Content for New Div Tag Goes Here
MORPHO-CHEMICAL DIVERSITY AND RAPD FINGERPRINTING IN WHITE FLESH GUAVA CULTIVARS
M. Usman1*, Q. Zaman1, B. Fatima1, I. A. Rana2 and F. S. Awan2
1Plant Tissue Culture Cell, Institute of Horticultural Sciences, University of Agriculture, Faisalabad-Pakistan 38040
2Centre of Agricultural Biochem. & Biotech. (CABB), University of Agriculture, Faisalabad-Pakistan 38040
Corresponding Author Email: m.usman@uaf.edu.pk
ABSTRACT
Different elite strains of Guava cultivars ‘Round’ and ‘Pyriform’ collected from a single locality were evaluated for phenotypic, physico-chemical and genetic variation. Phenotypic variability was observed in strains of ‘Round’ and ‘Pyriform’ cultivars in leaf and fruit physical traits. In ‘Round’ strains, variation in leaf shape and other traits were less compared to ‘Pyriform’ strains. Strains Gola No. 1 ‘G1’, Surahi ‘S’ and Large Surahi ‘LS’, had less seeds compared with other strains. Among ‘Round’ strain ‘G1’ and ‘Pyriform’ strain ‘LS’ were better for most of the physical traits. In chemical traits, TSS:TA and total sugars showed variation among strains and ‘Round’ strain Small Gola ‘SG’ and ‘Pyriform’ strain ‘S’ were better for TSS:TA and TS. Overall, non-reducing sugars were higher in ‘Round’ while other fruit physico-chemical traits were similar in both varieties. Five out of fifteen randomly amplified polymorphic DNA (RAPD) primers produced 37 bands. Molecular variance was higher (74%) among strains compared with varieties (26%). Different strains of ‘Round’ and ‘Pyriform’ shaped cultivars were clustered as two distinct groups in the dendrogram generated by UPGMA method. The primers K1, K4 and K7 were highly polymorphic for selected guava genotypes. This base line study could be helpful in germplasm characterization, varietal identification and development of sequence characterized amplified regions (SCAR) marker associated to different traits.
Keywords: Fruit size, fruit quality, Gola, Surahi, DNA marker.
https://doi.org/10.36899/JAPS.2020.2.0034 Published online March 02, 2020
INTRODUCTION
Guava (Psidium guajava L.) is widely grown nutraceutically important fruit crop in tropics and subtropics around the world. It is rich in vitamin C, B complex, flavonoids and dietary fiber (Prakash, 2002; Rai et al., 2010). Guava is being largely produced in India, Pakistan, Thailand, Brazil and Mexico. Guava production has increased 5-10 times in different leading countries in the last two decades. In Pakistan, Guava is ranked as 3rd most important tree fruit crop with annual production of 522 thousand tons out of 70 thousand hectares (Anonymous, 2016). Major crop share (80%) is being contributed by Punjab province. Despite Pakistan is the second largest producer of guava after India, its per hectare yield is stagnant (7-8 tons/ha) for the last 10 years. Less quantity and low-quality guava produce may be caused by lack of commercially successful clonal propagation system leading to the development of a seedling-based guava industry, lack of high-density plantations, poor germplasm resources, little focus on breeding and biotechnology interventions, wide spreading guava wilt disease and fruit fly. Collectively these elements lead to less production of domestic quality.
There are two main types of guava classified on the basis of fruit shape and commonly known as Gola (Round) and Surahi (Pyriform). Both these types have wide diversity in flesh color (red, pink, cream, white), skin color (pink, golden, cream and white) and fruit size (large, medium and small) (Sharma et al., 2010; Usman et al., 2012). Largely sexual propagation of Guava in Pakistan and prevalence of occasional cross pollination led to varietal degradation and higher heterogeneity in the progeny. Hence, great phenotypic diversity exists within the orchards (Adrees et al., 2010). Further, metaxenial effects on fruit quality by pollen parents has been reported (Usman et al., 2013). Together, these factors indicate existence of wide genetic diversity in the available strains which need to be investigated. Morphological and genotypic description of the available germplasm and development of genetic associations using molecular markers including random amplified polymorphic DNA (RAPD), restriction fragment length polymorphism (RFLP), inter-simple sequence repeat (ISSR) and simple sequence repeat (SSR) has been reported in different fruit crops including guava. Morphological characterization showed great variability and have not been effective to differentiate closely related guava germplasm (Chandra et al., 2004; Valera-Montero et al., 2016). Further, variability in the environmental conditions and climate changes could add towards phenotypic diversity. RAPD markers are easy to use for assessment of genetic diversity, construction of genetic linkage maps and essentially need no previous data. Primers less than ten nucleotides have been used for complex DNA fingerprinting profiles (Annoles et al., 1991). RAPD markers have been effectively used in almond (Martins et al., 2004), banana (Ray et al., 2006), guava (Ahmed et al., 2011; Valera-Montero et al., 2016), loquat (Fukuda et al., 2016) and citrus sp. (Biswas et al., 2010; Munankarmi et al., 2014). Hence, eight strains in two white fleshed cultivars Round and Pyriform were collected from single locality and were assessed for morphological, physico-chemical and genetic characterization using RAPD markers. Assessment of molecular variance and development of genotypic associations in these strains could be highly useful to determine genetic identity of genotypes, hybrids and for future biotechnology applications.
