NUCLEAR LOCALIZATION OF ARABIDOPSISˈS HEAT SHOCK FACTOR HSFA1D USING BIMOLECULAR FLORESCENCE COMPLEMENTATION (BIFC) SYSTEM
Z. Shah1, G S. Ali2, A. El-Sayed3, M. Hussain1, H. Shah4, M. Z. Ahmad5, N. Ahmad 1and A. Iqbal6
1Department of Biotechnology, University of Science and Technology Bannu, KP, Pakistan; 2MREC-University of Florida, 2725 Binion Rd, Apopka, FL, USA 32703; 3Botany and Microbiology Department, Faculty of Science, Zagazig University Egypt; 4Plant Science Division, Pakistan Agriculture Research Council, Islamabad, Pakistan; 5Guangdong Provincial Key Laboratory of Plant Molecular Breeding, College of Agriculture, South China Agriculture University Guangzhou 510642, Gaungdong, China; 6Department of Botany, Islamia College Peshawar, KP, Pakistan. arshad.iqbal@icp.edu.pk
Corresponding author’s email: zamarud_gd@yahoo.com
ABSTRACT
Response to thermal stress in plants is mostly regulated by heat shock factors (hsfs). Among hsfs, HsfA1d is one of the key players in protecting plants against heat stress. Subcellular localization of gene is prerequisite for its characterization. Bimolecular Fluorescent Complementation (BiFC) provides the most advance and reliable tool to visualize protein-protein interactions and to explore their location in living system. Objective of the present study was to accomplish sub cellular localization of HsfA1d by using BiFC. HsfA1d was cloned in-frame with complementary halves of yellow florescent proteins (YFP) using gateway cloning system. The BiFC constructs along with positive control were introduced into plant cells through agro-infiltration. The non-florescent fragments of YFP tagged with HsfA1d produced no signal when they were infiltrated alone in tobacco leaves. The N and C termini of YFP fused with HsfA1d resulted in florescence upon co-expression in tobacco leaf epidermal cells. The intact YFP tagged with HsfA1d, after DAPI staining, was observed under 3 different laser beams of YFP, DAPI and merged. The same florescent signals detected under different channels of laser scanning confocal microscope confirmed the nuclear localization of HsfA1d. It is concluded, that HsfA1d is localized in the nucleus of cell.
Key words: HsfA1d, subcellular localization, yellow florescent proteins, gateway cloning, BiFC
https://doi.org/10.36899/JAPS.2021.4.0313
Published online December 18, 2020
INTRODUCTION
Proteins interact during every physiological process to perform their biological function. For example, DNA replication, assembly of translation apparatus, regulation of gene expression at different levels, DNA packaging and signal transduction involve protein-protein interaction (Scott et al. 2016). Thus, the investigation and characterization of various types of proteins interactions occur in the cell are important to understand the various aspect of cytology. Variety of techniques has been designed for the detection and analysis of proteins interactions both in vitro and in vivo (Xing et al. 2016). Every technique has its own pros and cons (Citovsky et al. 2006). For example, co-immunoprecipitation is one of the widely used methods to analyze invitro proteins interactions (Zhu et al. 2017). However, the requirements for specific antibodies against target proteins make it more expensive and laborious process. Moreover, Co-immunoprecipitation is associated with cell lysis which hinders the subcellular localization of the test proteins (Yang et al. 2014). Cross-linking and co-fractionation allows detection of in vivo protein–protein interactions, but the involvement of complex biochemical procedures makes it difficult to apply (Chen et al. 2010). Yeast two-hybrid (Y2H) is another assay used to visualize in vivo proteins interactions. The high occurrence of false-positives results and the non-availability of sub-cellular compartments for specific proteins interaction, add to the several drawbacks of this system (Mehla et al. 2017). Sos and Ras offers a different version of two hybrid system for detection of protein interaction. In this system, the complementary fragments of target protein (N & C) are fused to the cytoplasmic proteins of interest. The reconstitutions of such fragments during protein interaction degrade the URA3 reporter protein eventually leading to uracil auxotrophy and resistance to 5-fluoroorotic acid. Though, these assays do not need the entry of test proteins to the nucleus, yet drawback lies in allowing the interacting proteins to follow native patterns of subcellular localization (Cruz-Migoni et al. 2019). Bimolecular Fluorescent Complementation (BiFC) offers the most advance and novel approach to detect protein interactions directly within the living cell. Proteins interactions occur in any compartment of aerobic cell can be detected by using BiFC (Miller et al. 2015). In this assay, the desired protein is tagged