GENETIC DIVERSITY AND DNA FINGERPRINTING OF POTATO VARIETIES USING SIMPLE SEQUENCE REPEAT (SSR) MARKERS
S. U. Rahman, S. Jamil*, R. Shahzad, E. Yasmeen, S. Sattar and M. Z. Iqbal
Agricultural Biotechnology Research Institute, AARI Faisalabad 38040, Pakistan
*Correspondence: Shakrajamil29@gmail.com
ABSTRACT
DNA fingerprinting is a tool for plant breeder rights protection, and variety registration in Plant Breeder Rights Repository. In the present study, we developed a DNA fingerprinting profile of 12 potato cultivars grown in Punjab Pakistan using 214 informative Simple Sequence Repeat (SSR) markers. A total of 1720 alleles were amplified by 214 SSR with an average of 8.04 alleles per marker. Approximately 72% of amplified alleles (1329 alleles) were polymorphic with 6.88 polymorphic alleles per SSR marker. The number of alleles ranged from 1 to 31. Similarly, polymorphic alleles per marker ranged from 0 to 24. A maximum number of alleles and polymorphic alleles were reported by IBR13 marker. The Polymorphic information content (PIC) value ranged from 0 to 0.96. The average PIC value for 214 amplified markers was 0.73. Collectively, 72 SSR markers amplified unique allelic patterns for DNA fingerprinting. Potato varieties Rubby and Sadaf were identified by 15 SSR markers whereas Faisalabad Red and SH-5 were identified by 12 SSR markers. Cluster and structure analysis classify the potato genotypes into two distinct groups. This information will be useful for the variety registration process and will provide a platform for future DNA fingerprinting and genetic diversity studies for the choice of SSR markers.
Keywords: Cluster analysis; Genetic diversity; Polymerase Chain Reaction; Polymorphic Information Content; Variety identification,
http://doi.org/10.36899/JAPS.2022.3.0479
Published first online October 19. 2021
INTRODUCTION
Potato (Solanum tuberosum L.) is the fourth most important and non-cereal principal cash crop globally (Devaux et al. 2014). It is high yielding and high nutritive vegetable, possessing carbohydrates and several minerals, fibers, fats, and vitamins contributing a total of 390 KJ 100g ⁻¹of baked potatoes (Majeed and Muhammad, 2018). Pakistan is among the large potato producing countries yielding 45.66 tonnes per hectares. According to Economic Survey of Pakistan, in 2020, Pakistan produced a total of 4,609, 600 tones with a total yield of 245,159 kg/ha of potatoes with 95% of production mainly originated from Punjab. Pakistan has been largely exporting potatoes to Afghanistan, Middle East, Russia, and Europe. In 2018, the country exported potatoes worth 120.9 million USD (GOP, 2020).
With ever increasing population and growing food demands (Shahzad et al., 2021a, b) potato varieties are evolving at faster rates with improved taste, enhanced pathogenic resistance, and high yield. Ever-increasing crop varieties need to be distinguished from each other. Traditionally, potato varieties were identified using morphological and physiological traits on the basis of Distinctiveness, Uniformity and Stability (DUS) testing through color, texture, sprouting, growth habit, and disease resistance. These traits are high affected by environment and lead towards false identification hence generating high risk of germplasm mixing (van Eck, 2007). Furthermore, biochemical markers such as isozymes are affected by developmental stages and growth conditions of a plant. Hence, rapid, reliable, and accurate identification of varieties is necessary for development of new cultivars with minimal laboratory instruments (Singh et al., 2019; Jamil et al., 2021a).
Molecular marker or DNA based markers are widely accepted for identification due to their immense benefits over morphological and biochemical markers (Vreugdenhil et al., 2011). Genetic markers are most efficient tool for identification of cultivar and estimation of relatedness. Further, Plant breeders’ Rights Rules 2018, issued by Ministry of National Food Security and Research, Pakistan clearly states the necessity of DNA profiling before registration of new variety for varietal protection (Sadaf, 2018; Iqbal et al., 2021a). Various PCR based markers including randomly amplified polymorphic DNA (RAPD), Amplified fragment length polymorphism (AFLP), Inter Simple Sequence Repeat (ISSR), Restriction Fragment Length Polymorphism (RFLP), and Single Nucleotide Polymorphism (SNPs) are the promising tools to assess genetic diversity and fingerprinting (Iqbal et al., 2019; Jamil et al., 2020 a, b).
Microsatellites or SSR (Simple sequence repeats) markers are comprised of short 1-6 bp repetitive DNA motifs that act as a powerful tool for studying genetic diversity, phylogeny, cultivar discrimination, marker-assisted selection for breeding and genome mapping (Lan et al., 2012; Vieira et al., 2016). They are simple, highly informative, reproducible, abundant, co-dominant, frequently, and randomly spread all over genome and have a specific location on a chromosome that can be easily analysed using PCR (Kalia et al., 2011). SSR was first used for genetic analysis of anther-derived potatoes and later on several studies demonstrated use of SSR markers to differentiate potato cultivars e.g. 38 accessions of potatoes in Brazil were distinguished using SSR markers (Veilleux et al., 1995; Favoretto et al. 2011). 589 native Chilean potato accessions were analysed for genetic diversity using SSR markers (Muñoz et al., 2016).
Although various studies relating to potato DNA fingerprinting are reported worldwide but no such report is available in Pakistan to date. In the present study, 214 SSR markers were used to analyse, distinguish, and generate fingerprints of twelve potato varieties in Punjab Pakistan. The SSR markers were selected based on polymorphism and their ability to discriminate. Present study will provide vivid information for breeding programs, genotype verification, cultivar registration, genetic assessment, and variety protection.
MATERIALS AND METHODS
Twelve potato genotypes (Table 1) were surveyed using SSR markers for varietal discrimination and genetic diversity assessment. The cultivars were grown in growth chambers at Agriculture Biotechnology Research Institute (ABRI), Faisalabad, Pakistan under standard growth conditions. The fresh young leaflets were collected and stored at -40℃ till proceeded further.