MATERIALS AND METHODS
Plant Materials and Morphological Characterization: Twenty-four bearing plants of uniform age (8-10 years) in eight different strains of Round and Pyriform cultivars (Table 1) were selected in Sharqpur region (31.2748° N 74.60° E) Punjab, Pakistan. Plant materials including leaf and fruit samples were collected from the selected plants for further evaluations following UPOV (1987) descriptors. A comprehensive phenotypic characterization was performed including tree, foliar and fruit traits.
Fruit Analysis
Physical traits: Ten fruits at market maturity were randomly selected from healthy plants and different physical traits were studied including fruit size (mm) viz. fruit length (FL), fruit diameter (FD); fruit weight FW (g), flesh weight FlW (g), seed cavity weight SCW (g) and seed cavity diameter SCD (mm).
Chemical traits: Selected fruits were peeled using sharp knife. Fruit juice was extracted from each sample and homogenized for chemical analysis. Juice total soluble solids TSS (°Brix) was estimated using refractometer Model RX 5000 (ATAGO, Japan). Hortwitz (1960) method was followed for determination of titratable acidity TA (%). TSS values were divided to the corresponding TA values and TSS:TA was calculated. Total sugars TS (non-reducing sugars NRS and reducing sugars RS) available in fruit juice were measured following Hortwitz (1960). Oxalic acid solution (0.4%) was added to dilute 10 ml of juice and 5 ml of filtered aliquot was titrated against light pink color end point. Ascorbic acid AA contents (mg per 100 ml juice) were determined by titration method (Ruck, 1969).
Data Analysis: Experiment was planned in randomized complete block design (RCBD) and treatment means were analyzed by Duncan’s Multiple Range Test using MSTAT-C software (Russell and Eisensmith, 1983; Damon and Harvey, 1987).
DNA Extraction, PCR Procedure and Data Analysis: Genomic DNA was extracted from fresh young guava leaf tissues using CTAB method (Doyle and Doyle, 1990) with some modifications following Khan et al. (2004). DNA was quantified using NanoDrop (ND-1000) spectrophotometer. The DNA was amplified using fifteen primers from K-series oligonucleotide decamer primers (GeneLink Co., UK) selected due to their higher polymorphic nature (Table 2). PCR conditions were optimized using 25-30 ng template genomic DNA, PCR buffer (10X), dNTPs 200 µM each, 2 mM MgCl2, 15 ng of each decamer primer, 2.5 µL gelatin and 1.5 units of Taq DNA polymerase (Fermentas, UK) in a total volume of 25 µL (Williams et al., 1990). The amplification was performed in a T-100 thermocycler (Bio-Rad, USA). The complete reaction consisted of 40 cycles, each cycle consisting of denaturation at 95°C for 1 min, primer annealing at 36°C for 1 min, extension at 72°C for 2 min, with an initial denaturation at 95°C for 5 min and a final extension at 72°C for 2 min, followed by cooling at 4°C. PCR products (5 µL) were separated on 1.2% (w/v) agarose (Biochem, France) gels in 0.5X Tris-Borate-EDTA buffer containing ethidium bromide (0.5 g) per ml of agarose and 1 Kb DNA Ladder (Fermentas, UK) was used as marker. The fingerprints were observed and photographed in Dolphin-Doc UV Transilluminator (Wealtec, Germany).
The amplified bands from all the primers which were found reproducible were counted manually as absent (0) and present (1). The binary data produced were aligned together for cluster analysis and developing similarity matrix using software Popgene32 (version 1.44) following Yeh et al., (2000) based on UPGMA (Nei, 1987). Analysis of molecular variance (AMOVA) was performed in GenAlEx 6.1 (Peakall and Smouse, 2006).
RESULTS
Morphological Diversity in Different Strains of Guava Cultivars: In cultivar Round, most of the strains showed erect tree habit except ‘WG’ which had spreading nature. Leaf shape variation (trullate) was less compared to Pyriform strains (Fig. 1, 2). Leaf twisting was absent in ‘GG’ and ‘G1’ while curvature of midrib was absent in ‘G1’. Leaf shape of base and tip was round to obtuse while other leaf traits showed less variability (Table 3). Fruit shape at stalk end was broadly rounded in all strains except ‘WG’ which was round. Neck was absent in all strains suggesting these strains as true Round shaped. Seeds were few in ‘G1’compared with other strains and seed size was medium. Strain ‘SG’ had larger seed size compared with other strains (Table 4).