with complementary fragments of florescent protein and the plant cells are transformed with both the constructs using agrobacterium. The association of non-florescent complementary fragments results in the restoration of florescent signals and eventually detected in the living plant cells (Kerppola, 2008). Initially, BiFC was limited to study the interacting transcription factors belong to bZIP and Rel family (Hu et al. 2002). Later on, the use of this assay was extended to detect protein interaction in mammalian (Hynes et al. 2004) bacterial (Tsuchisaka and Theologis, 2004) and plants cells (Stolpe et al. 2005). In plants, several expression vectors have been constructed to detect protein interaction in epidermal cells of tobacco and Arabidopsis (Walter et al. 2004). The BiFC vector, that were initially in use, suffered several drawbacks including inadequate cloning site, the requirement of two selectable markers for developing transgenic plants and non-availability of space for the expression of additional reporter genes (Bracha-Drori et al. 2004). To come across such problems, a large number of BiFC vectors with expanded MCS have been designed (Walter et al. 2004). The expanded MCS helps in tagging target proteins to complementary halves of auto florescent proteins. Maximum diversity in cloning machinery is important as precise association and integration of the interacting proteins is essential for reconstitution of florescent YFP. In addition to, such vector design provides an opportunity for the BiFC expression cassettes to use the agrobacterium binary plasmids for co-expression of interacting partners (Lai and Chiang, 2013). Moreover, these vectors help us to incorporate more autofluorescent proteins as reporter genes that may act as internal controls and markers of subcellular compartments. BiFC provides the opportunity to monitor transiently interacting proteins without the aid of any sophisticated equipment. Tobacco (Nicotiana benthamiana) offers a model system to be transformed with generated constructs and visualize protein-proteins interaction. Agrobacterium uses Ti plasmid (tumor inducing) as vehicle for the transport of target gene into the plant genome. The present study aimed at determining subcellular compartment of HsfA1d using gateway compatible BiFC system.
MATERIALS AND METHODS
Cloning of HsfA1d in BiFC Vectors: Arabidopsisˈs HsfA1d was tagged with N and C termini of YFP in BiFC vectors using LR reaction (Supp. 1). The LR reaction for cloning HsfA1d in frame with N terminus of YFP was initiated by mixing pUC57GW-HsfA1d (entry clone) and pGSA002-nYFPn (destination vector) in ratio of 4and 1 Fmoles respectively followed by the addition of clonaseII enzyme (0.5 µl) and placed for 1 hour at room temperature. Similarly, for cloning HsfA1d in frame with C terminus of YFP, the reactants (pUC57GW-HsfA1d and pGSA002-nYFPc) were added in 3.5:1Fmoles. The resultant mixture was kept for 1 hour at 25 ºC. Heat shock method was used to transform stellar cells with cloning product and applied on spectinomycin (50 mg/l) containing LB plates. The LB plates were incubated overnight at 37 ºC. Plasmids were extracted from transformed bacteria using plasmid extraction kit and subjected to digestion with restriction enzyme NOCI for confirmation. The digested DNA fragments were separated through electrophoresis using agarose gel (1%).
Preparation of Chemically Competent Agrobacteria (GV3101): Single colony of Agrobacteria was inoculated in 5 ml nutrient broth and subjected to overnight shaking (200 rpm) at 28 ºC. One ml of the resultant culture was mixed with fresh nutrient broth (50 ml) and grown at 28 ºC till the OD600 reached to 0.6. Further bacterial growth was stopped by keeping the culture on ice (30 min) and distributed in 2 sterile tubes equally. The bacterial culture was subjected to centrifugation (4000 g) for 5 min. The pellet was re-suspended in 15 ml CaCl2 (50 mM) and placed on ice for 15 min followed by centrifugation (3800 g) for 5 min. The pellet was re-suspended in 2 ml CaCl2 (50 mM) supplemented with glycerol (15%). Aliquots were made and stored at -80 ºC.
Transformation of competent Agrobacteria: Competent agro-bacterial cells (10 µl), after thawing, were mixed with plasmid DNA (1 µl) and placed on ice for 8 min. The cells were first subjected to cold treatment (-80 ºC for 5 min) followed by incubation for 5 min at 37 ºC and kept on ice for 1 min. Fresh nutrient broth (1 ml) was added to the cells and grown at 28 ºC under 200 rpm. The cells were centrifuged (8,000 g) for 1 min and the pellet was re-suspended in nutrient broth (50 μl). Cells were applied to nutrient agar plates with spectinomycin (50 mg/l) and placed for 2 days at 28 ºC. In this way, agrobacteria were transformed with cloned BiFC vectors along with pGWB442-YFP-HsfA1d (positive control).
