain: Almeria
|
C. sativus
|
90.0
|
89.8
|
89.8
|
90.0
|
|
FJ492808
|
USA
|
Melon
|
90.0
|
89.8
|
89.8
|
90.0
|
|
KC633820
|
Lebanon
|
C. sativus
|
90.0
|
89.8
|
89.8
|
90.0
|
|
U67170
|
Spain
|
Melon
|
90.0
|
89.8
|
89.8
|
90.0
|
|
KY123231
|
Saudi Arabia
|
Citrullus lanatus
|
100
|
100
|
100
|
100
|
|
KY123232
|
Saudi Arabia
|
C. lanatus
|
100
|
100
|
100
|
100
|
|
KY123233
|
Saudi Arabia
|
C. lanatus
|
100
|
100
|
100
|
100
|
|
KY123234
|
Saudi Arabia
|
C. lanatus
|
100
|
100
|
100
|
100
|
DISCUSSION
CYSDV is considered important disease of cucurbits in many parts of the world including Middle-East, Mediterranean Basin, Lebanon, Egypt, North Africa, Southern Europe and the Canary Islands (Abou-Jawdah et al. 2000; Kao et al., 2000; Kheyr-pour et al., 2000; Louro et al., 2000; Bananej et al., 2002; El-Rahmany et al., 2014). The natural hosts of CYSDV are restricted to the Cucurbitaceae including watermelon, melon and cucumber. In addition, the following experimental host plants have also been identified: Cucurbita maxima and Lactuca sativa. It has been successfully spread to many cucurbit-producing regions throughout the world (Abou-Jawdah et al. 2000; Kheyr-pour et al. 2000). In Saudi Arabia, yellowing symptoms have been observed in cucurbits crop in open field and greenhouses (Al-Saleh et al., 2015 a, b; Shakeel et al., 2016). Similar symptoms are reported to be produced by BPYV, CCYV, and CYSDV (Celix et al., 1996, Wisler et al., 1998, Okuda et al., 2010). All of these viruses are transmitted by whitefly (Bemisia tabaci) and cause brittleness of the older leaves (Abrahamian and Abou-Jawdah, 2014) and the vector is well established in most of the regions of Saudi Arabia (Shakeel et al. 2018). Although the results showed that samples with mixed infection were significantly less than those that had single infection of WmCSV and CYSDV, whereas, in previous studies mixed infections with two or three whitefly transmitted viruses were very common in cucurbits fields in Lebanon, Palestine and Jordan (Al-Musa et al., 2011; Ali-Shtayeh et al., 2014). Hence, these viruses pose a greatthreat to cucurbit crops in Saudi Arabia. In the southern regions of Saudi Arabia, these crops are feb by rain water only and farmer doesn’t put much efforts to harvest a productive and disease free crop. Perhaps this is the reason why the disease incidence is so high. In most of these areas, farmers commonly grow watermelon throughout the season. The main purpose of molecular studies was to have an idea about the genetic makeup of Saudi isolates of CYSDV and their percentage identity and phylogenetic relationship with other isolates reported worldwide. Therefore, RT-PCR, which is specific and sensitive method for detection of plant viruses based on nucleic acid (Uehara-lchiki et al., 2013), was performed to diagnose the Saudi isolates of CYSDV. The presence of 465 (HSP70 region of CYSDV genome) base pair bands in all the subjected samples confirmed the presence of CYSDV in these samples. The identity of CYSDV of viral isolates was established using sequence analysis of HSP70 gene. The four CYSDV isolates have 100% similarity percentage among themselves, all of which are falling in clad B while other isolates obtained from Genbank were placed in clad A. CYSDV Saudi isolates are justifying the phenomena of two sub populations, Western and Eastern group, as the phylogenetic tree shows that Saudi Arabian isolates have a separate cluster referring to “Eastern subpopulation” showing a greater difference percentage with the isolates submitted from other parts of the world. These isolates shared highest percentage identity (90.3%) with an isolate from Portugal. A possible explanation for evolution of Saudi Arabian isolates into distinct eastern subpopulation may be attiributed to its survival in the harsh weather conditions that mantains very high temperature with low humidity since its first discovery in 2001 in Saudi Arabia (Rubio et al., 2001). They characterized the genetic variability of CYSDV isolates from different countries of the world through single-strand conformation polymorphism and nucleotide sequence analysis of the CP gene. Based on their results, they divided the isolates into two genetic groups: a ‘Western’ group containing samples from Spain, Jordan, Turkey, Lebanon, and North America (Rio Grande Valley of Texas and Mexico), and an ‘Eastern’ group containing samples from Saudi Arabia (Rubio et al., 1999, 2001). Both percentage similarity and phylogenetic position of all four CYSDV isolates clearly depicts that the virus is endemic to the region and has no evolutionary relationship to isolates reported from other regions and rule out the impact of host in the evolution of CYSDV.
