MORPHOLOGICAL AND MORPHOMETRIC ANALYSIS OF GENUS ERIOBOTRYA LINDL. (ROSACEAE)
M. Idrees1, 2, M. L. Pathak1,, N. H. Memon2, S. Khan3, Z. Y. Zhang2*and X. F. Gao1*
1CAS Key Laboratory of Mountain Ecological Restoration and Bioresource Utilization & Ecological Restoration and Biodiversity Conservation Key Laboratory of Sichuan Province, Chengdu Institute of Biology, Chinese Academy of Science, P.O Box 416, Chengdu 61004, China.
2College of Life Science, Neijiang Normal University, Neijiang 641000, Sichuan, China
3School of Geography and Resources Science, Neijiang Normal University, Neijiang 641000, Sichuan, China
* Corresponding author’s email: xfgao@cib.ac.cn, zhangzyong219@126.com
ABSTRACT
Morphometric analysis based on 40 morphological (quantitative and qualitative) characters were used to evaluate the patterns of morphological variations and to better understand the taxonomic relationship of the whole genus of Eriobotrya. Cluster analysis, principal component analysis and principal coordinate analysis were performed for both quantitative and qualitative morphological characters. Result of factor analysis revealed that leaf blade length and width, bract shape, bract length and width, sepal length, petal length and width, petal state, style number, fruit length and width were highly positive significant, whereas pedicel length and style state were highly negative significant. Cluster analysis revealed that all the taxa of the genus grouped into 4 clusters. Cluster 1 included 18 taxa (E. bengalensis, E. bengalensis var. angustifolia, E. bengalensis var. intermedia, E. stipularis, E. dubia, E. serrata, E. obovata, E. poilanei, E. elliptica var. petelotii, E. longifolia, E. angustifolia, E. petiolata, E. deflexa, E. deflexa var. buisanensis, E. kwangsiensis, E. fulvicoma, E. cavaleriei and E. glabrescens). Cluster II included 16 taxa (E. henryi, E. seguinii, E. hookeriana, E. japonica, E. × daduheensis, E. malipoensis, E. elliptica, E. grandiflora, E. fragrans, E. fragrans var. furfuracea, E. prinoides, E. prinoides var. laotica, E. salwinensis, E. tengyuehensis, E. latifolia and E. glabrescens var. victoriensis). Two species (E. wardii and E. platyphylla) formed a distinct cluster and considered to be doubtful to the genus. The quantitative and qualitative characters studies will be particularly useful for further identification and systematic of the genus Eriobotrya.
Keywords: Cluster analysis, Eriobotrya, morphometric analysis, species, systematic, taxonomy.
https://doi.org/10.36899/JAPS.2021.4.0306
Published online January 04, 2021
INTRODUCTION
Morphological characterization has been widely used for taxonomic studies as well as plant classifications (Sokal and Sneath 1973). The method of cluster analysis (CA) based on similarity matrixes which produce hierarchical classification of taxa. Principal component analysis (PCA) used for reducing the dimensions of the original data and Principal coordianate analysis (PCoA) indicate the position of point relative to each other’s, both are commonly used for screening the accession collections, taxonomic relationships and botanical systematic (Badenes et al. 2000). However, such morphometric studies have been conducted in the genera of Potamogeton (Kaplan and Marhold 2012), Clerodendrum L. (Deshmukh et al. 2012), Hedera L. (Ackerfield and Wen 2002).
Eriobotrya Lindl. (1821), is a genus belongs to the family Rosaceae (Potter et al. 2007), comprises about more than 30 taxa, and are widely distributed in China and southeast Asia (Vidal 1965, Yang et al. 2005). Taxon relationships, delimitations and classification within this genus are controversial and unclear. Several early treatments for the classification of Eriobotrya were based on leaf morphology (Yang 2005). Yu (1974) classified (13 taxa of the genus) them into 2 groups, based on leaf blade tomentose and without tomentose. Zhang et al. (1990) classified (15 taxa of the genus) them into 2 groups, based on autumn and winter flowering group. Gu and Spongberg (2003) described 14 species of Eriobotrya and classified them in China. Yang et al. (2007) proposed a new classification system and divided (15 taxa of the genus) into 3 groups, based on style number, stamens and leaf blade size. All of these studies were based on Eriobotrya species which is mostly recorded in China, whereas the species found in Southeast Asia were not studies and investigated. Recently, Yang et al. (2017) classified (22 taxa of the genus) them into 3 groups, based on leaf length (small, medium, and large size). Zhang et al. (2017) classified (25 taxa of the genus) them into 5 group based on 53 morphological characters. These studies were included some of the taxa from Southeast Asia, but the completed classification and species relationships of the whole genus are still unclear.
The aims of the present study are to use a standard statistical method of multivariate analysis: 1) to clarify morphological patterns of variations within the whole genus of Eriobotrya; 2) to reveal the most informative characters for the determination of taxa, and to assess their taxonomic relationship.