In cultivar Pyriform strains, tree shape was spreading in nature in all strains except ‘RS’ which was erect. There was little variation in the stem color while leaf blade length and leaf blade width showed variation (Fig. 1, 2). Among different leaf traits, more variability was observed in leaf shape compared with other traits (Table 3). In fruit traits, higher diversity was observed in fruit shape, diameter of calyx cavity, flesh thickness in relation to core diameter and seed size. Strains ‘S’ and ‘LS’ had less seeds compared with ‘SS’ and ‘RS’ strains (Table 4).
Physico-chemical Diversity in Fruit of Different Strains of Guava Cultivars
Fruit size (mm) and seed cavity diameter (mm): Fruit length was higher in Pyriform strains ‘LS’ and ‘S’ compared with Round and other Pyriform strains whereas ‘RS’ fruit was even shorter in length than Round strains (Table 4, 5). Fruit diameter was higher in ‘LS’, ‘S’ and ‘G1’ strains while least diameter was observed in ‘SG’. Seed cavity diameter was less in ‘S’ and ‘WG’ strains and was maximum in ‘LS’ (Fig. 1, 2).
Fruit weight (g) and seed cavity weight (g): Among Round and Pyriform strains, fruit weight of ‘LS’, ‘S’ and ‘G1’ was significantly higher compared with other strains. The lowest fruit weight was found in ‘RS’ and ‘SG’ strains. Seed cavity weight was more in ‘S’ and ‘GG’ while ‘WG’ had the lowest cavity weight (Table 5, 6).
Total soluble solids (°Brix), titratable acidity (%) and total sugars (%): TSS was significantly higher in Round strains compared with Pyriform strains and it was the lowest in ‘SS’ and ‘RS’. TA was higher in ‘LS’ and ‘WG’ while it was the lowest in ‘S’ and ‘SG’. Sugar content was higher in ‘SS’, ‘G1’ and the lowest in ‘RS’ (Table 5, 6).
Overall, little variation was noted for mean physical and biochemical attributes among the Round and Pyriform cultivars (data not shown) except NRS which were higher in cultivar Round (3.82%) compared with Pyriform (3.08%).
Molecular Variance and Genotypic Characterization using RAPD Markers: Different guava strains were estimated for genetic diversity using 10 decamer oligonucleotide RAPD primers. Five out of fifteen primer sets produced highly polymorphic and reproduceable bands (30) showing great variability among strains and were used to develop systematic relationships. The genetic diversity analysis precision was directly related to number of polymorphic fragments amplified. Primer K1 showed higher polymorphism and produced more amplified fragments bands compared with other primers (Fig. 3). Size of the amplified fragments ranged from 150 bp to 2500 bp. Total molecular variance estimated based on allelic distance matrix for F-Statistics analysis was higher within varieties (among strains) compared with variance among Round and Pyriform varieties (Fig. 4). The developed dendrogram for genetic similarities clearly divided the guava populations into two main groups classified on the base of fruit shape viz. Round (Gola) and Pyriform (Surahi) strains. These findings indicate efficiency of RAPD markers in identifying fruit shape diversity among two main cultivars (Fig. 5). The similarity coefficients ranged from 0.594 to 1.000.Table 1. List of strains in different Guava Cultivars
Cultivars |
Strains |
Round |
Golden Gola |
‘GG’ |
|
Small Gola |
‘SG’ |
|
Gola No. 1 |
‘G1’ |
|
White Gola |
‘WG’ |
Pyriform |
Sadabahar Surahi |
‘SBS’ |
|
Round Surahi |
‘RS’ |
|
Large Surahi |
‘LS’ |
|
Surahi |
‘S’ |
Table 2. Synthetic oligonucleotide decamer random primers used for molecular diversity analysis
Primer Name |
Code |
Nucleotide Sequence (5’-3’) |
GL DecamerA-01 |
K1 |
CAGGCCCTTC |
GL DecamerA-02 |
K2 |
TGCCGAGCTG |
GL DecamerA-03 |
K3 |
AGTCAGCCAC |
GL DecamerA-04 |
K4 |
AATCGGGCTG |