Table 1. List of Potato genotypes used in the study along with pedigree parentage
|
Sr. No.
|
Name
|
Pedigree Parentage
|
|
1
|
Faisalabad Red
|
Desiree × Laal-e-Faisal
|
|
2
|
Faisalabad White
|
CIP Clone No. 386043 (Introduction)
|
|
3
|
SH-5
|
Bartina × Cardinal
|
|
4
|
PRI-Red
|
FD44-24 × FD 12-24
|
|
5
|
Ruby
|
384636-1 × FD 1-8
|
|
6
|
Sadaf
|
FD 3-15 × FD 35-36
|
|
7
|
Sialkot Red
|
SH-5 × Cardinal
|
|
8
|
Sahiwal White
|
FD 35-36 × SH-5
|
|
9
|
Cosmo
|
FD 3-15 × FD 35-36
|
|
10
|
Sahiwal Red
|
FD 3-15 × SH-5
|
|
11
|
Ravi Red
|
FD 35-36 × SH-5
|
|
12
|
FD 81-1
|
N-9619 × FD 3-15
|
Genomic DNA was extracted from 2 grams of young leaflets by grinding samples in liquid nitrogen. Finely powdered samples were preceded according to modified Cetyl Trimethyl Ammonium Bromide (CTAB) method (Kanwal et al., 2021). The samples were quantified using Nanodrop spectrophotometer (ND200, Thermo Scientific, U.S.A.). The extracted DNA with A260/A280 = 1.80 – 2.0 was considered as pure. Furthermore, the quality and quantity of each extract was assessed by loading DNA 20 ng/ µL on 0.8% (w/v) on agarose gel stained with ethidium bromide. DNA samples were stored at -40℃. 226 polymorphic SSR markers (Table S1) were selected and synthesized according to previous studies (Buteler et al., 1999; Hwang et al., 2002; He et al., 2003; Ghislain et al., 2009; Salimi et al., 2016). Polymorphic SSR markers were used to fingerprint and analyse genetic diversity of 12 potato cultivars.
PCR amplifications were performed in a 25 μL reaction mixture, containing 2.0 μL of 20 ng/μL DNA template, 0.6 μM forward primer, 0.6 μM of reverse primer, 12 μL green master mix and volume was adjusted using double distilled deionized water (d3H2O) for each SSR marker. Following PCR program was used: initial denaturation 5 min at 94 °C, 35 cycles of denaturation of 30 s at 94 °C, 1 min of annealing at SSR specific annealing temperature (46-60 °C) and 45 s at 72 °C; with a final extension step of 7 min at 72 °C. The PCR products were stored at 4°C.
All amplified products were fractioned on vertical gel electrophoresis System model POWERPRO-3AMP (cleaver scientific limited) using 6% PAGE performed at 16 watts power using 50 bp DNA ladder as a reference. PAGE gels were further stained by Silver nitrate staining for visualization according to previously described staining protocol (Jamil et al., 2020b). Finally, images were captured using Syngene trans-illuminator.
The data file for SSR markers was constructed in the form of a binary matrix by scoring 0 for absence and 1 for presence of specific amplification of allele. For cluster analysis, a distance matrix was generated using NTSYSpc 2.0 and Un-weighted pair Group Method of Arithmetic Means (UPGMA) was used to construct dendrogram. Moreover, population structure and genetic diversity levels of potato genotypes were estimated using a model-based Bayesian clustering approach implemented in STRUCTURE v. 2.3.4 (Pritchard et al., 2000) by assigning accessions to population based on SSR markers. Genotyping data of 214 microsatellites were used to determine population’s structure of various potato varieties. Population structure analysis was performed according to parameters previously mentioned in our recent paper (Jamil et al., 2020b). Further Polymorphic Information contents (PIC) were also calculated for all SSR markers along with number of alleles, polymorphic alleles, and different allelic diversity parameters.
RESULTS
SSR Polymorphism: Among 226 SSR markers used in the study, twelve markers i.e. CB330645, CB330657, IB2·45A, Ib-255F1, IB2-66, Ib3/31, Ib-316, IBC1, IBSSR16, IBSSR23, IBSSR24, and IBSSR25 were not amplified remaining 214 markers were used for fingerprinting and genetic diversity studies. Among 214 amplified markers 21 were monomorphic and 193 were polymorphic. A total of 1720 alleles were amplified with an average of 8.04 alleles per marker. Approximately 72% of amplified alleles (1329 alleles) were polymorphic with 6.88 polymorphic alleles per SSR marker. The number of alleles ranged from 1 (for 23 SSR markers) to 31 (IBR13). Similarly, polymorphic alleles per markers ranged from 0 (15 markers) to 24 (IBR13). The average number of alleles and polymorphic alleles was 8.2 and 6.2 respectively. The PIC value for markers ranged from 0 (12 SSR markers) to 0.96 (IBR13). The average PIC value for 214 amplified markers was 0.73. Whereas size of amplified products ranged from 80 to 1000 bp (Table 2).
DNA Fingerprinting: The 193 polymorphic markers were used for fingerprinting of 12 potato genotypes. Twelve potato genotypes were identified successfully by using different seventy-two markers. SSR markers BU691268 (Fig 1) and IBJ5446 each distinguished three potato genotypes (SH-5, Ruby & Sadaf) and (Faisalabad-Red, Ruby, and Cosmo) respectively. Similarly, 14 markers i.e., STM0019a, STM1052, STM5114, STM5127, Ib-286, IBN-35, IBE-5, CB330283, CB330200, BM878740, BU692896, CB330762, IBE32 and IBJ62 identified two genotypes each as given in Table 3. The genotypes Ruby and Sadaf were uniquely identified by 15 markers whereas Faisalabad Red and SH-5 were identified with 12 markers each. Similarly, Sahiwal White was identified with the help of 11 SSR markers and Faisalabad White by 10 markers. The remaining genotypes i.e., Ravi Red, Sahiwal Red, PRI Red, Sialkot Red, Cosmo, and FD 81-1 were uniquely identified using 6, 4, 3, 2, 2 and 1 SSR markers respectively (Table 3)
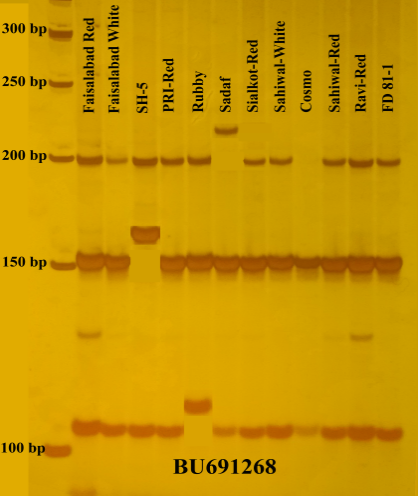
Fig 1. Polyacrylamide gel electrophoresis result of BU691268 SSR marker containing DNA fingerprints of three genotypes i.e., SH-5, Rubby, and Sadaf.