Conclusion and Recommendations: CYSDV is the most important and damaging virus to cucurbits in Saudi Arabia. Four isolates were collected from watermelon and sequenced for the first time in Saudi Arabia using specific heat shock protein 70 (HSP70) homologous gene. Low genetic variabilities among Saudi isolates were detected in these samples. Because of their ability to establish in entirely different and extreme climatic conditions and a different set of plant species that play a role as overwintering host. To understand the epidemiology of CYSDV in Saudi Arabia, collection of a greater number of samples from different hosts including arable weeds need to be done and analyzed especially in other regions and locations which were not surveyed. Screening of commercial watermelon cultivars for the presence of resistance against CYSDV should be done. Complete genome sequencing and analysis of genetic diversity for CYSDV Saudi Arabian isolates should be done and Construction of infectious clones for this virus will help to find resistant cultivars against this virus. These studies it may provide a possible explanation to the reason for Eastern population of CYSDV to exist.
Acknowledgments: The authors would like to extend their sincere appreciation to the Deanship of Scientific Research, King Saud University, Saudi Arabia, for its funding of this research group no. RG-1438-065.
Compliance with ethical standards: All authors declare that they have no conflict of interest.
Ethical approval: This article does not contain any studies with human participants or animals performed by any of the authors.
REFERENCES
- Abou-Jawdah, Y., H. Sobh, A. Fayad, H. Lecoq, B. DelTcolle, and T.J. Trad-Ferr (2000). Cucurbit yellow stunting disorder virus -a new threat to cucurbits in Lebanon. J. Plant Pathol. 82:55-60.
- Abrahamian, P.E., and Y. Abou-Jawdah (2014). Whitefly-transmitted criniviruses of cucurbits: current status and future prospects. Virus 25: 26-38.
- Aguilar, J.M., M. Franco, C.F. Marco, B. Berdiales, E. Rodriguez-Cerezo, V. Truniger, and M.A. Aranda (2003). Further variability within the genus Crinivirus, as revealed by determination of the complete RNA genome sequence of Cucurbit yellow stunting disorder virus. J. Gen. Virol. 84: 2555-2564.
- Ali-Shtayeh, M.S., R.M. Jamous, O.B. Mallah, and S.Y. Abu-Zeitoun (2014). Molecular characterization of Watermelon chlorotic stunt virus (WmCSV) from Palestine. Viruses 6: 2444-2462.
- Al-Musa, A., G. Anfoka, A. Al-Abdulat, S. Misbeh, F. Haj Ahmed, and I. Otri (2011). Watermelon chlorotic stunt virus (WmCSV): A serious disease threatening watermelon production in Jordan. Virus Genes 43: 79-89.
- Al-Saleh, M.A., I.M. Al-Shahwan, M.A. Amer, M.T. Shakeel, A. Kamran, X. Christos, C. Orfanidou, and N.I. Katis (2015b). First report of Cucurbit aphid-borne yellows virus in cucurbit crops in Saudi Arabia. Plant Dis. 99: 894.
- Al-Saleh, M.A., I.M. Al-Shahwan, M.A. Amer, M.T. Shakeel, O.A. Abdalla, C.G. Orfanidou, and N.I. Katis (2015a). First report of Cucurbit chlorotic yellows virus in cucumber in Saudi Arabia. Plant Dis. 99: 734.
- Bananej, K., G. A. Dafalla, A. Ahoonmanesh, and A. Kheyr-Pour (2002). Host range of an Iranian isolate of Watermelon chlorotic stunt virus as determined by whitefly-mediated inoculation and agroinfection, and its geographical distribution. Journal of Phytopathology 150: 423-430.
- Berdiales, B., J.J. Bernal, E. Sáez, B. Woudt, F. Beitia, and E. Rodrfguez-Cerezo (1999). Occurrence of Cucurbit yellow stunting disorder virus (CYSDV) and Beet pseudo-yellows virus in cucurbit crops in Spain and transmission of CYSDV by two biotypes of Bemisia tabaci. Eur. J. Plant Pathol. 105(2):211-215.
- Celix, A., A. Lopez-Sese, N. Almarza, M.L. Gomez-Guillamon, and E. Rodriguez-Cerezo (1996). Characterization of Cucurbit yellow stunting disorder virus, a Bemisia tabaci transmitted Closterovirus. Phytopathology 86: 1370-1376.