GL DecamerA-05 |
K5 |
AGGGGTCTTG |
GL DecamerA-20 |
K6 |
GTTGCGATCC |
GL DecamerC-01 |
K7 |
TTCGAGCCAG |
GL DecamerC-02 |
K8 |
GTGAGGCGTC |
GL DecamerC-03 |
K9 |
GGGGGTCTTT |
GL DecamerC-04 |
K10 |
CCGCATCTAC |
GL DecamerC-05 |
K11 |
GATGACCGCC |
GL DecamerC-06 |
K12 |
GAACGGACTC |
GL DecamerD-01 |
K13 |
ACCGCGAAGG |
GL DecamerD-02 |
K14 |
GGACCCAACC |
GL DecamerD-03 |
K15 |
GTCGCCGTCA |
Table 3. Morphological diversity in tree, shoot and leaf traits of different strains of Guava cultivars Round and Pyriform
Organs |
Plant Traits |
Round strains |
Pyriform strains |
Golden
Gola |
Small
Gola |
Gola
No.1 |
White
Gola |
Sadabahar
Surahi |
Round
Surahi |
Large
Surahi |
Surahi |
Tree |
Attitude of branches |
Erect |
erect |
erect |
spreading |
spreading |
erect |
spreading |
spreading |
Young Shoot |
color of stem |
yellow green |
green |
yellow green |
green |
yellow green |
yellow green |
yellow green |
green |
Leaf |
Blade length |
Long |
short |
medium |
long |
medium |
medium |
short |
short |
|
Blade width |
medium |
narrow |
broad |
broad |
broad |
narrow |
broad |
broad |
|
Length/width ratio |
High |
low |
medium |
high |
medium |
medium |
low |
medium |
|
Shape |
Trullate |
obtrullate |
trullate |
trullate |
obtrullate |
trullate |
oblong |
ovate |
|
Curvature in cross section |
Strong |
strong |
strong |
strong |
strong |
strong |
medium |
present |
|
Twisting |
Absent |
present |
absent |
present |
absent |
absent |
absent |
absent |
|
Curvature of midrib |
present |
present |
absent |
present |
present |
absent |
absent |
absent |
|
Degree of curvature of midrib |
medium |
weak |
- |
medium |
weak |
- |
- |
- |
|
Variegation |
present |
absent |
present |
absent |
present |
absent |
present |
absent |
|
Color |
yellow green |
yellow green |
yellow green |
yellow green |
green |
green |
dark green |
yellow green |
|
Color of midrib on lower side |
reddish |
yellow |
yellow |
cream |
reddish |
cream |
reddish |
yellow |
|
Spacing of secondary veins |
Wide |
close |
medium |
medium |
wide |
wide |
medium |
close |
|
Relief of surface of upper side |
smooth |
smooth |
smooth |
smooth |
smooth |
smooth |
smooth |
smooth |
|
Pubescence on lower side |
Sparse |
medium |
sparse |
sparse |
sparse |
absent or very sparse |
absent or very sparse |
sparse |
|
Undulation of margin |
present |
present |
present |
present |
present |
present |
present |
present |
|
Degree of undulation of margins |
Weak |
strong |
weak |
strong |
weak |
weak |
strong |
strong |
|
Shape of base |
Obtuse |
rounded |
rounded |
obtuse |
obtuse |
rounded |
cordate |
obtuse |
|
Shape of tip |
rounded |
obtuse |
rounded |
obtuse |
obtuse |
obtuse |
rounded |
rounded |
Table 4. Morphological diversity in fruit traits of different strains of Guava cultivars Round and Pyriform
Fruit Traits |
Round strains |
Pyriform strains |
Golden
Gola |
Small
Gola |
Gola
No.1 |
White
Gola |
Sadabahar
Surahi |
Round
Surahi |
Large
Surahi |
Surahi |
Shape at stalk end |
broadly rounded |
broadly rounded |
broadly rounded |
rounded |
broadly rounded |
broadly rounded |
truncate |
truncate |
Width of neck in relation to that of fruit |
_ |
_ |
_ |
_ |
medium |
broad |
broad |
broad |
Color of skin |
Red |
pale yellow |
pale yellow |
pale yellow |
pale yellow |
pale yellow |
pale yellow |
pale yellow |
Relief of surfaces |
Smooth |
Bumpy |
bumpy |
rough |
smooth |
rough |
smooth |
rough |
Longitudinal ridges |
Present |
Absent |
present |
absent |
Absent |
absent |
present |
absent |
Prominence of longitudinal ridges |
medium |
_ |
weak |
_ |
- |
- |
weak |
- |
Longitudinal grooves |
Present |
Absent |
present |
absent |
present |
absent |
absent |
absent |
Size of sepal |
Small |
Medium |
medium |
small |
Small |
medium |
small |
small |