MATERIALS AND METHODS
The present study was based on 36 taxa of Eriobotrya (Table 1), with more than 300 specimens were analyzed. The examined number of specimens for each taxon ranged from 2 to 20, depending upon availability and a suitable structure of both vegetative and reproductive characters were included to decrease missing data for analysis. Although in each taxa, the type specimens were examined. These specimens deposited in the relevant herbaria (A, B, BM, CDBI, CANT, HITBC, HENU, HUH, HGAS, IBK, IBSC, K, KATH, KUN, K, L, LINN, M, N, NAS, NYBG, P, PE, SYS, TAIF, U, US, UPS, WU, WUK, YU).
A total of 40 quantitative and qualitative morphological characters were recorded and included in the numerical analysis (Table 2). Each specimen characters were measured 4-8 time and means were recorded. Measurements were performed on each specimen, using ImageJ software or a ruler. The macro-morphological and micro-morphological observations were analyzed and examined under optical microscope (Stemi DV4 and LEICA S8 AP0).
Cluster analysis and scatter plot analysis were performed using PAST software (Hammer et al. 2001). A principal coordinate analysis was conducted on the basis of all morphological characters by using the methodology of Nobis et al. (2016). A principal component analysis was performed on the basis of correlation matrix (Sokal and Sneath 1963). Each accession was marked with the symbol on the scatter plot. Factors with eigenvalues >1 were chosen according to the Kaiser criterions (Kaiser 1960). Quantitative data were analyzed to determined mean analysis, analysis of variance, standard variation and coefficient of variation using SPSS software.
Table 1. List of investigated taxa and their type specimens, herbarium and collection site
|
No
|
Taxa
|
Type specimens
|
Herbaria with barcode
|
Collection site
|
|
1
|
E. angustissima Hook.f.
|
J.D. Hooker & T. Thomson s.n.
|
K000758406
|
India, Khasia
|
|
2
|
E. bengalensis (Roxb.) Hook.f.
|
Wallich 668/2
|
K001111550
|
India
|
|
3
|
E. bengalensis (Roxb.) Hook.f. var. angustifolia (Card.) J.E. Vidal
|
F. Ducloux 4719
|
P02143257
|
China, Yunnan
|
|
4
|
E. bengalensis (Roxb.)Hook.f. var. intermedia J.E. Vidal
|
G. Forrest 17845
|
E00072977
|
Myanmar
|
|
5
|
E. cavaleriei (H.Lév.) Rehder
|
J. Cavalerie 3220
|
E00011330
|
China, Guizhou
|
|
6
|
E. × daduheensis H.Z. Zhang ex W.B. Liao, Q. Fan & M.Y. Ding
|
Q. Fan, 9292
|
SYS00168451
|
China, Sichuan
|
|
7
|
E. deflexa (Hemsl.) Nakai
|
A. Henry 498
|
K000758389
|
Taiwan, Formosa
|
|
8
|
E. deflexa (Hemsley) Nakai var. buisanensis (Hayata) Hayata
|
S. Sasak s.n.
|
TAI!
|
Taiwan, Buizan
|
|
9
|
E. dubia (Lindl.) Decne.
|
Wallich 668/1
|
K001111549
|
Nepal
|
|
10
|
E. elliptica Lindl.
|
F. Buchanan-Hamilton s.n.
|
BM000521994
|
Nepal, Narainhetty
|
|
11
|
E. elliptica Lindl. var. petelotii J.E. Vidal
|
Petelot s.n.
|
P02143261
|
Vietnam, Kao Lay
|
|
12
|
E. fragrans Champ. ex Benth.
|
J.G. Champion s.n.
|
K000758384
|
China, Hong Kong
|
|
13
|
E. fragrans Champ. ex Benth. var. furfuracea J.E. Vidal
|
A. Chevalier 38893
|
P02143263
|
Vietnam, Nha Trang
|
|
14
|
E. fulvicoma W. Y. Chun ex W.B. Liao, F.F. Li & D.F. Cui
|
Z. Huang 32257
|
WUK0109531
|
China, Guangdong
|
|
15
|
E. grandiflora Rehder & E.H. Wilson
|
Veitch Exped. 3506
|
A00026472
|
China, Sichuan
|
|
16
|
E. glabrescens J.E. Vidal
|
F. Kingdon-Ward 20616
|
BM000602189
|
Myanmar, Burma
|
|
17
|
E. glabrescens J.E. Vidal var. victoriensis J.E. Vidal
|
F. Kingdon-Ward 21915
|
BM000602190
|
Myanmar, KachinState
|
|
18
|
E. henryi Nakai
|
A. Henry 13018
|
A00026474
|
China, Yunnan
|
|
19
|
E. hookeriana Decne.
|
M.D. Anderson 490
|
P02143268
|
India, Sikkim
|
|
20
|
E. japonica (Thunb.) Lindl.
|
Thunberg s.n.