Transient Transformation of Tobacco Leaves
Inoculums preparation: Single colony of transformed Agrobacteria was added to nutrient broth (5 ml) with spectinomycin (50 mg/l) and subjected to overnight shaking (200 rpm) at 28 ºC. The resultant culture was centrifuged (3000 g) for 5 min and the pellet was re-suspended in autoclaved water (20 ml) followed by re-centrifugation, using same rpm and time. The pellet was re-suspended in MMA solution (5 ml) and kept for 2 hours at 25 ºC. Final volume of inoculums was adjusted to 10 ml with OD600 of 0.6 using MMA solution. Equal volumes (5 ml) of agro inoculums harboring pGSA002-NYFP-HsfA1d and pGSA002-CYFP-HsfA1d were mixed.
Infiltration of tobacco leaves: For BiFC, two weeks old tobacco seedlings (Sup. 1) were divided into 4 groups. Each group of seedlings was infiltrated with 5 ml of different constructs i.e pGWB442-YFP-HsfA1d (positive control), pGSA002-NYFP-HsfA1d (negative control), pGSA002-CYFP-HsfA1d (negative control) and mixture of pGSA002-NYFP-HsfA1d+ pGSA002-CYFP-HsfA1d (test). The infiltrated seedlings were exposed to dark for 40 hours and observed for florescence. The florescent leaf segments (test samples) were stained with DAPI (4′,6-diamidino-2-phenylindole) reagent for 20 min and re-observed under 3 different channels ((YFP, DAPI and merged), using confocal microscopy.
Confocal microscopy: One cm of infiltrated leaf segment was placed on microscopic slide with upside down and about 30 μl H2O was added. The leaf piece was covered with cover glass and fixed using adhesive tape on both sides. Olympus laser scanning confocal microscope was used for images.
RESULTS
Heat shock factor (HsfA1d) was tagged with complementary halves of YFP using gateway cloning system (LR reaction; Fig 1A & B).
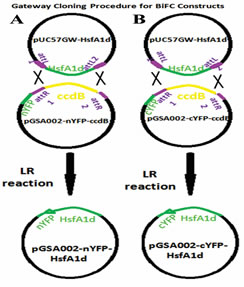
Figure1. Gateway procedure (LR reaction) for cloning HsfA1d in frame with, (A) N terminus of YFP and (B) C
terminus of YFP.
coli competent cells were transformed with product of the both LR reactions (Supp.2). Three fragments (10702, 1121 and 391 bp) were observed upon digestion of pGSA002-nYFP-HsfA1d with restriction enzyme NcoI while the empty vector (pGSA002-nYFP), used as control, resulted in 2 fragments (10557 and 1840 bp). Similarly, restriction digestion-based confirmation of pGSA002-cYFP-HsfA1d with NcoI produced 3 fragments (10702, 797 and 391 bp) while 2 fragments (10557 and 1516 bp) were observed when the control vector (pGSA002-cYFP) was subjected to NOCI as shown in the Supp.3. The BiFC vectors where HsfA1d has been cloned in frame with N & C termini of YFP shown in the Figure 2A & B.

Figure 2. Cloned BiFC vectors, (A) pGSA002-nYFPn-HsfA1d and (B) pGSA002-nYFPc-HsfA1d.
CaMV35S = Promoter, Spectinomycin adenyletransferase= bacteria marker, NOS =
terminator sequence, EYFPn= N terminus of YFP, EYFPc = C terminus of YFP,HsfA1d= gene
of interest
Competent Agrobacterial cells were successfully transformed with pGSA002-nYFP-HsfA1d and pGSA002-cYFP-HsfA1d (Supp. 4). Tobacco leaves infiltrated pGWB442-YFP-HsfA1d, used as positive control, shown florescence under confocal microscope (Fig. 3A) while no florescence was recorded when pGSA002-nYFP-HsfA1d and pGSA002-cYFP-HsfA1d were expressed alone in tobacco leaf samples (Fig. 3B & C). Similarly, florescence was observed when pGSA002-nYFP-HsfA1d and pGSA002-cYFP-HsfA1d were co-expressed in tobacco leaves (Fig 3D).