Table 2. List of SSR markers used in the study along with polymorphism status (PS), Polymorphic Information contents (PIC), No. of alleles (NOA), Polymorphic Alleles (PA), and annealing temperature (AT).
|
Sr. No.
|
Marker Name
|
PS
|
PIC
|
NOA
|
PA
|
TA
|
Sr. No.
|
Marker Name
|
PS
|
PIC
|
NOA
|
PA
|
TA
|
Sr. No.
|
Marker Name
|
PS
|
PIC
|
NOA
|
PA
|
TA
|
|
1
|
BM878740
|
Polymorphic
|
0.91
|
14
|
9
|
45
|
73
|
IB-255
|
Polymorphic
|
0.79
|
5
|
5
|
45
|
144
|
IBR12
|
Polymorphic
|
0.94
|
19
|
14
|
46
|
|
2
|
BM878757
|
Polymorphic
|
0.89
|
11
|
9
|
42
|
74
|
Ib-275
|
Polymorphic
|
0.72
|
4
|
1
|
46
|
145
|
IBR13
|
Polymorphic
|
0.96
|
31
|
24
|
44
|
|
3
|
BM878879
|
Polymorphic
|
0.65
|
3
|
1
|
45
|
75
|
Ib-286
|
Polymorphic
|
0.89
|
13
|
4
|
42
|
146
|
IBR14
|
Polymorphic
|
0.74
|
5
|
2
|
46
|
|
4
|
BU690134
|
Polymorphic
|
0.96
|
27
|
22
|
42
|
76
|
Ib-297
|
Polymorphic
|
0.83
|
7
|
6
|
46
|
147
|
IBR16
|
Polymorphic
|
0.90
|
12
|
10
|
44
|
|
5
|
BU690375
|
Polymorphic
|
0.79
|
6
|
5
|
42
|
77
|
Ib3/24
|
Polymorphic
|
0.79
|
5
|
1
|
44
|
148
|
IBR19
|
Polymorphic
|
0.75
|
7
|
5
|
44
|
|
6
|
BU690615
|
Polymorphic
|
0.72
|
4
|
4
|
42
|
78
|
Ib3/28
|
Monomorphic
|
0.00
|
1
|
0
|
42
|
149
|
IBR20
|
Polymorphic
|
0.90
|
13
|
10
|
44
|
|
7
|
BU690708
|
Polymorphic
|
0.63
|
3
|
3
|
44
|
79
|
Ib-318
|
Polymorphic
|
0.64
|
4
|
2
|
44
|
150
|
IBR21
|
Polymorphic
|
0.82
|
6
|
6
|
46
|
|
8
|
BU690750
|
Polymorphic
|
0.93
|
17
|
16
|
44
|
80
|
IbC10
|
Polymorphic
|
0.77
|
5
|
4
|
48
|
151
|
IBS01
|
Monomorphic
|
0.00
|
1
|
0
|
44
|
|
9
|
BU690910
|
Polymorphic
|
0.68
|
4
|
3
|
44
|
81
|
IbC11
|
Polymorphic
|
0.80
|
7
|
3
|
42
|
152
|
IBS02
|
Polymorphic
|
0.68
|
4
|
3
|
44
|
|
10
|
BU690977
|
Monomorphic
|
0.00
|
1
|
0
|
46
|
82
|
IbC12
|
Polymorphic
|
0.79
|
5
|
2
|
44
|
153
|
IBS07
|
Polymorphic
|
0.62
|
3
|
2
|
46
|
|
11
|
BU691143
|
Polymorphic
|
0.91
|
14
|
12
|
42
|
83
|
IBC13
|
Polymorphic
|
0.73
|
4
|
1
|
44
|
154
|
IBS09
|
Polymorphic
|
0.78
|
8
|
7
|
46
|
|
12
|
BU691268
|
Polymorphic
|
0.89
|
7
|
7
|
42
|
84
|
IBC3
|
Polymorphic
|
0.47
|
4
|
3
|
44
|
155
|
IBS10
|
Polymorphic
|
0.77
|
5
|
3
|
46
|
|
13
|
BU691341
|
Polymorphic
|
0.92
|
19
|
15
|
42
|
85
|
IBC4
|
Polymorphic
|
0.52
|
4
|
3
|
44
|
156
|
IBS11
|
Polymorphic
|
0.63
|
3
|
3
|
50
|
|
14
|
BU691547
|
Polymorphic
|
0.93
|
19
|
15
|
42
|
86
|
IBC5
|
Polymorphic
|
0.47
|
3
|
3
|
46
|
157
|
IBS17
|
Polymorphic
|
0.88
|
12
|
11
|
44
|
|
15
|
BU691662
|
Monomorphic
|
0.00
|
1
|
1
|
42
|
87
|
IBC8
|
Polymorphic
|
0.74
|
5
|
3
|
44
|
158
|
IBS18
|
Polymorphic
|
0.70
|
5
|
3
|
44
|
|
16
|
BU691762
|
Monomorphic
|
0.00
|
1
|
0
|
42
|
88
|
IBC9
|
Polymorphic
|
0.77
|
5
|
3
|
46
|
159
|
IBSR01
|
Polymorphic
|
0.44
|
2
|
2
|
42
|
|
17
|
BU691865
|
Polymorphic
|
0.85
|
8
|
4
|
45
|
89
|
IBCIP-1
|
Polymorphic
|
0.90
|
12
|
6
|
42
|
160
|
IBSSR02
|
Polymorphic
|
0.32
|
2
|
1
|
46
|
|
18
|
BU691949
|
Polymorphic
|
0.85
|
11
|
9
|
42
|
90
|
IBCIP-12
|
Monomorphic
|
0.80
|
5
|
0
|
46
|
161
|
IBSSR03
|
Monomorphic
|
0.00
|
1
|
0
|
45
|
|
19
|
BU691984
|
Monomorphic
|
0.00
|
1
|
1
|
42
|
91
|
IBCIP-13
|
Polymorphic
|
0.85
|
8
|
6
|
42
|
162
|
IBSSR04
|
Polymorphic
|
0.92
|
12
|
12
|
42
|
|
20
|
BU692061
|
Monomorphic
|
0.00
|
1
|
1
|
46
|
92
|
IBCIP-2
|
Polymorphic
|
0.92
|
16
|
6
|
42
|
163
|
IBSSR05
|
Polymorphic
|
0.82
|
6
|
6
|
45
|
|
21
|
BU692090
|
Polymorphic
|
0.85
|
8
|
5
|
45
|
93
|
IBCIP-5
|
Polymorphic
|
0.87
|
9
|
8
|
42
|
164
|
IBSSR06
|
Polymorphic
|
0.79
|
5
|
5
|
44
|
|
22
|
BU692095
|
Polymorphic
|
0.93
|
18
|
12
|
45
|
94
|
IBCIP-7
|
Polymorphic
|
0.87
|
10
|
7
|
42
|
165
|
IBSSR07
|
Polymorphic
|
0.15
|
2
|
2
|
48
|
|
23
|
BU692154
|
Polymorphic
|
0.63
|
5
|
5
|
42
|
95
|
IBCIP-8
|
Monomorphic
|
0.00
|
1
|
0
|
46
|
166
|
IBSSR08
|
Polymorphic
|
0.84
|
8
|
6
|
42
|
|
24
|
BU692227
|
Polymorphic
|
0.85
|
10
|
9
|
46
|
96
|
IBCIP-9
|
Polymorphic
|
0.87
|
8
|
3
|
46
|
167
|
IBSSR09
|
Polymorphic
|
0.76
|
5
|
4
|
46
|
|
25
|
BU692248
|
Polymorphic
|
0.38
|
2
|