|
UPS-THUNB11908
|
Japan
|
|
21
|
E. kwangsiensis Chun
|
Z. Huang 39423
|
IBK00061038
|
China, Guangxi
|
|
22
|
E. latifolia Hook.f.
|
T. Lobb s.n.
|
K000758400
|
Myanmar, Moalmayne
|
|
23
|
E. longifolia (Decne.) Hook.f.
|
Griffith 2093
|
P02143220
|
Bangladesh, Mishmi Hills
|
|
24
|
E. malipoensis K.C. Kuan
|
C.W. Wang & Y. Liu 86318
|
PE00004573
|
China, Yunnan
|
|
25
|
E. obovata W.W. Sm.
|
E.E. Maire 2450
|
E00011331
|
China, Yunnan
|
|
26
|
E. petiolata Hook.f.
|
J.D. Hooker s.n.
|
K000758394
|
India, Sikkim
|
|
27
|
E. platyphylla Merr.
|
F. Kingdon-Ward 10205
|
A00026485
|
Myanmar, Upper Burma
|
|
28
|
E. poilanei J.E. Vidal
|
E. Poilanei 22591
|
P02143226
|
Vietnam, Haut Donnai
|
|
29
|
E. prinoides Rehder & E.H. Wilson
|
A. Henry 9878
|
A00026476
|
China, Yunnan
|
|
30
|
E. prinoides Rehder & E.H. Wilson var. laotica J.E. Vidal
|
E. Poliane 2243
|
P02143229
|
Laos, Xieng Khouang
|
|
31
|
E. salwinensis Hand.-Mazz.
|
H.R.E. von Handel-Mazzetti 9573
|
WU0059392
|
China, Yunnan
|
|
32
|
E. seguinii (H.Lév.) Cardot ex Guillaumin
|
J. Séguin & R.P. Bodinier 2617
|
E00011359
|
China, Guizhou
|
|
33
|
E. serrata J.E. Vidal
|
E. Poilane 2345
|
P02143235
|
Laos, Xièng Khouang
|
|
34
|
E. stipularis Craib
|
A.F.G. Kerr 14125
|
K000758408
|
Thailand, Siam
|
|
35
|
E. tengyuehensis W.W. Sm.
|
G. Forrest 9857
|
E00011333
|
China, Yunnan
|
|
36
|
E. wardii C.E.C. Fisher
|
F. Kingdon-Ward 7618
|
K000758392
|
Myanmar, Namkia Moutains
|
Table 2. Morphological characters, taxonomic code, mean/average, standard deviation and variance measure in the genus Eriobotrya.
|
No
|
Character
|
Unit
|
Taxonomic Code
|
Min.
|
Max.
|
Mean
|
st. deviation
|
Variance
|
|
|
1
|
Leaf blade length
|
Cm
|
|
4.43
|
29.83
|
15.2384
|
5.85
|
34.2
|
|
|
2
|
Leaf blade width
|
Cm
|
|
1.58
|
15.36
|
5.66
|
3.01
|
9.12
|
|
|
3
|
Petiole length
|
Cm
|
|
entire or .5
|
4.49
|
2.2676
|
1.114
|
1.24
|
|
|
4
|
Lateral vein
|
Count
|
<10 (0); 10-20 (1); >20 (2)
|
|
|
|
|
|
|
|
5
|
Leaf serration
|
Count
|
<15 (0); 15-30 (1); >30 (2)
|
|
|
|
|
|
|
|
6
|
Blade length/Blade width
|
Ratio
|
|
|
|
|
|
|
|
|
7
|
Petiole Length/Blade width
|
Ratio
|
|
|
|
|
|
|
|
|
8
|
Leaf blade abaxially tomentose
|
Code
|
Glabrous (0); Tomentose (1)
|
|
|
|
|
|
|
|
9
|
Petiole state
|
Code
|
Glabrous (0); Tomentose (1)
|
|
|
|
|
|
|
|
10
|
Inflorescence size
|
Cm
|
|
2.42
|
16.77
|
8.06
|
3.46
|
12.03
|
|
|
11
|
Inflorescence type
|
Code
|
Contracted (0);
Spreading (1)
|
|
|
|
|
|
|
|
12
|
Bract shape
|
Code
|
Linear or lanceolate (0);
Ovate (1), Other (2)
|
|
|
|
|
|
|
|
13
|
Bract length
|
Cm
|
|
0.2
|
0.61
|
0.3532
|
0.117
|
0.014
|
|
|
14
|
Bract width
|
Cm
|
|
0.04
|
0.33
|
0.1548
|
0.079
|
0.006
|
|
|
15
|
Pedicel length
|
Cm
|
|
sessile or 0
|
0.66
|
0.2239
|
0.1666
|
0.028
|
|
|
16
|
Pedicel state
|
Code
|
Glabrous (0); Tomentose (1)
|
|
|
|
|
|
|
|
17
|
Calyx (glabrous or tomentose)
|
Code
|
Glabrous (0); Tomentose (1)
|
|
|
|
|
|
|
|
18
|
Sepal shape
|
Code
|
Triangular or trangular ovate (0); Ovate or rounded (1)
|
|
|
|
|
|
|
|
19
|
Sepal length
|
Cm
|
|
0.13
|
0.35
|
0.2216
|
0.0617
|
0.004
|
|
|
20
|
Sepal width
|
Cm
|
|
0.09
|
0.65
|
0.1873
|
0.094
|
0.009
|
|
|
21
|
Sepal inside surface
|
Code
|
Glabrous (0); Tomentose (1)
|
|
|
|
|
|
|
|
22
|
Sepal outside surface
|
Code
|