Diameter of calyx cavity in relation
to that of fruit |
Large |
Medium |
medium |
small |
medium |
large |
large |
small |
Ridged collar around calyx cavity |
inconspicuous |
conspicuous |
inconspicuous |
inconspicuous |
conspicuous |
inconspicuous |
inconspicuous |
inconspicuous |
Length of stalk |
Medium |
Long |
medium |
short |
Short |
medium |
medium |
medium |
Color of flesh |
Cream |
White |
white |
white |
White |
white |
white |
white |
Evenness of color of flesh |
Even |
Even |
even |
even |
Even |
even |
even |
mottled |
Discoloration of flesh after cutting |
Absent |
Present |
absent |
absent |
Absent |
absent |
present |
present |
Grittiness of outer flesh |
Present |
Absent |
absent |
absent |
Absent |
present |
absent |
present |
Thickness of outer flesh in relation
to core diameter |
Medium |
Medium |
thick |
thick |
Thick |
medium |
thick |
very thick |
Puffiness |
Absent |
Present |
absent |
absent |
Absent |
absent |
absent |
absent |
Juiciness |
Medium |
Juicy |
medium |
juicy |
Juicy |
medium |
juicy |
medium |
Number of seeds |
Many |
Many |
few |
medium |
medium |
medium |
few |
very few |
Seed size |
Medium |
Large |
medium |
medium |
medium |
large |
large |
medium |
Table 5. Genotypic variation in fruit physical and biochemical attributes in guava cultivar Round strains
Strains |
Fruit Size (mm) |
Fruit Weight (g) |
Seed Cavity
Diameter (mm) |
Fruit Quality |
FL |
FD |
FL:FD |
FW |
FlW |
SCW |
FlW:
SCW |
TSS
(°Brix) |
TA (%) |
TSS:
TA |
AA
(mg 100 ml-1) |
RS (%) |
NRS (%) |
TS (%) |
Golden Gola (GG) |
63.94 ± 1.77 ab |
63.46 ± 2.20 ab |
1.00
± 0.02b |
50.97 ± 4.00 b |
20.36
± 2.22 c |
30.71 ± 1.24 |
0.66
± 0.04 c |
35.04
± 1.13 a |
10.22 ± 0.85 |
0.690 ± 0.033 |
14.81
± 0.43 b |
170.73 ± 10.85 |
3.76 ± 0.22 |
4.08 ± 0.43 |
8.05
± 0.25 ab |
Small Gola (SG) |
55.61 ± 2.83 b |
52.49 ± 3.95 b |
1.05
± 0. 05 a |
40.31 ± 0.94 b |
18.51
± 0.98 cd |
22.16 ± 0.83 |
0.83
± 0.03 b |
30.90
± 1.80 ab |
9.84
± 0.77 |
0.610 ± 0.022 |
16.13
± 0.32 a |
157.11 ± 16.49 |
3.91 ±0.21 |
3.43 ± 0.08 |
7.46
± 0.28 ab |
Gola No.1 (G1) |
72.35 ± 1.88 a |
70.65 ± 3.31 a |
1.02
± 0.03 a |
75.98 ± 3.99 a |
48.98
± 3.02 a |
27.08 ± 3.73 |
1.80
± 0.07 a |
34.80
± 0.71 a |
9.61
± 0.49 |
0.655 ± 0.020 |
14.67
± 0.29 b |
140.10 ± 15.73 |
3.72 ± 0.16 |
4.49 ± 0.24 |
8.44
± 0.25 a |
White Gola (SG) |
66.83 ± 5.70 ab |
59.99 ± 2.63 ab |
1.11
± 0.06 a |
45.19 ± 6.73 b |
29.17
± 3.81 b |
16.19 ± 3.71 |
1.80
± 0.06 a |
26.97
± 1.41 b |
9.17
± 0.44 |
0.734 ± 0.041 |
12.52
± 0.34 c |
146.61 ± 04.02 |
3.72 ± 0.13 |
3.29 ± 0.17 |
7.18
± 0.10 b |
Means sharing similar letters in columns are statistically non-significant (P<0.05)
Table 6. Genotypic variation in fruit physical and biochemical attributes in guava cultivar Pyriform strains
Strains |
Fruit Size (mm) |
Fruit Weight (g) |
Seed Cavity
Diameter SCD |
Fruit Quality |
FL |
FD |
FL:FD |
FW |
FlW |
SCW |
FlW:
SCW |
TSS
(°Brix) |
TA (%) |
TSS:
TA |
AA
(mg 100 ml-1) |
RS (%) |
NRS (%) |
TS (%) |
Sadabahar
Surahi (SBS) |
67.19
± 3.04 b |
60.67
± 1.88 b |
1.10
c |
46.16
± 3.73 b |
23.59
± 2.70 c |
22.57
± 1.71 |
1.04
bc |
34.20
± 0.59 ab |
8.67
± 0.62 |
0.678 ± 0.050 |
12.78
± 0.335 c |
148.33 ± 14.87 |
4.29 ± 0.37 |
3.77 ± 0.40 |
8.25
± 0.56 |
Gola
Surahi (GS) |
39.99
± 0.92 c |
59.21
± 1.08 b |
0.67
d |
39.99
± 0.92 b |
13.76
± 0.73 d |
26.23
± 1.05 |
0.52
c |
37.67
± 0.57 ab |
8.63
± 0.32 |
0.642 ± 0.032 |
13.44
± 0.17 b |
140.83 ± 05.22 |
3.78 ± 0.20 |
2.58 ± 0.48 |
6.49
± 0.33 |
Large
Surahi (LS) |
91.73
± 0.62 a |
68.79
± 1.58 a |
1.33
a |
74.35
± 5.35 a |
48.19
± 1.73 a |
26.16
± 2.56 |
1.84
a |
39.43
± 3.01 a |
9.63
± 0.37 |
0.784 ± 0.022 |
12.28
± 0.19 d |
140.00 ± 12.14 |
3.93 ± 0.30 |
3.13 ± 0.37 |
7.22
± 0.09 |
Surahi (S) |
84.74
± 5.21 a |