2
|
46
|
97
|
IBE1
|
Polymorphic
|
0.84
|
8
|
8
|
44
|
168
|
IBSSR10
|
Polymorphic
|
0.90
|
11
|
11
|
44
|
|
26
|
BU692403
|
Polymorphic
|
0.90
|
11
|
11
|
42
|
98
|
IBE10
|
Polymorphic
|
0.85
|
9
|
5
|
44
|
169
|
IBSSR11
|
Polymorphic
|
0.86
|
9
|
9
|
44
|
|
27
|
BU692471
|
Polymorphic
|
0.67
|
4
|
3
|
46
|
99
|
IBE12
|
Polymorphic
|
0.90
|
13
|
13
|
42
|
170
|
IBSSR12
|
Polymorphic
|
0.83
|
9
|
9
|
46
|
|
28
|
BU692496
|
Monomorphic
|
0.63
|
3
|
3
|
46
|
100
|
IBE14
|
Polymorphic
|
0.95
|
25
|
20
|
42
|
171
|
IBSSR13
|
Polymorphic
|
0.41
|
3
|
2
|
44
|
|
29
|
BU692566
|
Polymorphic
|
0.70
|
4
|
1
|
46
|
101
|
IBE15
|
Polymorphic
|
0.86
|
10
|
8
|
42
|
172
|
IBSSR14
|
Polymorphic
|
0.44
|
2
|
1
|
50
|
|
30
|
BU692646
|
Monomorphic
|
0.00
|
1
|
0
|
46
|
102
|
IBE2
|
Polymorphic
|
0.63
|
3
|
3
|
48
|
173
|
IBSSR15
|
Polymorphic
|
0.66
|
3
|
2
|
44
|
|
31
|
BU692658
|
Polymorphic
|
0.91
|
13
|
10
|
42
|
103
|
IBE24
|
Polymorphic
|
0.89
|
11
|
9
|
42
|
174
|
IBSSR17
|
Polymorphic
|
0.57
|
3
|
2
|
46
|
|
32
|
BU692739
|
Polymorphic
|
0.89
|
13
|
11
|
42
|
104
|
IBE27
|
Polymorphic
|
0.91
|
15
|
14
|
42
|
175
|
IBSSR18
|
Polymorphic
|
0.83
|
8
|
7
|
44
|
|
33
|
BU692763
|
Polymorphic
|
0.79
|
5
|
4
|
48
|
105
|
IBE28
|
Polymorphic
|
0.90
|
14
|
14
|
42
|
176
|
IBSSR19
|
Polymorphic
|
0.68
|
5
|
3
|
45
|
|
34
|
BU692780
|
Polymorphic
|
0.81
|
6
|
2
|
42
|
106
|
IBE29
|
Polymorphic
|
0.72
|
4
|
4
|
42
|
177
|
IBSSR20
|
Monomorphic
|
0.00
|
1
|
1
|
46
|
|
35
|
BU692800
|
Monomorphic
|
0.00
|
1
|
0
|
46
|
107
|
IBE3
|
Polymorphic
|
0.64
|
4
|
4
|
44
|
178
|
IBSSR21
|
Polymorphic
|
0.81
|
7
|
7
|
46
|
|
36
|
BU692858
|
Polymorphic
|
0.85
|
9
|
7
|
42
|
108
|
IBE30
|
Polymorphic
|
0.81
|
6
|
4
|
42
|
179
|
IBSSR22
|
Polymorphic
|
0.83
|
6
|
2
|
46
|
|
37
|
BU692896
|
Polymorphic
|
0.92
|
17
|
9
|
42
|
109
|
IBE32
|
Polymorphic
|
0.71
|
6
|
5
|
42
|
180
|
IBSSR26
|
Polymorphic
|
0.90
|
11
|
5
|
48
|
|
38
|
BU692914
|
Polymorphic
|
0.91
|
14
|
13
|
42
|
110
|
IBE33
|
Polymorphic
|
0.89
|
10
|
7
|
42
|
181
|
IBSSR27
|
Polymorphic
|
0.86
|
9
|
9
|
44
|
|
39
|
BU692940
|
Polymorphic
|
0.88
|
10
|
10
|
42
|
111
|
IBE34
|
Polymorphic
|
0.87
|
9
|
7
|
42
|
182
|
IBSSR331
|
Polymorphic
|
0.84
|
7
|
3
|
42
|
|
40
|
CB329940
|
Monomorphic
|
0.00
|
1
|
0
|
45
|
112
|
IBE4
|
Polymorphic
|
0.85
|
9
|
6
|
46
|
183
|
SSR128
|
Polymorphic
|
0.88
|
11
|
7
|
50
|
|
41
|
CB329965
|
Monomorphic
|
0.92
|
12
|
0
|
42
|
113
|
IBE5
|
Polymorphic
|
0.90
|
14
|
3
|
46
|
184
|
SSR270
|
Polymorphic
|
0.89
|
12
|
12
|
45
|
|
42
|
CB330083
|
Monomorphic
|
0.89
|
9
|
0
|
48
|
114
|
IBE7
|
Polymorphic
|
0.86
|
12
|
12
|
44
|
185
|
SSR38
|
Polymorphic
|
0.88
|
10
|
7
|
45
|
|
43
|
CB330141
|
Polymorphic
|
0.74
|
6
|
4
|
42
|
115
|
IBE8
|
Polymorphic
|
0.82
|
8
|
3
|
46
|
186
|
SSR4
|
Polymorphic
|
0.78
|
6
|
5
|
50
|
|
44
|
CB330144
|
Monomorphic
|
0.00
|
1
|
0
|
48
|
116
|
IBJ1525
|
Polymorphic
|
0.92
|
16
|
8
|
44
|
187
|
SSR450
|
Monomorphic
|
0.80
|
5
|
5
|
45
|
|
45
|
CB330200
|
Polymorphic
|
0.87
|
11
|
9
|
46
|
117
|
IBJ1798E
|
Polymorphic
|
0.90
|
13
|
9
|
45
|
188
|
SSR555
|
Polymorphic
|
0.90
|
11
|
3
|
45
|
|
46
|
CB330223
|
Polymorphic
|
0.63
|
3
|
1
|
48
|
118
|
IBJ199
|
Polymorphic
|
0.93
|
18
|
18
|
46
|
189
|
SSR578
|
Polymorphic
|
0.70
|
5
|
3
|
50
|
|
47
|
CB330283
|
Polymorphic
|
0.41
|
4
|
3
|
46
|
119
|
IbJ206a
|
Polymorphic
|
0.60
|
5
|
4
|
42
|
190
|
SSR92
|
Polymorphic
|
0.64
|
3
|
1
|
45
|
|
48
|
CB330285
|
Polymorphic
|
0.84
|
8
|
8
|
48
|
120
|
IBJ206b
|
Polymorphic
|
0.79
|
5
|
1
|
44
|
191
|
STG0001
|
Polymorphic
|
0.90
|
12
|
12
|
52
|
|
49
|
CB330296
|
Polymorphic
|
0.84
|
7
|
3
|
42
|
121
|
IBJ27
|
Polymorphic
|
0.72
|
4
|
4
|
42
|
192
|
STG0010
|
Polymorphic
|
0.59
|
3
|
3
|
55
|
|
50
|
CB330416
|
Polymorphic
|
0.86
|
10
|
8
|
42
|
122
|
IBJ290
|
Polymorphic
|
0.81
|
6
|
4
|
46
|
193
|
STG0016
|
Polymorphic
|
0.87
|
9
|
9
|
53
|
|
51
|
CB330456
|
Polymorphic
|
0.38
|
2
|
1
|
48
|
123
|
IBJ302
|
Polymorphic
|
0.91
|
17
|
15
|
42
|