Figure 3. BiFC based detection of HsfA1d in N. benthamiana leaf epidermal cells, (A) YFP tagged with HsfA1d (positive control), (B) HsfA1d tagged with N terminus of YFP (negative control), (C) HsfA1d tagged with C terminus of YFP (negative control) and (D) Co-expression of N and C termini of YFP tagged with HsfA1d (test) Scale bar = 10 µm.
The tobacco epidermal cells, detected positive for florescence, revealed florescence under observing 3 laser beams (YFP, DAPI and merged) using confocal microscopy (Fig. 4).

Figure 4. Nuclear localization of HsfA1d in N. benthamiana leaf epidermal cell, Detection of interacting N and C termini of YFP tagged with HsfA1d under, (A) YFP channel, (B) DAPI channel and (C) Merged channel
DISCUSSION
The integration of Gateway cloning system and BiFC provide powerful tool to the researchers for understanding protein interactions in the cells. Gateway cloning technique allows the genetic engineers to rapidly design a variety of vectors with high degree of precision (Karna et al. 2018) while the BiFC helps in detection of protein-protein interaction and their subcellular localization (Gookin and Assmann, 2014).The attachment (att) sites lies on the vectors provide a platform for recombination that ultimately leads to gene shuffling (Reece-Hoyes and Walhout, 2018). In the current study, recombination at the attachment sites present on entry clone (aaL1, attL2) and destination vector (attR1, attR2) helped in cloning target gene through LR reaction. The BiFC vectors (pGSA002-nYFP-HsfA1d and pGSA002-cYFP-HsfA1d) constructed in the present study strengthened the earlier reports of Karimi et al. (2007), that gateway accurately clone gene cassettes into multiple vectors. Agrobacterium mediated infiltration was successfully used for introduction of BiFC vectors into the tobacco plant. These results are in line with the reports of Li et al. (2005). The mentioned results are also in agreement with Leckie and Stewart (2011) that agrobacteria extend valuable source for the transport of target gene into host plant. Epidermal cells of 4 weeks old Nicotiana benthamiana were transformed with HsfA1d tagged with YFP. Similar results exhibited by Schweiger and Schwenkert (2014), further endorsed the utility of Nicotiana benthamiana as model system for transient expression of the target gene. The young age of explants, inoculums OD (0.6) and post infiltration period (42 hours) contributed in successful delivery of HsfA1d fused with auto-florescent protein into tobacco (Bernaudat et al. 2011). Among advanced techniques, BiFC represents one of the most powerful tools for detection and studying protein interactions in living cell (Citovsky et al. 2006). The foundation of this assay lies on reconstitution of fluorescent complex of two complementary fragments of YFP fused to the interacting molecules (Schütze et al. 2009). In the current study, the co-expression of cloned BiFC vectors (pGSA002-HsfA1d-nYFPand pGSA002-HsfA1d-cYFP) in tobacco cells, extended opportunity for non-florescent C and N termini to reconstitute YFP and hence signals were detected. Similar results were also shown by Horstman et al. 2014. No florescence was observed when these vectors were expressed alone, as negative control, in the host cells. These results are in agreement with the reports of Peter et al., (2017). Signals were detected when pGWB442-HfsA1d tagged with intact YFP was expressed in epidermal cells as positive control. The leaf segments already detected positive for YFP were stained with DAPI and examined under 3 distinct laser beams (DAPI, YFP, merged) for florescence. As DAPI only binds to the nuclear DNA, therefore appearance of florescent proteins tagged with target gene, on the same location under 3 different laser beams accomplished that HsfA1d lies in nuclear compartment of the cell. Hernández-Sánchez et al. (2015), ended up with same results upon exposing the host tissues to nuclear dye (DAPI) for subcellular localization of the target gene.
Acknowledgments: The Research facilities were provided by MREC University of Florida USA while financial support was extended by Higher Education Commission Pakistan and University of Science and Technology, Bannu, Pakistan.
Author’s contribution: Zamarud Shah conducted the main experiment and collected data. Gul Shad Ali supervised the research. Arshad Iqbal and Hussain Shah helped in manuscript writing. Muhammad Zulfiqar Ahmed and Masroor Hussain contributed in figure setting and data analysis. Ashrf El-Sayed and Nisar Ahmad helped in making the revised manuscript.