Glabrous (0); Tomentose (1)
|
|
|
|
|
|
|
|
23
|
Petal shape
|
Code
|
Obovate (0); Ovate (1);
other (2)
|
|
|
|
|
|
|
|
24
|
Petal at base
|
Code
|
Glabrous (0); slightly pilose or woolly (1); Villous (2)
|
|
|
|
|
|
|
25
|
Petal Length
|
Cm
|
|
0.27
|
0.94
|
0.562
|
0.188
|
0.036
|
|
26
|
Petal width
|
Cm
|
|
0.2
|
0.64
|
0.3835
|
0.132
|
0.018
|
|
27
|
Petal length/Petal width
|
Ratio
|
|
|
|
|
|
|
|
28
|
Indulendum of Ovary
|
Code
|
Glabrous (0); Villous (1)
|
|
|
|
|
|
|
29
|
Style number
|
Code
|
Consistently 2 (0); 2-4 (1); upto 5 (2), consistently 5 (3)
|
|
|
|
|
|
|
30
|
Style state
|
Code
|
Free (0); Connate/fused (1)
|
|
|
|
|
|
|
31
|
Style at base
|
Code
|
Glabrous (0); Villous (1)
|
|
|
|
|
|
|
32
|
Fruit length
|
Cm
|
|
1.25
|
2.01
|
1.522
|
0.204
|
0.042
|
|
33
|
Fruit width
|
Cm
|
|
0.53
|
2.29
|
1.277
|
0.4529
|
0.205
|
|
34
|
Fruit length/Fruit width
|
Ratio
|
|
|
|
|
|
|
|
35
|
Tree habit
|
M
|
<5 (0); 5-10 (1); >10 (2)
|
|
|
|
|
|
|
36
|
Stamens
|
Code
|
10-15 (0); 15-20 (1); >20 (2)
|
|
|
|
|
|
|
37
|
Stipule state
|
Code
|
Cuducous (0); Persistent (1)
|
|
|
|
|
|
|
38
|
Stipule shape
|
Code
|
Linear or lanceolate (0);
elliptic (1), Other (2)
|
|
|
|
|
|
|
39
|
Stipule length
|
Cm
|
|
0.33
|
2.05
|
0.989
|
0.4117
|
0.17
|
|
40
|
Stipule width
|
Cm
|
|
0.25
|
1.24
|
0.695
|
0.3188
|
0.102
|
RESULTS
Cluster analysis: Cluster analysis was performed on 40 quantitative and qualitative characters resulted, the separation of 36 taxa of the genus grouped into 4 clusters. The first main cluster was further divided into 4 subgroups; Subgroup I consist of E. bengalensis, E. bengalensis var. angustifolia, E. bengalensis var. intermedia, E. stipularis and E. dubia (Plate 1).

Plate 1. Morphological relationships among closely related species A). E. bengalensis, B). E. bengalensis var. intermedia, C). E. bengalensis var. angustifolia, D). E. dubia and E). E. stipularis. See Table 1 for details of the type specimens.
Subgroup II consists of E. serrata, E. obovata, E. poilanei, and E. elliptica var. petelotii (Plate 2). Subgroup III consists of E. longifolia, E. angustifolia, and E. petiolata (Plate 3). Subgroup IV consists of E. deflexa, E. deflexa var. buisanensis, E. cavaleriei, E. fulvicoma, E. glabrescens, and E. kwangsiensis (Plate 4). In cluster analysis, the taxa were clearly separated on the basis of first or second component or both (Figure 1). Both PCA and PCoA analysis were strongly support the cluster analysis (Figure 2 and 3).

Plate 2. Morphological relationships among closely related species A). E. serrata, B). E. elliptica var. petelotii, C). E. poilanei, D). E. dubia, E) E. obovata. See Table 1 for details of the Type specimens.
The second main cluster was also further divided into 4 subgroups; Subgroup V consists of E. japonica, E. × daduheensis and E. malipoensis (Plate 5). Subgroup VI consists of E. henryi, E. seguinii, E. prinoides, E. prinoides var. laotica and E. hookeriana (Plate 6 and 7). Subgroup VII consists of E. fragrans, E. fragrans var. furfuracea, E. elliptica, E. grandiflora, and E. salwinensis (Plate 8). Subgroup VIII consists of E. tengyuehensis, E. latifolia and E. glabrescens var. victoriensis (Plate 9). E. wardii and E. platyphylla formed a distinct cluster (Cluster III and IV) (Plate 8).