194
|
STG0025
|
Polymorphic
|
0.76
|
6
|
5
|
55
|
|
52
|
CB330471
|
Polymorphic
|
0.92
|
16
|
16
|
48
|
124
|
IBJ3816E
|
Polymorphic
|
0.71
|
4
|
2
|
44
|
195
|
STI0001
|
Polymorphic
|
0.86
|
9
|
9
|
55
|
|
53
|
CB330554
|
Polymorphic
|
0.87
|
12
|
10
|
42
|
125
|
IBJ462E
|
Polymorphic
|
0.85
|
7
|
7
|
55
|
196
|
STI0003
|
Polymorphic
|
0.90
|
11
|
9
|
55
|
|
54
|
CB330601
|
Polymorphic
|
0.91
|
16
|
14
|
42
|
126
|
IBJ530
|
Polymorphic
|
0.89
|
11
|
7
|
48
|
197
|
STI0004
|
Polymorphic
|
0.84
|
8
|
7
|
55
|
|
55
|
CB330627
|
Polymorphic
|
0.90
|
14
|
14
|
42
|
127
|
IBJ5446b
|
Polymorphic
|
0.87
|
13
|
10
|
44
|
198
|
STI0012
|
Polymorphic
|
0.87
|
10
|
9
|
55
|
|
56
|
CB330636
|
Polymorphic
|
0.86
|
13
|
9
|
42
|
128
|
IBJ559
|
Polymorphic
|
0.90
|
15
|
12
|
44
|
199
|
STI0014
|
Polymorphic
|
0.76
|
5
|
5
|
55
|
|
57
|
CB330643
|
Polymorphic
|
0.74
|
5
|
3
|
42
|
129
|
IBJ566
|
Polymorphic
|
0.89
|
11
|
9
|
44
|
200
|
STI0030
|
Polymorphic
|
0.59
|
3
|
3
|
58
|
|
58
|
CB330649
|
Monomorphic
|
0.00
|
1
|
0
|
46
|
130
|
IBJ62
|
Polymorphic
|
0.93
|
23
|
21
|
42
|
201
|
STI0032
|
Polymorphic
|
0.88
|
10
|
9
|
60
|
|
59
|
CB330675
|
Monomorphic
|
0.67
|
3
|
0
|
46
|
131
|
IBJ766E
|
Polymorphic
|
0.93
|
17
|
11
|
44
|
202
|
STI0033
|
Polymorphic
|
0.86
|
10
|
10
|
60
|
|
60
|
CB330693
|
Polymorphic
|
0.83
|
10
|
8
|
43
|
132
|
IBJ90
|
Polymorphic
|
0.86
|
8
|
6
|
46
|
203
|
STM0019a
|
Polymorphic
|
0.83
|
9
|
9
|
54
|
|
61
|
CB330694
|
Polymorphic
|
0.89
|
15
|
14
|
44
|
133
|
IBN18
|
Monomorphic
|
0.67
|
3
|
0
|
42
|
204
|
STM0031
|
Polymorphic
|
0.89
|
14
|
14
|
48
|
|
62
|
CB330729
|
Polymorphic
|
0.82
|
7
|
7
|
42
|
134
|
IBN21
|
Polymorphic
|
0.91
|
14
|
13
|
42
|
205
|
STM0037
|
Polymorphic
|
0.75
|
5
|
5
|
56
|
|
63
|
CB330762
|
Polymorphic
|
0.88
|
11
|
10
|
42
|
135
|
IBN22
|
Monomorphic
|
0.00
|
1
|
0
|
44
|
206
|
STM1052
|
Polymorphic
|
0.85
|
13
|
13
|
48
|
|
64
|
CB330798
|
Polymorphic
|
0.57
|
3
|
2
|
42
|
136
|
IBN24
|
Polymorphic
|
0.82
|
7
|
2
|
44
|
207
|
STM1053
|
Polymorphic
|
0.73
|
4
|
2
|
60
|
|
65
|
CB330817
|
Polymorphic
|
0.93
|
19
|
14
|
42
|
137
|
IBN34
|
Polymorphic
|
0.85
|
9
|
8
|
42
|
208
|
STM1064
|
Polymorphic
|
0.82
|
7
|
6
|
45
|
|
66
|
CB330917
|
Polymorphic
|
0.91
|
16
|
14
|
42
|
138
|
IBN35
|
Polymorphic
|
0.81
|
7
|
7
|
44
|
209
|
STM1104
|
Polymorphic
|
0.88
|
9
|
9
|
60
|
|
67
|
IB 2-38
|
Polymorphic
|
0.76
|
6
|
5
|
42
|
139
|
IBN36
|
Polymorphic
|
0.91
|
12
|
12
|
48
|
210
|
STM1106
|
Polymorphic
|
0.60
|
5
|
3
|
60
|
|
68
|
Ib2/30
|
Polymorphic
|
0.84
|
7
|
2
|
44
|
140
|
IBN37
|
Polymorphic
|
0.70
|
4
|
3
|
42
|
211
|
STM5114
|
Polymorphic
|
0.82
|
9
|
9
|
45
|
|
69
|
IB2.45B
|
Polymorphic
|
0.64
|
3
|
3
|
42
|
141
|
IBR03
|
Polymorphic
|
0.66
|
3
|
1
|
46
|
212
|
STM5121
|
Polymorphic
|
0.87
|
9
|
9
|
48
|
|
70
|
Ib2-27
|
Polymorphic
|
0.88
|
10
|
7
|
44
|
142
|
IBR04
|
Polymorphic
|
0.78
|
6
|
5
|
46
|
213
|
STM5127
|
Polymorphic
|
0.50
|
5
|
5
|
48
|
|
71
|
Ib-242
|
Polymorphic
|
0.92
|
15
|
14
|
42
|
143
|
IBR08
|
Polymorphic
|
0.83
|
10
|
9
|
44
|
214
|
STPoAc58
|
Polymorphic
|
0.87
|
8
|
2
|
63
|
|
72
|
Ib2-248
|
Polymorphic
|
0.92
|
12
|
4
|
46
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Genetic Diversity Analysis: Based on the results of 193 polymorphic SSR markers, SHAN similarity matrix was used to generate an UPGMA dendrogram to study extent of genetic diversity among 12 potato cultivars (Fig 2). The similarity coefficient between 12 potato varieties varied from 0.69 to 0.84. As shown in Fig 2, the dendrogram divided 12 potato genotypes into two distinct groups. Cluster I is represented by light green color and comprised of five genotypes i.e. Faisalabad Red, Faisalabad White, SH-5, PRI Red, and Rubby. Similarly, Cluster II was represented by light blue color and is comprised of seven genotypes i.e. Sadaf, Sialkot Red, Sahiwal White, Sahiwal Red, Ravi Red, and FD 81-1. Cosmo did not form any cluster with rest of genotypes and remained separated (Fig 2).