- Clark, M.F., and A.M. Adams (1977). Characteristics of the microplate method of enzyme-linked immunosorbent assay for the detection of plant viruses. J. Gen. Virol. 34: 475-483.
- Cohen, S., and R. Ben-Joseph (2000). The dynamics of viruses affecting cucurbits in Israel: 40 years since 1960. Acta Hortic. 510: 321-325.
- Desbiez, C., H. Lecoq, S. Aboulama, and M. Peterschmitt (2000). First report of Cucurbit yellow stunting disorder virus in Morocco. Plant Dis. 84:596; 4.
- El-Rahmany, R.G., A.K. El Attar, H.S. Zein, N.A. Abdallah, and H.M. Mazyad (2014). Characterization of an Egyptian isolate of the Cucurbit yellow stunting disorder virus. Arab. J. Biotechnol. 17: 29-42
- Hassan, A.A., and J.E. Duffus (1991). A review of a yellowing and stunting disorder of cucurbits in the United Arab Emirates. Emir. J. Food Agric. 2: 1-16.
- Hourani, H., and Y. Abou-Jawdah. (2003). Immunodiagnosis of Cucurbit yellow stunting disorder virus using polyclonal antibodies developed against recombinant coat protein. Plant Pathol. 85: 197-204.
- Kao, J., L. Jia, T. Tian, L. Rubio, and B.W. Falk (2000). First report of Cucurbit yellow stunting disorder virus (genus Crinivirus) in North America. Plant Dis. 84:101; 2.
- Karasev, A.V. (2000). Genetic diversity and evolution of closteroviruses. Rev. Phytopathol. 38: 293-324.
- Kheyr-Pour, A., K. Bananej, G.A. Dafalla, P. Caciagli, E. Noris, A. Ahoonmanesh, H. Lecoq, and B. Gronenborn (2000). Watermelon chlorotic stunt virus from the Sudan and Iran: Sequence comparisons and identification of a whitefly-transmissible determinant. Phytopathology 90: 629-635.
- Kuo, Y.-W., M. R. Rojas, R. L. Gilbertson, and W. M. Wintermantel (2007). First report of Cucurbit yellow stunting disorder virus in California and Arizona, in association with Cucurbit leaf crumple virus and Squash leaf curl virus. Plant Dis. 91:330.
- Liu, H.Y., G.C. Wisler, and J.E. Duffus (2000). Particle lengths of whitefly-transmitted Criniviruses. Plant Dis. 84: 803-805.
- Liu, L. Z., Y. Chen, and W. M. Zhu (2010). First Report of Cucurbit yellow stunting disorder virus on Melon in China. Plant Dis. 94(4):485.
- Livieratos, I. C., A.D. Avgelis, and R.A. Coutts (1999). Molecular characterization of the Cucurbit yellow stunting disorder virus coat protein gene. Phytopathology 89: 1050-1055.
- Livieratos, I.C., N. Katis, and R.H. Coutts (1998). Differentiation between Cucurbit yellow stunting disorder virus and Beet pseudo-yellows virus by a reverse transcription-polymerase chain reaction assay. Plant Pathol. 47: 362-369.
- Lopez-Sese, A. I., and M. L. Gomez-Guillamon (2000). Resistance to Cucurbit yellow stunting disorder virus (CYSDV) in Cucumis melo L. Hortscience 35:110-113.
- Louro, D., M. Vicente, A.M. Vaira, and G.P. Accotto (2000). Cucurbit yellow stunting disorder virus (genus Crinivirus) associated with the yellowing disease of cucurbit crops in Portugal. Plant Dis. 84:1156.
- Martelli, G.P., A.A. Agranovsky, M. Bar-Joseph, D. Boscia, T. Candresse, R.H.A. Coutts, V.V. Dolja, J.E. Duffus, B.W. Falk, D. Gonsalves, W. Jelkmann, A.V. Karasev, A. Minafra, A. Murant, S. Namba, C.L. Niblett, H.J. Vetten, and N. Yoshikawa (2000). Family Closteroviridae. pp. 943-952. In: Van Regenmortel, M.H.V., C.M. Fauquet, D.H.L. Bishop, E.B. Carstens, M.K. Estes, S.M. Lemon, J. Maniloff, M.A. Mayo, D.J. McGeoch, C.R. Pringle, and R.B. Wickner (eds.) Seventh Report of the International Committee on Taxonomy of Viruses. New York & San Diego, Academic Press.
- Nelson, S.C., and B.C. Bushe (2006). Collecting plant disease and insect pest samples for problem diagnosis. Soil and crop management 14: 1-10.
- Okuda, M., S. Okazaki, S. Yamasaki, S. Okuda, and M. Sugiyama (2010). Host range and complete genome sequence of Cucurbit chlorotic yellows virus, a new member of the genus Crinivirus. Phytopathology 100: 560–566.