Plate 3. Morphological relationships among closely related species A). E. angustifolia, and B). E. longifolia. See Table 1 for details of the type specimens.

Plate 4. Morphological relationships among closely related species A). E. deflexa, B). E. deflexa var. buisanensis, C). E. glabrescens, D). E. cavaleriei, E). E. kwangsiensis, andF). E. fulvicoma. See Table 1 for details of the type specimens.

Plate 5. Morphological relationships among closely related species A). E. japonica, B). E. malipoensis, and C). E. × daduheensis. See Table 1 for details of the type specimens.

Plate 6. Morphological relationships among closely related species A). E. henryi, and B). E. seguinii. See Table 1 for details of the type specimens.

Plate 7. Morphological relationships among closely related species A). E. prinoides, B). E. prinoides var. laotica, and C). E. hookeriana. See Table 1 for details of the type specimens.

Plate 8. Morphological relationships among closely related species A). E. fragrans, B). E. fragrans var. furfuracea, C). E. elliptica, D). E. grandiflora, and E). E. salwinensis. See Table 1 for details of the type specimens.

Plate 9. Morphological relationships among closely related species A). E. tengyuehensis, B). E. glabrescens var. victoriensis, and C). E. latifolia. See Table 1 for details of the type specimens.

Plate 10. Morphological distinct and doubtful species A). E. wardii, and B). E. platyphylla. See Table 1 for details of the type specimens.
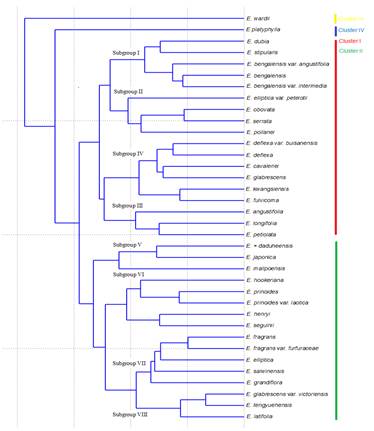
Figure 1. Cluster analysis on basis on both vegetative and floral characters using 36 taxa of Eriobotrya.
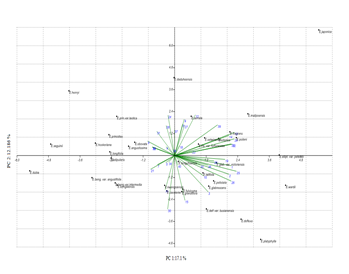
Figure 2. Principal components analysis according to the first 2 main factors performed on 40 morphological characters (PC1 x PC2).
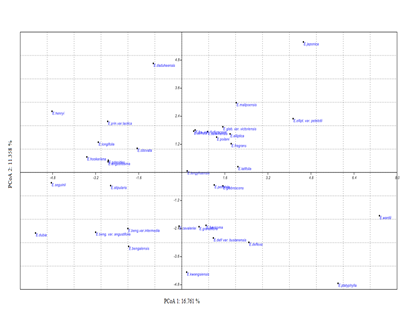
Figure 3. Principal coordinate analysis (PCoA) performed on 40 morphological characters
Principal component analysis (PCA) and Principal coordinate analysis (PCoA): Fourty quantitative and qualitative characters with taxonomic code, mean, standard deviation, and variance are illustrated in (Table 3). In principal component analysis, the first 12 principal component accounted for 84.8329 % of total variances, whereas the first 2 main principal components account for 17.1 % and 12.186 % respectively. The loading of the first to four principal’s components are presented in Table 2. The factor loading of > 0.6 was considered as significant. Ten characters showed highest correlation with the first axis, while two characters showed highest correlation with the second axis. Twelve characters showed that the positive factor loading > 0.6 on the first and second axis and included leaf length & width, bract shape, bract length and width, sepal length, petals length and width, petal state, style number, fruit length and width, whereas two characters showed negative factor loadings that > -0.6 on the second axis and included pedicel length and style state (free or connate). In addition, six characters showed that the positive factor loading > 0.6 on the third and fourth axis and these included pedicel state (glabrous or tomentose), calyx state (glabrous or tomentose), sepals outer surface, leaves abaxially tomentose or glabrous, petioles state (tomentose or glabrous), fruit length/fruit width, while there is no such characters that found to be listed in the negative factor loading that >-0.6 on the third and fourth axis.
Principal coordinate analysis (PCoA) was clearly separated the taxa on the basis of position on both side of the axis (PCoA eigenvalues and cumulative varianceare illustrated in Suppl. S2). PCoA scatter plot analysis supports the results of PCA analysis (Figure 1 and 2).