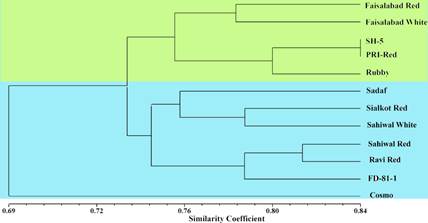
Fig 2. Dendrogram of 12 potato genotypes generated using data of 214 SSR markers through SHAN similarity matrix and unweighted pair group method with arithmetic mean clustering method.
Table 3. List of SSR markers that uniquely identify each genotype
|
Genotype Name
|
No of Markers
|
Marker
|
|
Faisalabad Red
|
12
|
STM1106, STM5127, Ib-24, Ib-286, IBE-7, IBS-17, BU691547, IBJ5446, BU690375, CB330200, CB330416, IBJ302
|
|
Faisalabad White
|
10
|
STM0019a, STM1052, STM5127, SSR38, IBCIP-5, IBR-13, CB330200, BM878740, CB330762, IBE32
|
|
SH-5
|
12
|
BU692227, IBSSR27, Ib-286, IBC-3, IBE-5, BU691268, IBJ1525, CB330554, BU692896, BU692095, IBE12, IBJ62
|
|
PRI-Red
|
3
|
STM5114, IBC-8, IBJ530
|
|
Ruby
|
15
|
BU692566, STM0031, STG0025, STI0012, IBN-35, BU691268, CB330283, CB330625, BU692896, CB330762, BU692914, IBE1, IBE15, IBE32, IBJ62
|
|
Sadaf
|
15
|
STM0019a, STM1052, STM5114, IBE-1, IBR-19, IBR-20, BU691268, IBJ559, CB330636, CB330771, BM8787757, CB330917, SSR270, IBCIP-1, IBJ5446
|
|
Sialkot Red
|
2
|
IBE-1, IBE-3
|
|
Sahiwal White
|
11
|
BU691949, IBR-21, IBE-5, IBE14, BU691143, IBJ559, CB330200, CB330601, BM878740, BU692739, IBN36
|
|
Cosmo
|
2
|
IBJ5446, BU690750
|
|
Sahiwal Red
|
4
|
STI0003, IBCIP-2, BU692858, IBN34
|
|
Ravi Red
|
6
|
IBN-21, CB330693, CB330285, CB330283, IBJ1798E, IBE34
|
|
FD 81-1
|
1
|
IBN-35
|
The maximum similarity was observed between SH-5 and PRI Red in cluster I sharing 84% of genetic loci. The lowest genetic similarity was observed between Cosmo and rest of 11 genotypes sharing 69% of genetic loci. There exists a domestic relationship between cultivar distribution and agro-ecological zone as observed in UPGMA dendrogram. Varieties Bred in Sahiwal i.e. Sadaf, Sahiwal White, Sahiwal Red, Ravi Red, Cosmo, FD 81-1 except PRI Red and Rubby shared same cluster. Similarly, varieties bred in Faisalabad i.e. Faisalabad White and Red also shared same clusters whereas varieties bred in Sialkot i.e. Sialkot White and SH-5 did not share same clusters (Fig 2).
In most of cases, dendrogram results fit well with pedigree/parentage information. Sialkot Red and Sahiwal White share a common parent (SH-5) and lie in same cluster. Sahiwal white and Sahiwal Red have both parents in common (FD 3-15 × SH-5). Both these genotypes share a common parent with Ravi Red (SH-5), Cosmo FD 81-1, and Sadaf (FD 3-15) hence all genotypes clustered together. However, in some cases, this was not true i.e. Sialkot Read and SH-5 have a common parent (Cardinal) but both genotypes are in different clusters. Faisalabad White and Faisalabad Red are in same clade but do not have a common parent. Sialkot White, Sialkot Red, Ravi Red, and Sahiwal Red are descendants of SH-5 but SH-5 lies in a separate cluster (Fig 2).
Population Structure of potato genotypes: Model-based cluster analysis using a Bayesian approach was carried out to infer population structure of twelve potato varieties using data of 193 polymorphic SSR markers. The LnP (D) scores for number of populations (K) increased up to 2 and showed inflation point at K2 which divides population into two groups. Similarly, ΔK-value also showed a peak at K = 2, indicating that genotypes comprised of two populations that are further subdivided into two subgroups. Population 1 (P1) contained five genotypes i.e. Faisalabad Red, Faisalabad White, SH-5, PRI Red, and Rubby. Whereas Population 2 (P2) contained seven genotypes Sadaf, Sialkot Red, Sialkot White, Cosmo, Sahiwal Red, Ravi Red, FD 81-8. Both populations were comprised of further two subgroups. The average distance between individuals in the same cluster (heterozygosity) was high in P2 (0.3008) as compared to P1 (0.2766). However genetic diversity in P1 was high with Fst value 0.16 as compared to P2 with Fst value 0.05. Results for Both K=2 and K=3 are given in Fig 3 for more accurate comparison (Fig 3)
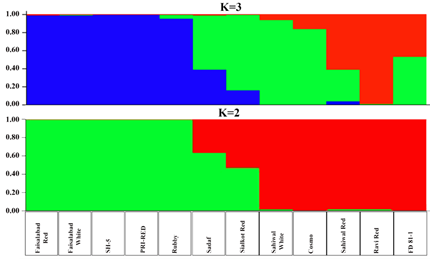
Fig 3. Structure Analysis of Potato varieties grown in Punjab Pakistan. Parameters: no admission model; K = 02; 10,000 Burn-in period; 100000 Rep.
DISCUSSION
Potato cultivar identification is of prime importance for germplasm maintenance and breeder’s right protection. Different types of DNA markers i.e. RFLPs, AFLPs, RAPDs, SSRs, and ISSRs were used in past for DNA fingerprinting, marker-assisted selection, phylogenetic and genetic diversity studies of potato (Onamu et al., 2016). Among these, SSR markers are more useful due to high polymorphism, ease of use, and high reproducibility (Jamil et al. 2020a, b; Iqbal et al., 2021b). Previously SSR markers have frequently been used for DNA fingerprinting, genetic diversity, and population structure studies of potato (Rodriguez-Bonilla et al., 2014; Song et al., 2016; Jian et al., 2017; Duan et al., 2019; Wang et al., 2019).