Conflict of interest: The authors declare that they have no conflict of interest
REFERENCES
- Bernaudat, F., A. Frelent-Barrand, N. Pochon, S. Dementin, P. Hivin, S. Boutigny, J. Rioux, D. Salvi, D. Seigneurin-Berni, P. Richaud, J. Joyard, D. Pignol, M. Sabaty, T. Desnos, E. Peby-Peyroula, E. Darrouzet, T. Vernet and N. Rolland. (2011). Heterologous expression of membrane proteins: choosing the appropriate host. PLOS ONE. 6: 12–20.
- Bracha-Drori, K., K. Shichrur, A. Katz, M. Oliva, R. Angelovici, S. Yalovsky and N. Ohad (2004). Detection of protein-protein interactions in plants using bimolecular fluorescence complementation. J. Plant. 40: 419–427.
- Chen, Z. A., A. Jawhari, L. Fischer, C. Buchen, S. Tahir, T. Kamenski, M. Rasmussen, L. Lariviere, J.Bukowski Wills, M. Nilges, P. Cramer and J. Rappsilber (2010). Architecture of the RNA polymerase II–TFIIF complex revealed by cross‐linking and mass spectrometry. EMBO J. 29: 717–726.
- Citovsky,V., L. Lee, S. Vyas, E. Glick, M. Chen, A. Vainstein, Y. Gafni, S.B. Gelvin and T. Tzfira (2006). Subcellular localization of interacting proteins by bimolecular fluorescence complementation in planta. J. M. Biol. 362 (5): 1120–1131.
- Cruz-Migoni, A., P. Canning, C. E. Quevedo, C.J.R. Bataille, N. Bery, A. Miller, A.J. Russell, S.E.V. Phillips, S.B. Carr and T.H. Rabbitts (2019). Structure-based development of new RAS-effector inhibitors from a combination of active and inactive RAS-binding compounds. PNAS 116 (7): 2545–2550
- Gookin, T.E. and S.M. Assmann (2014). Significant reduction of BiFC non-specific assembly facilitates in planta assessment of heterotrimeric G-protein interactors. Plant J. 80: 553–567
- Horstman, A., I. Tonaco, K. Boutilier and R. Immink (2014). A cautionary note on the use of split-YFP/BiFC in plant protein-protein interaction studies. Int J Mol Sci. 15: 9628–9643
- Hu, C.D., Y. Chinenov and T.K. Kerppola (2002). Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Cell. 9: 789–798.
- Hynes, T.R., L. Tang, S.M. Mervine, J.L. Sabo, E.A. Yost, P.N. Devreotes and C.H. Berlot (2004). Visualization of G protein beta gamma dimers using bimolecular fluorescence complementation demonstrates roles for both beta and gamma in subcellular targeting. J. Biol. Chem. 279: 30279–30286.
- Hernández-Sánchez, I.E., I. Maruri, A. Ferrando, J. Carbonell, S.P. Graether and J.F. Jimenez Bremont (2015). Nuclear localization of the dehydrin Opsdhn1 is determined by histidine-rich motif. Front Plant Sci. 6: 702
- Karimi, M., A. Depicker, and P. Hilson (2007). Recombinational Cloning with Plant Gateway Vectors. Plant Physiol 145(4): 1144-54.
- Karna, S.R., X. Zogaj, J.R. Barker, J. Seshu, S.L. Dove and K.E. Klose (2018). A bacterial two-hybrid system that utilizes Gateway cloning for rapid screening of protein-protein interactions. Biotechniques. 49 (5): 831-833
- Kerppola, T.K. (2008). Bimolecular fluorescence complementation (BiFC) analysis as a probe of protein interactions in living cells. Annu Rev Biophys. 37: 465–87.
- Leckie, B.M. and C.N. Stewart (2011). Agroinfiltration as a technique for rapid assays for evaluating Candidate insect resistance transgenes in plants. Plant Cell Rep. 30: 325–334
- Lai, H., and C. Chiang (2013). Bimolecular Fluorescence Complementation (BiFC) Assay for Direct Visualization of Protein-Protein Interaction in viv. Bio Protoc. 3(20): e935.
- Li, J., A. Krichevsky, M. Vaidya, T. Tzfira and V. Citovsky (2005). Uncoupling of the functions of the Arabidopsis VIP1 protein in transient and stable plant genetic transformation by Agrobacterium. Proc. Natl Acad. Sci. USA 102: 5733–5738.