Table 3. Eigenvalues and cumulative variance for 4 major factors obtained from PCA analysis and 40 characters for each factor within all taxa of Eriobotrya.
|
PC
|
Eigenvalues
|
% variance
|
|
|
1
|
6.84006
|
17.1
|
|
|
|
2
|
4.87428
|
12.186
|
|
|
|
3
|
3.93641
|
9.841
|
|
|
|
4
|
3.17074
|
7.9268
|
|
|
|
5
|
2.63888
|
6.5972
|
|
|
|
6
|
2.04928
|
5.1232
|
|
|
|
7
|
1.98776
|
4.9694
|
|
|
|
8
|
1.70867
|
4.2717
|
|
|
|
9
|
1.60908
|
4.0227
|
|
|
|
10
|
1.39198
|
3.4799
|
|
|
|
11
|
1.29935
|
3.2484
|
|
|
|
No
|
PC1
|
PC2
|
|
1
|
0.7005
|
-0.1519
|
|
2
|
0.6738
|
-0.281
|
|
3
|
0.4164
|
-0.5474
|
|
4
|
-0.1061
|
0.1318
|
|
5
|
-0.1099
|
-0.1064
|
|
6
|
-0.3355
|
0.2166
|
|
7
|
-0.2146
|
-0.137
|
|
8
|
0.09734
|
0.4714
|
|
9
|
0.1179
|
0.546
|
|
10
|
0.3586
|
-0.3044
|
|
11
|
-0.1182
|
-0.5159
|
|
12
|
0.2567
|
0.613
|
|
13
|
0.7425
|
0.3492
|
|
14
|
0.6742
|
0.3034
|
|
15
|
0.1279
|
-0.6733
|
|
16
|
-0.2766
|
0.126
|
|
17
|
-0.2766
|
0.126
|
|
18
|
0.05974
|
0.1481
|
|
19
|
0.625
|
-0.0578
|
|
20
|
0.489
|
-0.08166
|
|
21
|
-0.2975
|
-0.2057
|
|
22
|
-0.276
|
0.1297
|
|
23
|
0.2161
|
0.07432
|
|
24
|
-0.08278
|
0.6056
|
|
25
|
0.7672
|
-0.2367
|
|
26
|
0.7024
|
-0.3841
|
|
27
|
0.1211
|
0.4546
|
|
28
|
-0.1051
|
0.4504
|
|
29
|
0.7408
|
0.2519
|
|
30
|
-0.08692
|
-0.8067
|
|
31
|
-0.2221
|
0.3634
|
|
32
|
0.7172
|
0.1713
|
|
33
|
0.701
|
0.1758
|
|
34
|
-0.07582
|
-0.09849
|
|
35
|
0.3179
|
-0.1436
|
|
36
|
0.4126
|
-0.1444
|
|
37
|
-0.00332
|
0.3861
|
|
38
|
-0.02257
|
0.09264
|
|
39
|
0.5322
|
0.4618
|
|
40
|
0.03905
|
-0.1388
|
* Number indicates 40 morphological characters.
DISCUSSION
The present study focused on the morphological patterns of variations and interspecies relationships reported here for the first time among 36 taxa of the genus Eriobotrya. Factor analysis revealed that leaf length and leaf width, bract shape, bract length and width, petal length and width, petal state (glabrous or villous), style number, and style state (free or connate), fruit length and width were the most informative characters for determination and identification of Eriobotrya taxa. The first and second component analysis using 40 characters clearly separated this genus into 4 clusters with 2 main clusters. The first main cluster consists of 18 species and was further divided into 4 subgroups. E. bengalensis, E. bengalensis var. angustifolia, E. bengalensis var. intermedia, E. stipularis, E. dubia formed a subgroup I. The morphological relationships with each other based on leaf blade abaxially glabrous, leaf size, 10 pairs of lateral veins, bract shape lanceolate, sepal shape triangular ovate, petal shape obovate and slightly pilose or glabrous, style connate at base and style number 2. Our results support the former classification and are consistent with previous analysis by using ITS sequences (Idrees et al. 2018, Idrees et al. 2020), and also with morphometric analysis of Zhang et al. (2017) that showed E. bengalensis, E. bengalensis var. angustifolia formed subgroup 1 in the group 5. E. bengalensis and E. stipularis totally close resembles to each other, and there are no such characters which can distinguish both species from each other except for stipule persistent or caducious. In addition, one of the sheets (isotype) of E. bengalensis “Wallich 668/2” kept at P barcode 02143254, the stipule is persistent and can be seen clearly. Another subgroup formed by E. obovata, E. serrata, E. poilanei, and E. elliptica var. petelotii. The relationship among these species mostly based on leaf blade abaxially glabrous, leaf length 15–25 cm, leaf width 4.5–9.5 cm, 10–20 pairs of lateral veins, bract obovate or elliptic, sepal shape ovate or rounded, petal shape obovate or cochelate, style connate at base and style number 2–5, leaf serration of all these taxa almost same. E. angustissima, E. longifolia and E. petiolata formed a subgroup and the relationships among these species based on leaf blade abaxailly glabrous, leaf length 15–20 cm, bract shape lanceolate, petal obovate, style connate at base and style number 2–5. E. deflexa, E. deflexa var. buisanensis, E.cavaleriei, E. kwangsiensis, and E. fulvicoma formed subgroup and the relationship among these species based on leaf blade abaxailly glabrous, leaf margin serrated, petal glabrous, style connate at base and style number 2–5. These results are consistent with ITS sequences analysis in our present work (Idrees et al. 2020), and also agreement with previous studies (Li et al. 2009, Yang et al. 2012, Zhang et al. 2017).