Among 226 markers, 193 polymorphic markers were used for further genetic analysis (Table 2). Up to our knowledge, no such studies were conducted in Pakistan for DNA fingerprinting of potato cultivars previously. However one report is available about genetic diversity studies but varieties used in that study are only six and RAPD markers were used which are not reproducible and reliable (Abbas et al., 2008). Unlike previous study of Duan et al., (2019), all varieties used in present study were distinguishable from each other on the basis of 72 SSR markers. Two SSR markers i.e. BU691268 (Fig 1) and IBJ5446 identified three varieties each whereas 14 markers distinguished two varieties each (Table 3). Our results support the findings of Jamil et al., (2020 a, b) that SSR markers are a powerful tool for genotyping and DNA fingerprinting of crops.
In our study alleles per locus (1 to 23 with average 8.04 alleles) and PIC value (0 to 0.96) were different from previous studies (Salimi et al., 2016; Song et al., 2016; Duan et al., 2019; Wang et al., 2019) because of the genetic background of varieties and SSR markers applied. Number of markers used in previous studies (20-30) was not comparable with (193 polymorphic SSR markers) our study as our study offered more genome coverage (Table 2). Duan et al. (2019) proposed a set of 11 SSR markers that are sufficient for discrimination of 217 potato cultivars. Similarly, Karaagac et al. (2014) proposed a set of six SSR markers for distinguishing of 50 tetraploid potato varieties. Different markers identified in our studies i.e. IBR13 (24 PA), BU690134 (22 PA), IBJ62 (21 PA), IBE14 (20 PA), and IBJ199 (18 PA) (Table 2) are a useful source for DNA fingerprinting and genetic diversity studies in future.
The genetic diversity studies with help of cluster (Fig 2) and population structure analysis (Fig 3) indicated narrow genetic makeup of potato genotypes explored in this study. Except for Cosmo, all other genotypes shared 72% genetic loci. However, Cosmo was dissimilar and shared 69% genetic loci with other 11 genotypes. The reason behind high similarity among genotypes is shared parentage. Except for Faisalabad White and Faisalabad Red, all other genotypes have one parent in common with any other genotype. SH-5 was used as a present in breeding of four genotypes. Similarly, FD 3-15 and FD 35-36 parents were used for breeding of four genotypes each. In extreme cases, Sahiwal Red and Sahiwal White both were evolved from the same cross (FD 35-36 × SH-5) due to colour difference only (Table 1).
At the time of selection breeders usually pay attention to phenotypes (a combination of genetics and environment) without considering genetic makeup (Braun and Wenzel, 2004). This is a leading reason for narrow genetic makeup of cultivated varieties of potato and other crops which expose crops to different disease and insects in the form of pandemics (Jamil et al., 2020c). For any successful breeding program information of genetic diversity in base material should be known and kept in consideration while designing a crossing plan. Crossing among genetically similar but phenotypically different genotypes has narrowed down genetic diversity and stagnant yield potential of cultivated crops including potato (Ray et al., 2012). The practice of using similar parents in breeding program, again and again, should be discouraged (Jamil et al., 2021b).
Conclusion: DNA fingerprints were developed for 12 potato genotypes grown in Punjab Pakistan. The genetic diversity studies using cluster and structure analysis grouped 12 potato genotypes into 2 distinct groups. Cluster I was comprised of 05 genotypes and cluster II was comprised of 6 genotypes whereas 01 genotype Cosmo did not obey clustering. The results of cluster and structure analysis were complementary to each other. The polymorphism information of 214 amplified SSR markers was also reported in this study. Our study will provide a platform for protection of Potato Breeders Rights and will help in varietal registration in Plant Breeder Rights Registry. We proposed five informative SSR markers i.e., IBR13, BU690134, IBJ6, IBE14, and IBJ199 for future DNA fingerprinting and genetic diversity studies.
Acknowledgements: The authors are highly thankful to the scientists of Potato Research Institute Sahiwal and its allied stations for providing plant material and necessary information. Mr. Baber Ali and Mr. Ahmad Shahzad Lab Assistants for technical assistance over time.
Funding detail: This work was supported by Punjab Agricultural Research Board under the Grant No. PARB 908.
REFERENCES
- Abbas, S. J., G. Rasool, S.R.U. Shah and A. Iqbal (2008). Analysis of genetic diversity in Pakistani potato cultivars by using randomly amplified polymorphic DNA (RAPD) primers. Amer.-Eur. J. Sus. Agri. 2(1): 50-53.
- Braun, A. and G. Wenzel (2004). Molecular analysis of genetic variation in potato (Solarium tuberosum ). I. German cultivars and advanced clones. Potato Res. 47(1-2): 81-92.
- Buteler, M., R. Jarret and D. LaBonten (1999). Sequence characterization of microsatellites in diploid and polyploid Ipomoea. Theor. App. Genet. 99(1-2): 123-132.
- Devaux, A., P. Kromann, O. Ortiz (2014). Potatoes for sustainable global food security. Potato Res. 57(3-4): 185-199.
- Duan, Y., J. Liu, J. Xu, C. Bian, S. Duan, W. Pang, J. Hu, G. Li, L. Jin (2019). DNA fingerprinting and genetic diversity analysis with simple sequence repeat markers of 217 potato cultivars (Solanum tuberosum) in China. Amer. J. Potato Res. 96(1): 21-32.
- Favoretto, P., E.A. Veasey, P.C.T.D. Melo (2011). Molecular characterization of potato cultivars using SSR markers. Horti. Brasileira 29(4): 542-547.
- Ghislain, M., J. Núnez, M.D.R Herrera, J. Pignataro, F. Guzman, M. Bonierbale, D.M. Spooner (2009). Robust and highly informative microsatellite-based genetic identity kit for potato. Mol. Breed. 23(3): 377-388.
- GOP 2020. Agriculture. In: Ministry of Food AaL ed. Economic Survey of Pakistan. Pakistan, Government of Pakistan. Pp. 1-23.
- He, C., V. Poysa, K. Yu (2003). Development and characterization of simple sequence repeat (SSR) markers and their use in determining relationships among Lycopersicon esculentum cultivars. Theor. App Genet. 106(2): 363-373.
- Hwang, S.Y., Y.T. Tseng, H.F. Lo (2002). Application of simple sequence repeats in determining the genetic relationships of cultivars used in sweet potato polycross breeding in Taiwan. Sci. Horti. 93(3-4): 215-224.
- Iqbal, M.Z., S. Jamil, A. Mehmood, R. Shahzad (2019). Identification of seven olive varieties using RAPD molecular markers. J. Agric. Res. 57(1): 07-14.
- Iqbal, M.Z., S. Jamil, R. Shahzad, K. Bilal, R. Qaisar, A. Nisar, S. Kanwal, M.K. Bhatti (2021a). DNA Fingerprinting of crops and its application in the field of plant breeding. J. Agric. Res. 59(1): 13-28.
- Iqbal, M.Z., S. Jamil, R. Shahzad, S.U. Rahman (2021b). DNA Fingerprinting and Cultivar Identification of Olive (Olea europaea L.) using SSR markers. Adv. Life Sci. 8(2): 143-148.
- Jamil, S., R. Shahzad, S. Kanwal, E. Yasmeen, S.U. Rahman, M.Z. Iqbal (2020a). DNA Fingerprinting and Population Structure of Date Palm Varieties Grown in Punjab Pakistan using Simple Sequence Repeat Markers. Int. J. Agri. Bio. 23(5): 943-950.