- Mehla, J., J.H.Caufield, Sakhawalkar, and P.Uetz (2017). Chapter Seventeen - A Comparison of Two-Hybrid Approaches for Detecting Protein–Protein Interactions. Method Enzymol. 586: 333–358.
- Miller, K. E., Y. Kim, W. Huh, and H. Park (2015). Bimolecular fluorescence complementation (BiFC) analysis: advances and recent applications for genome-wide interaction studies. J Mol Biol. 427(11): 2039–2055.
- Reece-Hoyes, J.S., and A.J.M. Walhout (2018). Gateway Recombinational Cloning. Cold Spring Harb Protoc.
- 2018(1): pdb. top094912.
- Peter, S., S.Z. Oven-Krockhaus, M. Veerabagu, V.M. Rodado, K.W. Berendzen, A.J. Meixner, K. Harter and F.E. Schleifenbaum (2017). Chimeric Autofluorescent Proteins as Photophysical Model System for Multicolor Bimolecular Fluorescence Complementation. J Phys Chem B. 121(11): 2407-2419.
- Schütze, K., K. Harter,and C. Chaban (2009).Bimolecular Fluorescence Complementation (BiFC) to Study Protein–Protein Interactions in Living Plant Cells. Methods Mol Biol. 479: 189–202.
- Schweiger, R., and S. Schwenkert (2014). Protein-protein interactions visualized by bimolecular fluorescence complementation in tobacco protoplasts and leaves. J. Vis. Exp. (85): e51327.
- Scott, D. E., A.R. Bayly, C. Abell and J. Skidmore (2016). Small molecules, big targets: drug discovery faces the protein–protein interaction challenge. Nat. Rev. Drug Discov. 15: 533–550.
- Stolpe,T., C. Süsslin, K. Marrocco, P. Nick, T. Kretsch, and S. Kircher (2005). In planta analysis of protein–protein Interactions related to light signaling by bimolecular fluorescence complementation. Protoplasma. 226: 137–146.
- Tsuchisaka, A., and A. Theologis (2004). Heterodimeric interactions among the 1-amino-cyclopropane-1-carboxylatesynthase polypeptides encoded by the Arabidopsis gene family. Proc. Natl Acad. Sci. USA. 101: 2275–2280.
- Walter, M., C. Chaban, K. Schütze, O. Batistic, K. Weckermann, C. Näke, D. Blazevic, C. Grefen, K. Schumacherand C. Oecking(2004). Visualization of proteininteractions in living plant cells using bimolecular fluorescence complementation. Plant. 40: 428–438.
- Xing, S., N. Wallmeroth, K.W. Berendzen and C. Grefen (2016). Techniques for the Analysis of Protein-Protein Interactions in Vivo. Plant Physiol. 171(2): 727–758.
- Yang, J.W., J.X. Fu, J. Li, X.L. Cheng, F. Li, J. F. Dong, Z.L. Liu, and C.X. Zhuang (2014). A Novel Co- immunoprecipitation Protocol Based on Protoplast Transient Gene Expression for Studying Protein– protein Interactions in Rice. Plant Mol. Biol. Rep 32(1): 153–161
- Zhu, X., A. Zelmer, and S. Wellmann (2017). Visualization of Protein-protein Interaction in Nuclear and Cytoplasmic Fractions by Co-immunoprecipitation and In Situ Proximity Ligation Assay. J Vis Exp. (119):
SUPPLEMENTARY DATA
|
Supp 1. Tobacco leaves used for infiltration of GV3101 harboring BiFC constructs
|
|
Supp 2. E. coli cells transformed with, (a) pGSA002-nYFPn-HsfA1d and (b) pGSA002-nYFPc-HsfA1d
|
|
Supp 3. Agrobacterium GV3101 cells transformed with, (a) pGSA002- nYFPn-HsfA1d and (b) pGSA002-nYFPc-HsfA1dHsfA1d
nYFPn-HsfA1d and (b) pGSA002-nYFPc-HsfA1d.
|
|
Supp 4. Restriction digestion of cloned BiFC vectors pGSA002-nYFPn-HsfA1d and pGSA002- nYFPc-HsfA1d along with their empty vectors (control) using enzyme NcoI.
L= 1kb plus DNA ladder, 1= pGSA002-nYFPn, 2= pGSA002-nYFPn-HsfA1d, 3 = pGSA002-nYFPn-HsfA1d, 4= pGSA002-nYFPcand 5= pGSA002-nYFPc-HsfA1d
|
|