The second major cluster consists of 16 taxa and was further divided into 4 subgroups. E. japonica, E. malipoensis, and E. × daduheensis formed a subgroup and morphological relationship among these species based on leaf blade abaxailly tomentose, leaf size 16–22 cm, leaf width 4–8 cm, 12–18 pairs of lateral veins, petiole sessile and tomentose, large inflorescence 7–10 cm and contracted, bract subulate, pedicel sessile, style free at base and style number 3–5. Our results confirmed their close relationship, and are consistent with molecular study based on ITS sequences analysis (Idrees et al. 2020)) and also agreement with Yang (2005); Li et al. (2009); Yang et al. (2012); Yang et al. (2017); Zhang et al. (2017). E. henryi, E. seguinii, E. hookeriana, E. prinoides, and E. prinoides var. laotica formed a subgroup. Li et al. (2017) suggested that E. henryi and E. seguinii might be the most primitive taxa in the genus Eriobotrya. While E. hookeriana, E. prinoides and E. prinoides var. laotica formed a clade and have close relationship with each other’s. Our results confirmed their close relationship, and agreement with Yang et al. 2009, Li et al. 2009, Yang et al. 2017, and Idrees et al. 2020. E. fragrans, E. fragrans var. furfuracea, E. salwinensis, E. grandiflora, and E.elliptica formed a subgroup and morphological relationship among these species based on leaf blade axially tomentose, leaf length 12–16 cm, blade 3.5–6.5 cm, petiole >2 cm, 10–14 pairs of lateral veins, inflorescence size 4.5–9.5 cm, spreading, pedicel 0.2–6 mm, sepal ovate or rounded, petal obovate, petal length 7 × 5 cm, fruit length 1.2–1.3 × 0.9–1 cm. In addition, E. grandiflora was redescribed and confirmed to have a close relationship with E. fragrans (Idrees et al. 2020). Our results confirmed that E. grandiflora nested with E. fragrans, E. fragrans var. furfuracea, E. salwinensis, and E. elliptica. E. tengyuehensis, E. latifolia and E. glabrescens var. victoriensis formed a subgroups and relationship among these species based on leaf blade abaxailly tomentose, leaf width 5–7 cm, petal obovate and villous at base, petal length 4–5 × 3–4 mm, inflorescence size 7 cm. E. wardii and E. platyphylla formed a distinct cluster, however morphologically both species entirely different from the rest of the species. Both species can be identified on the basis of only type specimens and suggested here to be as doubtful species within this genus. Most of the species relationships were consistent with the phenetic analysis of morphological studies (Yang et al. 2017, Zhang et al. 2017), as well as molecular study (Li et al. 2009, Idrees et al. 2020).
Conclusions: We present here 40 morphological characters (quantitative and qualitative characters) using 36 taxa of the genus Eriobotrya. The morphometric analysis (cluster analysis, principal component analysis and principal coordinate analysis) distinguished well the whole taxa of the genus Eriobotrya. The most informative characters for determination and identification of taxa in the present study was leaf length and leaf width, bract shape, bract length and width, petal state (glabrous or villous), petal length and width, style number, and style state (free or connate), fruit length and width. Furthermore, E. wardii and E. platyphylla considered as the dubious species and it would be necessary to conduct molecular study to confirm its relationship with rest of the species. The quantitative and qualitative combinations of character studies will be particularly useful for further identification and systematic of the genus Eriobotrya.
Conflict-of-interest: All authors declared that there is no conflict of interest.
Acknowledgments: We are grateful to the curators and staffs of CENT, IBSC, KUN, PE, SCAU and SYS for their assistance in the study of specimens. Thanks are given to the reviewers for their helpful comments. This research was supported by grants from the National Natural Science Foundation of China (Grant Nos. 31670192 & 31110103911), and Science and Technology Basic Work, Project of the Ministry of Science and Technology of China (Grant No. 2013FY112100),Technology Department of Sichuan Province of China [2015JY0242], and the Major Scientific and Technological Achievements Transformation Project of Neijiang Normal University.
REFERENCES
- Ackerfield, J. and J. Wen (2002). A morphometric analysis of Hedera (The ivy genus, Araliaceae) and its taxonomic implications. Adansonia. Ser. 3, 24(2): 197-212.