- Jamil, S., R. Shahzad, E. Yasmeen, S.U. Rahman, M. Younas, M.Z. Iqbal (2020b). DNA Fingerprinting of Pakistani Maize Hybrids and Parental lines using Simple Sequence Repeat Markers. J. Bot. 52(6): 2133-45.
- Jamil, S., R. Shahzad, S. Ahmad, R. Fatima, R. Zahid, M. Anwar, M.Z. Iqbal, and X. Wang, (2020c). Role of Genetics, Genomics and Breeding approaches to combat stripe rust of wheat. Front. Nut. Article Number 580715 https://doi.org/10.3389/fnut.2020.580715.
- Jamil, S., R. Shahzad, M.A. Iqbal, E. Yasmeen, S.U. Rahman (2021a). DNA fingerprinting and genetic diversity assessment of GM cotton genotypes for protection of plant breeders rights. Int. J. Agric. Biol. 25 (4): 768-776.
- Jamil, S., R. Shahzad, S.U. Rahman, M.Z. Iqbal, M Yaseen, S. Ahmad, R. Fatima (2021b). The level of Cry1Ac endotoxin and its efficacy against armigera in Bt cotton at large scale in Pakistan. GM Crops Food 12(1): 1-17.
- Jian, W., H. Lu, R.Y. Wang, M.M. He, Q.C. Liu (2017). Genetic diversity and population structure of 288 potato (Solanum tuberosum) germplasms revealed by SSR and AFLP markers. J. Integ. Agri. 16(11): 2434-2443.
- Kalia, R.K., M.K. Rai, S. Kalia, R. Singh, A. Dhawan (2011). Microsatellite markers: an overview of the recent progress in plants. Euphytica 177(3): 309-334.
- Kanwal, S., S. Jamil, R. Shahzad, S.U. Rahman, M.Z. Iqbal (2021). Standarization of different protocols for genomic DNA isolation from Phoenix dactylifera. Pak. J. Bot. 53(5): 1665-1668.
- Karaagac, E., S. Yilma, A. Cuesta-Marcos, M.I. Vales (2014). Molecular analysis of potatoes from the Pacific Northwest tri-state variety development program and selection of markers for practical DNA fingerprinting applications. Amer. J Potato Res. 91(2): 195-203.
- Lan, Q.K., Q.J. Li, J.B. Zhang, X.M. Zhao, Z. Zhu, R. Chen, S.Y. XU, Y. Wang, Y.Z. Guo (2012). Construction of SSR-based molecular fingerprinting and analysis of genetic diversity for celery varieties from Tianjin. Tianjin Agric. Sci. 18: 7-11.
- Majeed, A., Z. Muhammad (2018). Potato production in Pakistan: challenges and prospective management strategies–a review. Pak. J. Bot. 50(5): 2077-2084.
- Muñoz, M., C. Folch, F. Rodriguez, J. Kalazich, S. Orena, J. Santos, R. Vargas, A. Fahrenkrog, A. Puga (2016). Genotype number and allelic diversity overview in the national collection of Chilean potatoes. Potato Res. 59(3): 227-240.
- Onamu, R., J. Legaria, J. Rodríguez, J. Sahagùn, J. Pèrez (2016). Molecular characterization of potato (Solanum tuberosum) genotypes using random amplified polymorphic DNA (RAPD) and inter simple sequence repeat (ISSR) markers. Afr. J. Biotech. 15(22): 1015-1025.
- Pritchard, J.K., M. Stephens, P. Donnelly (2000). Inference of population structure using multilocus genotype data. Genetics 155(2): 945-959.
- Ray, D.K., N. Ramankutty, N.D. Mueller, P.C. West, J.A. Foley (2012). Recent patterns of crop yield growth and stagnation. Nat. Commun. 3(1): 1-7.
- Rodriguez-Bonilla, L., H.E. Cuevas, M. Montero-Rojas, F. Bird-Pico, D. Luciano-Rosario, D. Siritunga (2014). Assessment of genetic diversity of sweet potato in Puerto Rico. PloS one 9(12): e116184.
- Sadaf, N. (2018). Legislative Developments in the Protection of Plant Breeders’ Rights in Pakistan. IIC-International Review of Intellectual Property and Competition Law 49(6): 700-713.
- Salimi, H., M. Bahar, A. Mirlohi, M. Talebi (2016). Assessment of the genetic diversity among potato cultivars from different geographical areas using the genomic and EST microsatellites. Iranian J. Biotech. 14(4): 270.
- Shahzad, R., S. Jamil, S. Ahmad., A. Nisar, Z. Amina, S. Saleem, M.Z. Iqbal, R.M. Atif, X. Wang (2021a). Harnessing the potential of plant transcription factors in developing climate resilient crops to improve global food security: Current and future perspectives. Saudi J. Biol. Sci. 28(4): 2323-2341
- Shahzad, R., S. Jamil, S. Ahmad, A. Nisar, S. Kanwal R.M. Atif M.Z. Iqbal, Z. Ali, A.I. Ghazy and A.A. Khan (2021b). Omics approaches for improving abiotic stress tolerance in rice: recent advances and future prospects. Frontiers in Plant-Soil Interaction, Elsevier publisher Netherlands pp.199-220.
- Singh, A., A. Singh, U. MeenakshiArya, A. Singh (2019). Utility of molecular markers in molecular breeding for integrated crop improvement. Bulletin of Environment Pharmacology Life Sci. 8(2): 1-8.
- Song, X., C. Zhang, Y. Li, S. Feng, Q. Yang, S. Huang (2016). SSR analysis of genetic diversity among 192 diploid potato cultivars. Horti. Plant J. 2(3): 163-171.
- van Eck, H.J. (2007). Genetics of morphological and tuber traits. Potato Biology and Biotechnology, Elsevier. Pp. 91-115.
- Veilleux, R.E., L.Y. Shen, M.M. Paz (1995). Analysis of the genetic composition of anther-derived potato by randomly amplified polymorphic DNA and simple sequence repeats. Genome 38(6): 1153-1162.
- Vieira, M.L.C., L. Santini, A.L. Diniz, C.D.F. Munhoz (2016). Microsatellite markers: what they mean and why they are so useful. Genet. Mol. Bio. 39(3): 312-328.
- Vreugdenhil, D., J. Bradshaw, C. Gebhardt, F. Govers, M.A. Taylor, D.K. MacKerron, H.A. Ross (2011). Potato biology and biotechnology: Advances and Perspectives, Elsevier.
- Wang, Y., M.A.R. Rashid, X. Li, C. Yao, L. Lu, J. Bai, Y. Li, N. Xu, Q. Yang, L. Zhang (2019). Collection and evaluation of genetic diversity and population structure of potato landraces and varieties in China. Frontiers in Plant Science 10: 139. https://doi.org/10.3389/fpls.2019.00139.
|