- Pallas, V., P. Mas, and J.A. Sanches-Navarro (1998). Detection of Plant RNA Viruses by Non-isotopic Dot-Blot Hybridization. pp. 461-468. In: Foster, G.D., & S.C. Taylor, (eds.) Methods in Molecular Biology. Springer. New York.
- Papayiannis, L.C., N. Ioannou, I.N. Boubourakas, C.I. Dovas, N.I. Katis, and B.W. Falk (2005). Incidence of viruses infecting cucurbits in Cyprus. J. Phytopathol. 153:530-535.
- Podleckis, E.V., R.W. Hammond, S.S. Hurtt, and A. Hadidi (1993). Chemiluminescent detection of potato and pome fruit viroids by digoxigenin-labeled dot blot and tissue blot hybridization. J. virol. Methods 43:147-158.
- Polston, J. E., L. L. Hladky, F. Akad, and W. M. Wintermantel (2008). First report of Cucurbit yellow stunting disorder virus in Cucurbits in Florida. Plant Dis. 92:1251.
- Robinson, R.W., and D.S. Decker-Walters (1997). Cucurbits. CAB International, Wallingford, UK.
- Rubio, L., J. Soong, J. Kao, and B.W. Falk (1999). Geographic distribution and molecular variation of isolates of three whitefly-borne closteroviruses of cucurbits: Lettuce infectious yellows virus, Cucurbit yellow stunting disorder virus, and beet pseudo-yellows virus. Phytopathology 89:707-711.
- Rubio, L., Y. Abou-Jawdah, H.X. Lin, and B.W. Falk. (2001). Geographically distant isolates of the Crinivirus Cucurbit yellow stunting disorder virus show very low genetic diversity in the coat protein gene. J. Gen. Virol. 82: 929-933.
- Ruiz, L., D. Janssen, L. Velasco, E. Segundo, and I.M. Cuadrado (2002). Quantitation of Cucurbit yellow stunting disorder virus in Bemisia tabaci (Genn.) using digoxigenin-labelled hybridisation probes. J. virol. Methods 101:95-103.
- Sambrook, J., and D.W. Russell (2001). Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbor: New York, NY, USA.
- A.L., M.L. Gomez-Guillamon, and J.R. Diaz-Ruiz (1994). Appearance of a possible new melon yellowing disease in Spain. Rep. Cucurbit Genet. Coop. 17: 72-73.
- Shakeel, M.T., M.A. Al-Saleh, M.A. Amer, I.M. Al-Shahwan, M.U. Umar, O.A. Abdalla, M.A. Zakri, and N.I. Katis (2018). Transmission of Cucurbit chlorotic yellows virus (CCYV) by whitefly biotype B in riyadh, Saudi Arabia. Int. J. Agric. Biol. 20: 241-248.
- Shakeel, M.T., M.A. Al-Saleh, M.A. Amer, I.M. Al-Shahwan, N. Dimou, C.G. Orfanidou, and N.I. Katis (2016). Etiology of yellowing diseases affecting greenhouse grown tomato and cucumber crops and the identification of involved Bemisia tabaci [Abstract]. In: Proceedings of 2nd International Whitefly Symposium; Feb 14-19. Arusha, Tanzania. pp-66.
- Steven, A, S, and N.M. Donald (2012). Cucurbits. pp. 286-316. In: Debbie, R., F. Graham, O. John. (eds.) Crop Post-Harvest: Science and Technology. Blackwell Publishing Ltd.
- Tian, T., V.A. Klaassen, J. Soong, G. Wisler, J.E. Duffus, and B.W. Falk (1996). Generation of cDNA specific to Lettuce infectious yellows closterovirus and other whitefly transmitted viruses by RT-PCR and degenerate oligonucleotide primers corresponding to the Closterovirus gene encoding the heat shock protein 70 homolog. Phytopathology 86: 1167-1173.
- Uehara-lchiki , T. Shiba, K. Matsukura, T. Ueno, M. Hirae, and T. Sasaya (2013). Detection and diagnosis of rice-infecting viruses. Front Microbiol. 4: 289.
- Wintermantel, W. M., L. L. Hladky, A. A. Cortez, and E. T. Natwick (2009). A new expanded host range of Cucurbit yellow stunting disorder virus includes three agricultural crops. Plant Dis. 93:685-690.
- Wisler, G.C., J.E. Duffus, H.Y. Liu, and R.H. Li (1998). Ecology and epidemiology of whitefly-transmitted Closteroviruses. Plant Dis. 82: 270-280.
|