68.75 ± 0.91 a |
1.23
ab |
65.61
± 4.02 a |
35.85
± 2.73 b |
29.76
± 1.65 |
1.20
b |
29.08
± 2.03 b |
9.10
± 0.97 |
0.618 ± 0.024 |
14.72
± 0.49 a |
142.19 ± 16.26 |
3.65 ± 0.38 |
2.83 ± 0.99 |
7.70
± 0.20 |
Means sharing similar letters in columns are statistically non-significant (P<0.05)

Fig. 1. Fruit and leaves of Guava cultivar Round (Gola) strains including A) Golden Gola B) White Gola C) Gola No. 1 and D) Small Gola from Sharqpur region

Fig. 2. Fruit and leaves of Guava cultivar Pyriform (Surahi) strains including A) Sadabahar Surahi B) Large Surahi C) Surahi and D) Round Surahi from Sharqpur region
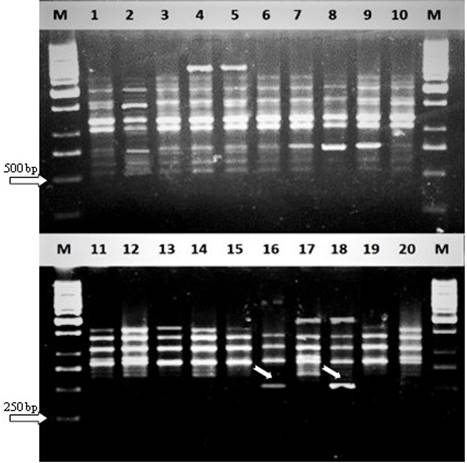 Fig. 3. RAPD markers produced in strains of Guava cultivars Pyriform and Round by primer set K1. Lanes show M= 1 Kb DNA ladder, 1-3 ‘SBS’, 4-6 ‘RS’, 7-9 ‘LS’, 10-11 ‘S’, 12-13 ‘GG’, 14-15 ‘G1’, 16-18 ‘WG and 19-20= SG. Arrows indicate unique bands (~500 bp) observed in cv. Round strain ‘WG’
Fig. 3. RAPD markers produced in strains of Guava cultivars Pyriform and Round by primer set K1. Lanes show M= 1 Kb DNA ladder, 1-3 ‘SBS’, 4-6 ‘RS’, 7-9 ‘LS’, 10-11 ‘S’, 12-13 ‘GG’, 14-15 ‘G1’, 16-18 ‘WG and 19-20= SG. Arrows indicate unique bands (~500 bp) observed in cv. Round strain ‘WG’

Fig. 4. Molecular variance (%) of Round and Pyriform guava cultivars as amplified with RAPD markers
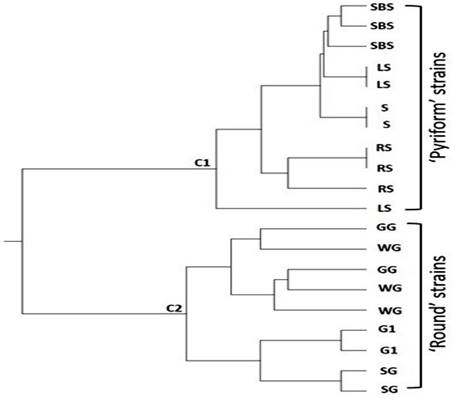
Fig. 5. Dendrogram showing relationship of different strains of Guava (Psidium guajava L.) cultivars Pyriform and Round based on similarity index. Pyriform strains included Sadabahar Surahi ‘SBS’, Large Surahi ‘LS’, Surahi ‘S’ and Round Surahi ‘RS’ (cluster 1) while Round strains included Golden Gola ‘GG’, White Gola ‘WG’, Gola No. 1 ‘G1’ and Small Gola ‘SG’ (cluster 2). Expected substitution per site is 10)
DISCUSSION
Guava orchards in Pakistan are mostly seedling based and show variability in different tree, foliar and fruit traits including size, shape and quality (Usman et al., 2012; 2013; 2018). Hence, there is huge potential of selection of elite strains for different fruit shapes, size, flesh color and fruit quality followed by clonal propagation (Pereira et al., 2017). Enormous foliage diversity found in different strains in the current study could be attributed to out breeding and sexual propagation. Number of seeds were less in strains ‘G1’, ‘S’ and ‘LS’ of Round and Pyriform cultivars. Different strains are classified by growers according to their size (small, medium and large) and shape (Round and Pyriform) under different names in different areas (Usman et al., 2018). The Round strains were truly round except ‘SG’ which showed little elongated fruit. In Pyriform strains, ‘RS’ had more FD compared with FL and was not truely pear shaped. Strains ‘G1’ and ‘LS’ had more FW. Round strains had higher TSS and TS compared with Pyriform strains. Wide variation in fruit size, weight and sugars have been reported in guava in different genotypes and seasons (Sharma et al. 2010; Adrees et al., 2010; Usman et al., 2012, 2018) and it could also be a metaxenial effect (Usman et al., 2013; Aslam et al., 2019). Genetic diversity assessment using molecular markers could eliminate the agro-climatic impact leading to phenotypic variation. Different DNA markers including RAPD have been used in guava for diversity assessment and development of phylogenetic relationships. RAPD markers have been very useful in finding polymorphism in guava (P. guajava L.) genotypes in India (Prakash et al., 2002), Bangladesh (Ahmed et al., 2011), Mexico (Valera-Montero et al., 2016) and in Feijoa (Acca sallowiana) strains (Dettori and Palombi, 2000). However, no previous study has reported RAPD based clustering of guava strains having different fruit shapes (Round and Pyriform). Overall, diversity observed in physico-chemical traits of Round and Pyriform varieties was low and correlated to the less molecular variance among varieties compared with variance within varieties (among strains). Low varietal variance could be attributed to the prevalence of much higher frequency of self-pollination than cross- pollination. Low molecular variance was also reported in populations of guava compared with individuals (Khrewar et al., 2018). The RAPD markers classified eight strains of two varieties into two separate clusters which were distinct in fruit shape. The distinctive RAPD markers could be effectively utilized for the development of SCAR markers which could help in identification and genetic characterization of intervarietial hybrids.
Conclusions: Guava strains Gola No. 1 and Large Surahi were found better in fruit size, weight, TSS and total sugars, however, Sadabahar Surahi is preferred due to its profusely regular bearing habit. Physico-chemical and molecular markers highlighted existence of higher variance among strains of both cultivars. The RAPD markers are cost-effective and have separately clustered guava strains having Round and Pyriform shapes. The potential of RAPD and other molecular markers shall be extensively utilized for genetic characterization in heterozygous guava populations for molecular breeding applications.
Acknowledgements: The authors are grateful to USDA Endowment Fund Secretariat, University of Agriculture, Faisalabad-Pakistan and ALP-PARC, Islamabad for partial funding of this research under Guava In Vitro Clonal Propagation and Guava Polyploidization (CS-121) Projects.
Novelty statement: The manuscript highlights: elite guava strains found better for fruit size and quality including Gola No. 1 and Large Surahi, existence of higher molecular variance among strains than varieties, RAPD based distinct clustering of elite strains of two varieties classified on the basis of fruit shape as Round and Pyriform.
REFERENCES
- Adrees, M., M. Younis, U. Farooq, and K. Hussain (2010). Nutritional quality evaluation of different guava varieties. Pak. J. Agri. Sci. 47: 1-4.
- Ahmed, B., M.A. Mannan, and S.A. Hossain (2011). Molecular characterization of guava (Psidium guajava L.) germplasm by RAPD analysis. Int. J. Nat. Sci. 1: 62-67.
- Annoles, G.C., B.J. Bassam, and P.M. Gresshoff (1991). DNA amplification and Fingerprinting using short arbitrary oligonucleotide primers. Biotechonol. 9: 553-557.
- Aslam, M.M., M. Usman, B. Fatima, M. Shahid, S.A.M. Bokhari, M.A. Rana (2019). Compatibility and Metaxenial Effects on Fruit Physico-Chemical Attributes in Intervaretial Crosses in Guava (Psidium guajava L.). 1st Int. Conf. on Hort. Crop Production & Protection, Univ. of Sargodha, Pakistan. Pp. 2.
- Biswas, M.K., Q. Xu, and X.X. Deng (2010). Utility of RAPD, ISSR, IRAP and REMAP markers for the genetic analysis of Citrus spp. Sci. Hort. 124: 254-261.
- Chandra, R.A., S.B. Gupta, and R.K. Tiwari (2004). Embryogenesis and plant regeneration from mesocarp of Psidium guajava L. Ind. J. Biotech. 3: 246-248.
- Damon, R.A. and W.R. Harvey (1987). Experimental Design, ANOVA and Regression. Harper and Row Publishers; New York.
- Dettori, M.T. and M.A. Palombi (2000). Identification of Feijoa sellowiana Berg. accessions by RAPD markers. Sci. Hort.86: 279-290.
- Doyle, J.J. and J.L. Doyle (1990). Isolation of DNA from small amounts of plant tissues. BRL. Focus. 12: 13-15.
- Anonymous (2016). Federal Board of Statistics, Govt. Pakistan., Islamabad.
- Fukuda, S., K. Ishimoto, S. Sato, S. Terakami, N. Hiehata, and T. Yamamoto (2016). A high-density genetic linkage map of bronze loquat based on SSR and RAPD markers. Tree Genet. Genomes. 12(4): 80.