- Badenes, M.L., J. Martinez-Calvo, and G. Llacer (2000). Analysis of a germplasm collection of loquat (Eriobotrya japonica). Euphyt. 114: 187-194.
- Deshmukh, S.A., S.R. Chavan, and D.K.Gaikwad (2012). Morphometric studies in the genus Clerodendrum Current Bot. 3(5): 41-44.
- Gu, S.Z. and S.A. Spongberg (2003). Eriobotrya Trans. Linn. Soc. London 1821, 13: 96, 102. F. China 9: 131-141.
- Hammer, O., D.A.T. Harper, and P.D. Ryan (2001). PAST: paleontological statistics software package for education and data analysis. 4(1): 9.
- Idrees, M., V.D. Troung, and X.F. Gao (2018). A new species of Eriobotrya (Rosaceae) from Con Dao National Park,southern Vietnam. Phytotaxa 365(3): 288-294.
- Idrees, M., A. Tariq, M.L. Pathak, X.F. Gao, S. Sadia, Z.Y. Zhang, and F.J. Zeng (2020). Phylogenetic relalationships of the genus Eriobotrya (Rosaceae) based on Nuclear Ribosomal DNA (ITS) Sequences. Pakistan J. Bot. 52(5): 1679-1684. http://dx.doi.org/10.30848/PJB2020-5(11).
- Kaiser, H.F. (1960). The application of electronic computers to factor analysis. Psychol. Meas. 20: 141-151.
- Kaplan, Z. and F.L.S.K. Marhold (2012). Multivariate morphometric analysis of the Potamogeton compressus group (Potamogetonaceae). Bot. J. Linn. Soc. 170: 112-130.
- Li, P., S.Q. Lin, X.H. Yang, G.B. Hu, and Y.M. Jiang (2009). Molecular phylogeny of Eriobotrya (loquat) infered from internal transcribed spacer sequences of nuclear ribosome. Pakistan J. Bot. 41(1): 185-193.
- Lindley, J. (1821). Observation on the natural group of plants called Pomaceae. Linn. Soc. London 13: 88-106.
- Nobis, M., K. Ewelina, N. Arkadiusz, D.G. Polina, and R. Kaja (2016). Multivariate morphometric analysis of the Stipa turkestanica group (Poaceae: Stipa sect. Stipa). Plant Syst. Evol. 302: 137-
- 13. Potter, D., T. Eriksson, R.C. Evans, S. Oh, J.E.E. Smedmark, D.R. Morgan, M. Kerr, K.R. Robertson, M. Arsenault, T.A. Dickinson, and C.S. Campbell (2007). Phylogeny and classification of Rosaceae. Plant Syst. Evol. 266: 5-43
- Sokal, P.R. and P.H. Sneath (1993). Principles of numerical taxonomy. W. H, Freeman, San Francisco.
- Vidal,E. (1965). Notes sur quelques Rosacées asiatiques (III) (Eriobotrya).Adansonia 5: 537-578.
- Yang, X.H. (2005). A systematic study of the genus Eriobotrya. Doctoral Dissertation of South China Agri. Uni., China. pp 1–99.
- Yang, X.H., K. Glakpe, S.Q. Lin, Y.L. Hu, Y. He, T.C.N. Nguyen, Y. Liu, G.B. Hu, and C.M. Liu (2005). Taxa of plants of the genus Eriobotrya around the world and native to Southeastern Asia. J. Fruit Sci. 22: 55-
- Yang, X.H. and S.Q. Lin (2007). New ideas on the classification of loquat. South China Fruits 36(3): 28-
- 19. Yang, X.H., C.M. Liu and S.Q. Lin (2009). Genetic relationships in Eriobotrya species as revealed by amplified fragment length polymorphism (AFLP) markers. Sci. Horti. 122: 264-268.
- Yang, X.H., P. Li, and Z.K. Zhang (2012). A Priliminarily phylogeny sudy of the Eriobtorya based on the nrDNA adh sequences. Bot. Horti Agrobot. Cluj-Napoca 40(2): 233-237.
- Yang, X.H., S.K. Najafabadi, M.Q. Shahid, Z. Zhang, Y. Jing, W. Wei, J. Wu, Y. Gao, and S. Lin (2017). Genetic relationships among Eriobotrya species revealed by genome-wide RAD sequence data. Ecol. Evol. 7: 2861-
- Yu, D.J. (1974). Delectis florae reipublicae popularis sinicae agenda academiae sinicae indita. In: Flora Reipublicae Popularis Sinicae, vol. 36, Science Press, Beijing, pp 260-
- Zhang, H.Z. and W.L. Qiu (1996). Flora of Chinese FrTit trees. Chapter Logan, Loquat. China Forestry Press, Beijing. pp 106-
- Zhang, Z., G.F. Li, X.H. Yang, and S.Q. Lin (2017). Taxonomic studies using multivariate analysis of Eriobotrya based on morphological traits. Phytotaxa 302(2): 122-
|