- Hortwitz, W. (1960). Official and tentative methods of analysis. Washington, D.C.Assoc. Office Agric. Chem. 9: 314-320.
- Khan, I. A., F. S. Awan, A. Ahmad and A. A. Khan (2004). A modified mini-prep method for economical and rapid extraction of genomic DNA in plants. Plant Mol. Biol. Rep. 22(1): 89-89.
- Kherwar, D., K. Usha, S.A. Mithra and B. Singh (2018). Microsatellite (SSR) marker assisted assessment of population structure and genetic diversity for morpho-physiological traits in guava (Psidium guajava L.). Plant Biochem. and Biotech. 27(3): 284-292.
- Martins, M., D. Sarmento, and M.M. Oliveira (2004). Genetic stability of micro propagated almond plantlets, as assessed by RAPD and ISSR markers. Plant Cell Rep. 23: 492-496.
- Munankarmi, N.N., R.L Shrestha, N. Rana, J.K.C. Shrestha, S. Shrestha, and R. Koirala (2014). Genetic Diversity Assessment of Acid Lime (Citrus aurantifolia Swingle) Landraces of Eastern Nepal Using RAPD Markers. Int. J. Appl. Sci. Biotech. 2: 315-327.
- Nei, M. (1987). Unweighted Paired Group of Arithmetic Means Averages (UPGMA) to estimate genetic distances and relatedness. Molecular Evolutionary Genetics. Columbia Univ. Press; New York.
- Prakash, D.P., P. Narayanaswamy, and S.N. Sondur (2002). Analysis of molecular diversity in guava using RAPD markers. J. Hortic. Sci. Biotechol. 77: 287-293.
- Pereira, F.M.P., M. Usman, N.A. Mayer, J.C. Nachtigal, O.R. Maphanga, and S. Willemse (2017). Advances in guava propagation. Rev. Bras. Frutic. 39(4): e-358.
- Peakall, R. and P.E. Smouse (2006). GenAlEx 6: Genetic analysis in excel. Population genetic software for teaching and research. Mol. Ecol. Notes 6: 288-295.
- Rai, M.K., P. Asthana, V.S. Jaiswal, and U. Jaiswal (2010). Biotechnological advances in guava (Psidium guajava L.): recent developments and prospects for further research. Struct. Funct. Genomics. 24: 1-12.
- Ray, T., I. Dutta, P. Saha, S. Das, and S.C. Roy (2006). Genetic stability of three economically important micropropagated banana (Musa spp.) cultivars of lower Indo-Gangetic plains, as assessed by RAPD and ISSR markers. Plant Cell Tissue Organ Cult. 85: 11-21.
- Ruck, J.A. (1969). Chemical methods for analysis of fruits and vegetables, Summerland Res. Stat, Dept. Agri., Canada.
- Russell, D.F. and S.P. Eisensmith (1983). MSTAT-C. Crop Soil Sci. Deptt., Michigan State Univ., USA.
- Sharma, A., S.K. Sehrawat, R.S. Singhrot, and A. Tele (2010). Morphological and Chemical characterization of Psidium Species. Not. Bot. Hort. Agrbot. Cluj. 38: 28-32.
- UPOV. (1987). Guava descriptors. International Union for the Protection of new Varieties of plants. Geneva, Sweden.
- Usman, M., M. Butt, and B. Fatima (2012). Enhanced in vitro multiple shoot induction in elite Pakistani guava cultivars for efficient clonal plant multiplication. Afr. J. Biotechol. 11:10182-10187.
- Usman, M., W.A. Samad, B. Fatima, and M.H. Shah (2013). Pollen parent enhances fruit size and quality in intervarietal crosses in guava (Psidium guajava L.). Int. J. Agric. Biol. 15: 125-129.
- Usman, M. R. Arif, B. Fatima and M. Awais (2018). Screening of ‘Round’ Guava strains for Physico-chemical Traits. 2nd Int. Conf. on Plant Sciences, GCU, Lahore, Pakistan. Pp. 102.
- Valera-Montero, L.L., P.J. Muñoz-Rodríguez, H. Silos-Espino, and S. Flores-Benítez (2016). Genetic diversity of guava (Psidium guajava L.) from Central Mexico revealed by morphological and RAPD markers. Phyton-Int. J. Exp. Bot. 85: 176-183.
- Williams, J.K.W., A.R. Kubelik, K.J. Livak, J.A. Rafalski, and S.V. Tingey (1990). DNA Polymorphisms Amplified by Arbitrary Primers Are Useful as Genetic Markers. Nucleic Acids Res. 18(22): 6531-6535.
- Yeh, F.C., R. Yang, T.J. Boyle, Z. Ye, and J.M. Xiyan (2000). Popgene32, Microsoft Windows based freeware for population genetic analysis. Mol. Biol. Biotechnol. Centre, Univ. Alberta, Edmonton, Canada.
|





